1. Introduction to Molecular Cytogenetics
The transition from classical cytogenetics—characterized by gross chromatin staining and morphological observation—to the current era of molecular DNA techniques represents a strategic pivot in genomic sciences. While traditional banding methods provided an overview of chromosome behavior in mitosis and meiosis, they were fundamentally limited in their ability to resolve specific DNA sequences or discern the complex parental origins within allopolyploid nuclei. Modern molecular cytogenetics, primarily through In Situ Hybridization (ISH), bridges the gap between cytological observation and molecular sequence data. This shift is vital for mapping specific cloned sequences and “painting” whole genomes to identify alien gene introgression and intergenomic homogenization.
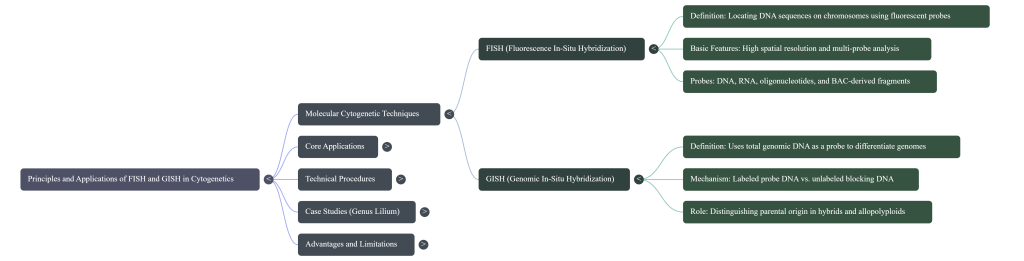
This technical guide synthesizes the mechanics and strategic applications of Fluorescence In Situ Hybridization (FISH) and Genomic In Situ Hybridization (GISH). Key themes include the analysis of genome structure in complex hybrids, the visualization of physical landmarks in allopolyploid evolution, and the role of these tools in interrogation breeding. We conclude with a discussion on the operational boundaries of these techniques, including the phenomenon of “genome turnover” in ancient polyploids.
The Strategic Shift: From Classical to Molecular Cytogenetics
The evolution of agricultural biotechnology necessitates a strategic transition from classical chromatin staining to molecular in situ hybridization (ISH). While classical techniques provided the foundation for our understanding of chromosomal diversity, they are largely limited to observing gross karyotypic alterations, such as changes in chromosome number and basic morphology. This shift enables researchers to move beyond these broad observations to precise, sequence-specific genomic interrogation. By utilizing ISH, we can anchor genetic data to physical chromosomal locations, providing high-resolution insights into genome organization and the behavior of specific genetic loci during mitosis and meiosis. This precision is the prerequisite for modern breeding strategies, particularly in identifying recombinant chromosomes and tracking alien chromatin segments during introgression.
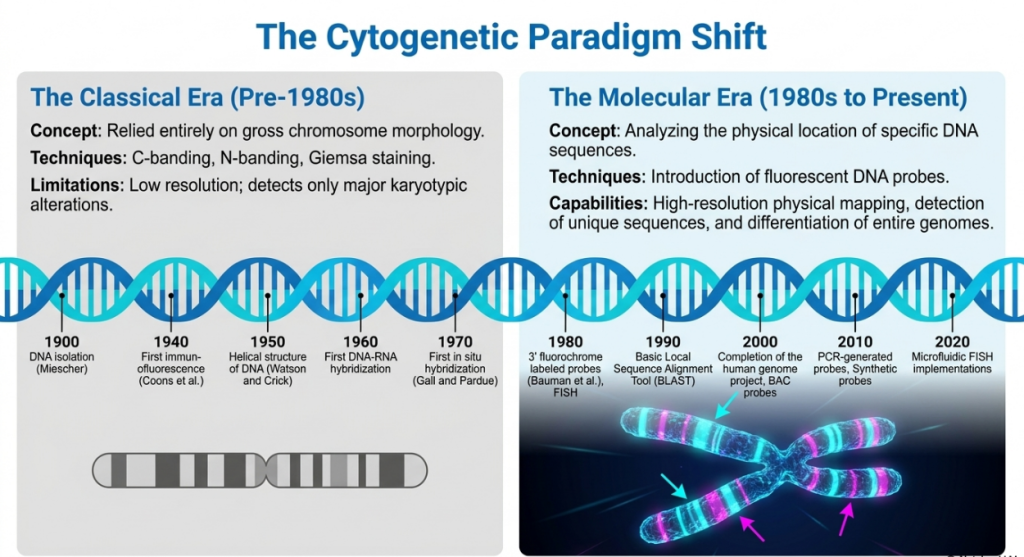
This transition addresses several critical limitations inherent in classical methods, as detailed below:
| Feature | Classical Cytogenetics (Chromatin Staining) | Molecular ISH (FISH and GISH) |
|---|---|---|
| Resolution | Restricted to gross morphology and large-scale alterations. | High-resolution mapping of specific DNA sequences and genes. |
| Reproducibility | Limited reproducibility; often highly labor-intensive. | High precision; utilizes consistent “monotonous markers” for identification. |
| Specificity | Inability to distinguish between chromosomes of similar size/shape. | Sequence-specific identification; distinguishes parental origins in complex hybrids. |
| Probes Used | General chemical dyes and stains (e.g., Giemsa, DAPI). | Sequence-specific DNA/RNA fragments or total genomic DNA (gDNA). |
| Application | Broad overview of chromosome number and meiotic behavior. | Identification of alien gene introgressions; localization of transgenes. |
| Data Depth | Limited to visible physical structures. | Integrated imaging of genetics, epigenetics, and chromatin folding. |
This refined molecular precision provides the technical framework for pinpointing genetic markers within a complex chromosomal landscape.
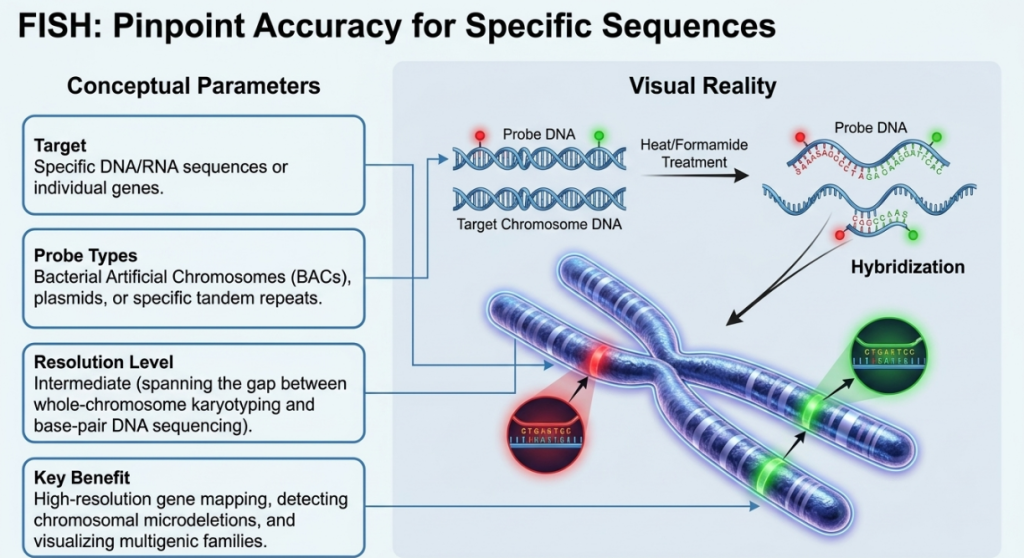
2. Principles and Technical Framework of FISH
Fluorescence In Situ Hybridization (FISH) functions as a high-resolution diagnostic and mapping tool that translates abstract molecular data into a spatial visual reality. It is predicated on the fundamental molecular properties of DNA: its double-stranded, paired characteristic and the ability of complementary sequences to anneal. By exposing organizationally preserved chromosomes to a short DNA sequence (a probe) labeled with a fluorescent molecule, we can physically map their exact location. This targeted approach provides a level of accuracy that classical banding cannot achieve, as it relies on high-degree sequence complementarity to “pinpoint” specific features within the genome.
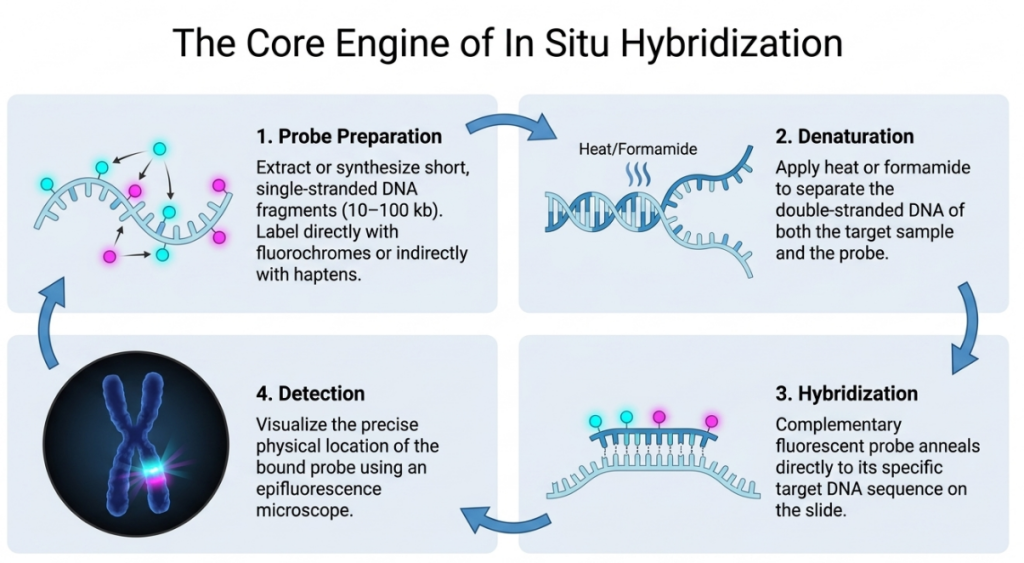
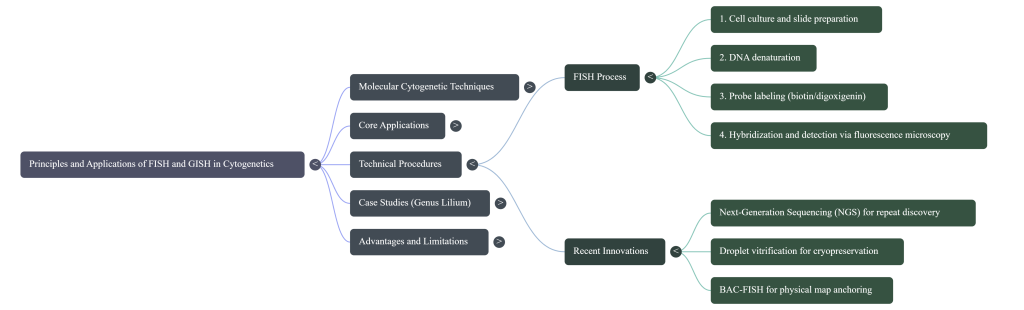
The FISH Protocol: A Step-by-Step Walkthrough
The FISH technique identifies particular DNA sequences by targeting them within an organizationally preserved cell. Here is the rigorous pathway we follow:
- Preparation (Metaphase and Denaturation): We prioritize cells in metaphase because the chromosomes are at maximum condensation, providing a discrete “map” for our probes. We then use Formamide (at 42°C) and heat (up to 75°C) for denaturation, which separates the double helix into single strands to make the target DNA accessible.
- Probing & Nick Translation: Probes are short fragments (typically 10 to 100 kb) often sourced from BAC (Bacterial Artificial Chromosome) libraries like the RPCI-11. We label these fragments using Nick Translation, a method that uses DNAase and DNA polymerase to incorporate labeled nucleotides into the probe.
- Hybridization: The labeled probe is applied to the slide, where it searches the genome and binds to its complementary sequence through annealing.
- Detection: Using an epifluorescent microscope, we visualize the results. If the sequence is present, it appears as a bright, glowing signal against the dark background of the chromatin.

Pro Tip: Direct vs. Indirect Staining
- Direct Staining: The fluorochrome is attached directly to the DNA probe, providing an immediate, though sometimes faint, signal.
- Indirect Staining: The probe is labeled with a hapten (Biotin/Digoxigenin). After hybridization, we add an antibody-fluorochrome conjugate (like Avidin or anti-Dig linked to a fluorophore). This antibody “finds” the hapten, amplifying the signal for superior clarity.
Moving from the individual “word” of a gene to the entire “dialect” of a species brings us to GISH.
Strategic Classification of ISH Probes
The selection of a probe foundation is a strategic decision based on the required diagnostic resolution and the scale of the target sequence.
| Probe Type | Foundation & Specific Diagnostic Function |
|---|---|
| DNA Probes | Short fragments (10–100 kb) used for detecting copy number variations and physical mapping of unique sequences. |
| RNA Probes | Targeted at mRNA/miRNA; critical for defining spatial-temporal patterns of gene expression and cellular activity. |
| Oligonucleotide Probes | High-specificity pairs (20 pairs covering 40–50 bp) for pinpointing miRNA and lncRNA locations. |
| BAC Probes | Bacterial Artificial Chromosomes (e.g., RPCI-11 library) (~100 kb); essential for anchoring contigs to physical maps and “painting” large segments. |
While FISH is peerless for identifying individual loci, the analysis of entire parental genomes within a hybrid nucleus requires the broader application of Genomic In Situ Hybridization.
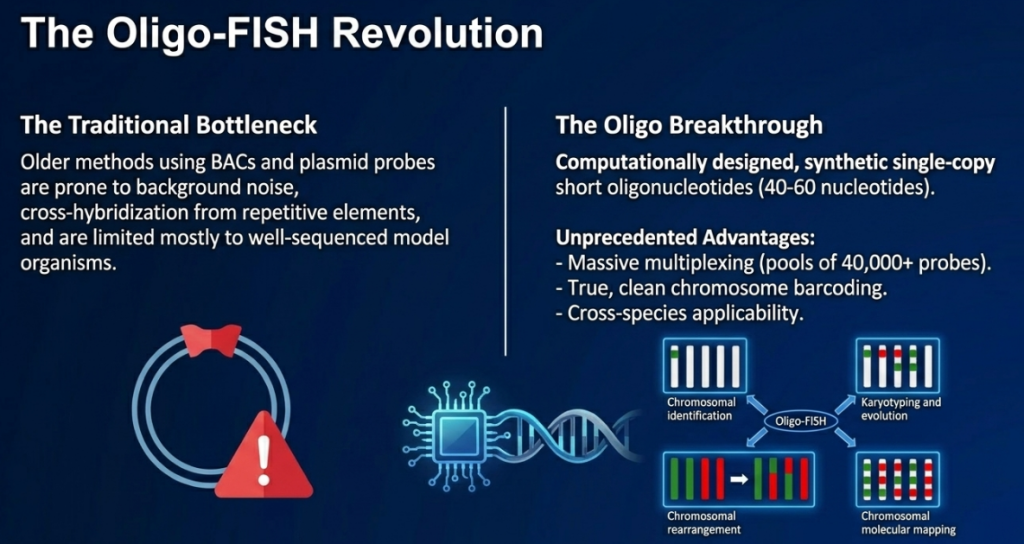
Primary Probe Targets and Their Strategic Utility
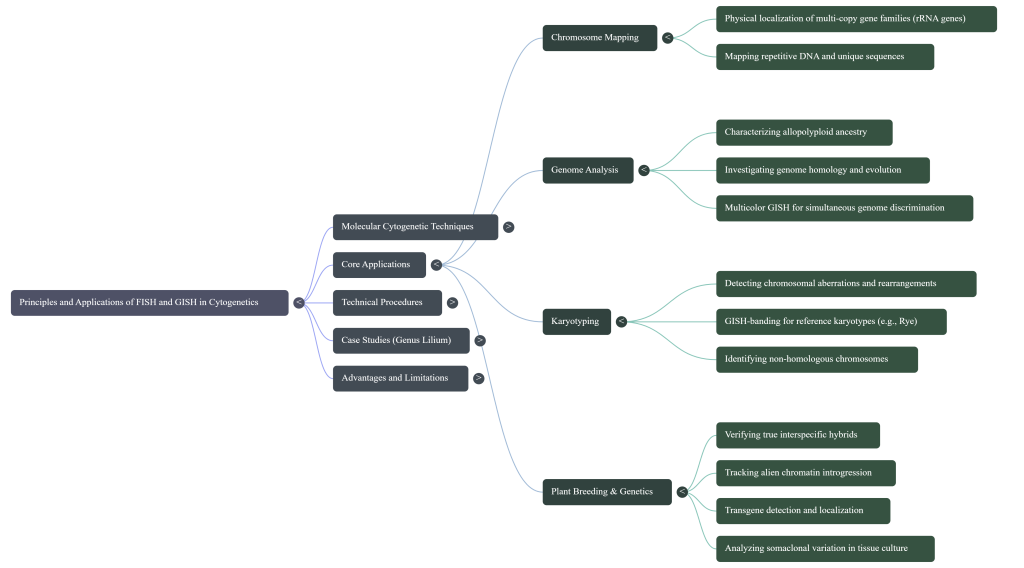
- Multi-copy Gene Families (rDNA): The 5S and 45S (18S-5.8S-26S) ribosomal RNA genes are among the most valuable landmarks for karyotype analysis. Because of their universal occurrence and high redundancy, they serve as reliable physical markers for identifying non-homologous chromosomes and characterizing evolutionary relationships across taxa.
- Repetitive DNA Sequences: Species-specific repetitive sequences, such as centromeric or telomeric repeats, are utilized to identify chromosomes accurately. These sequences are particularly useful in comparative genome studies and for tracking microstructural variations that occur during evolutionary development.
- Unique and Low-Copy Sequences: FISH is the primary method for localizing individual functional loci and transgene insertion sites. Utilizing probes derived from artificial chromosomes (e.g., BAC clones), researchers can determine the exact chromosomal position of specific genes, which is vital for understanding gene expression patterns.
- RNA Targets: Beyond DNA, FISH can localize mRNA and miRNA targets, allowing us to define spatial-temporal patterns of gene expression within tissues, bridging the gap between genomics and physiology.
Labeling and Visualization
The visualization of these targets is mediated by the incorporation of haptens—specifically biotin (Vitamin H) and digoxigenin (a steroid from Digitalis purpurea). These modified nucleotides are incorporated into the DNA probe through enzymatic labeling. Following hybridization, these haptens are detected using antibodies or avidin linked with fluorochromes. Under an epifluorescence microscope, these emit specific wavelengths of light, revealing the precise physical location of the target sequence.
While FISH identifies specific sequences, a specialized variant is required to “paint” and analyze entire subgenomes in complex plant hybrids.
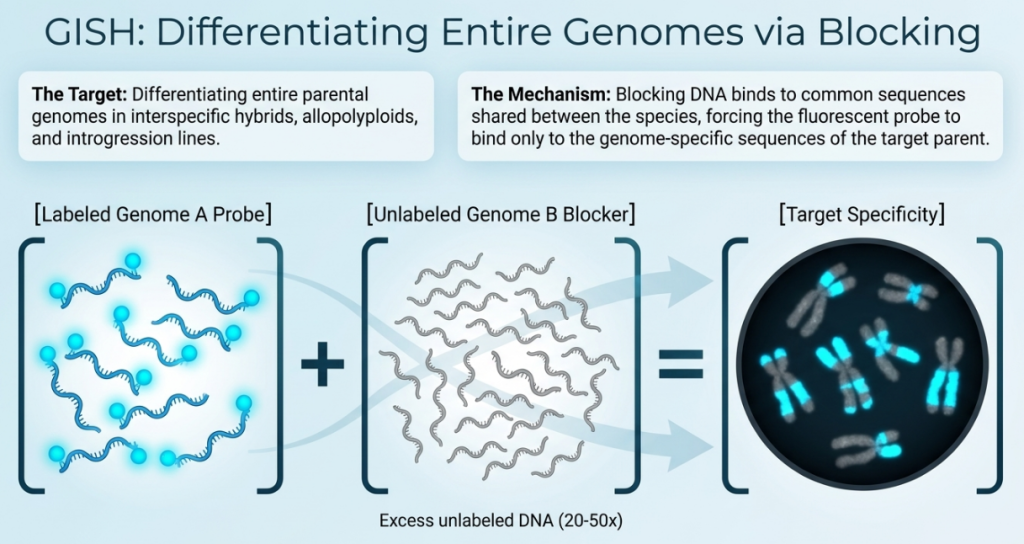
3. Genomic In Situ Hybridization (GISH): Mechanics and Parental Differentiation
Genomic In Situ Hybridization (GISH) is a specialized application of the FISH protocol that uses total genomic DNA from a parent species as a probe to distinguish genomes within a hybrid. By targeting species-specific dispersed repetitive sequences, GISH acts as a celestial map, pinpointing which parent contributed specific chromatin to complex allopolyploids. This makes it a primary tool for navigating the evolutionary history of our most vital crops and wild plants. GISH is an indispensable tool for investigating interspecific hybridization and allopolyploidy. By using total genomic DNA (gDNA) as a probe, GISH allows researchers to discriminate between the constituent genomes of a hybrid, a process critical for tracking alien gene introgression in “interrogation breeding.”
The case of the missing parent. A landmark example of GISH as a “molecular time machine” involved the study of Milium montianum, a species with 22 chromosomes. Using genomic painting, researchers identified a clear homology between its chromosomes and those of M. vernale (2n=8). This analysis didn’t just clarify a relationship; it successfully predicted the existence of an extinct or undiscovered ancestor, treating modern DNA as a living record of the past.
A tool for paleoreconstruction. Because GISH can pinpoint exactly where an ancestor’s DNA resides in a modern plant, it serves as a powerful method for reconstructing ancient genetic lineages. It allows scientists to look back through the fog of time to see how different lineages merged to form the diverse species we see today.
GISH has become a “powerful tool for analyzing interspecific and intergeneric hybrids and allopolyploid species as well as introgression, addition and substitution lines.”
Technical Variables and Blocking Ratios
The primary variable in GISH is the use of unlabeled “blocking” or “competitor” DNA. To prevent the probe from labeling common repetitive sequences shared by parental species—a frequent challenge due to homeologue similarity—blocking DNA is added in excess. In species with high homology, such as Brassica napus, a 10-fold to 100-fold excess of blocking DNA is required to achieve sufficient resolution and inhibit non-specific hybridization.
The Mechanism of “Blocking DNA”
The most critical technical distinction in GISH is the use of Blocking DNA. When analyzing hybrids of closely related species, high sequence homology can cause the probe to label both parental genomes simultaneously, obscuring results. To mitigate this, an excess of unlabeled genomic DNA from the non-target parent is added to the hybridization mixture. This unlabeled DNA “blocks” or pre-hybridizes to the shared common sequences, ensuring the labeled probe binds only to the sequences unique to the target genome. This significantly enhances the resolution of alien chromatin segments and translocation breakpoints.
Success in these applications depends on the rigorous execution of the experimental protocol.
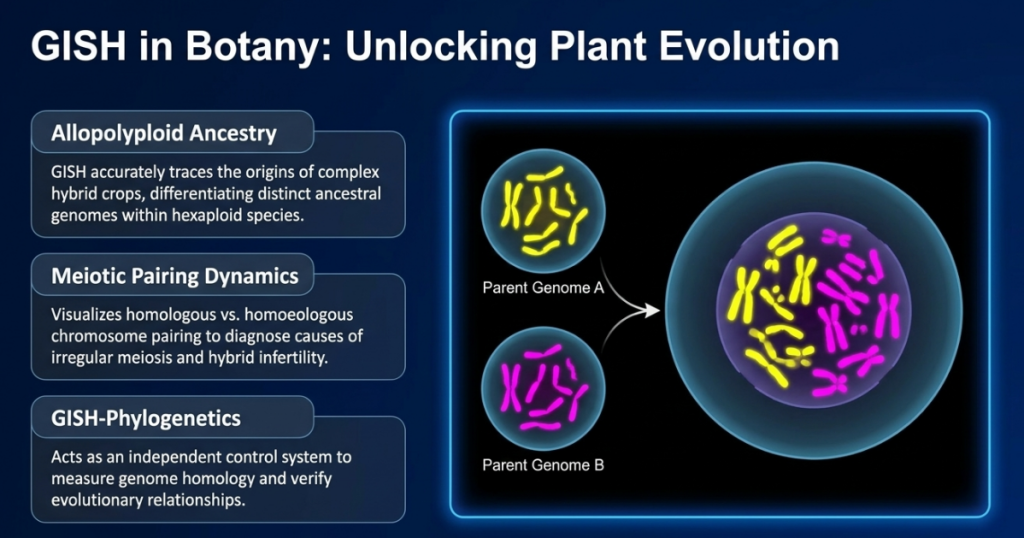
The Five-Million-Year “Memory Loss” of Plants
The limits of the luminescent map. Even the most powerful tools have their hurdles, and GISH faces two primary technical thresholds. First, it typically only works on natural polyploids if the hybridization event occurred less than five million years ago. Beyond this window, parental genomes undergo a “memory loss” known as homogenization, where DNA sequences evolve or are replaced so much that they can no longer be distinguished.
The size of the canvas. A second critical limitation involves the physical size of the genome being studied. Research suggests a 0.6 pg 1C genome size threshold; if the genome is smaller than this limit, the GISH labeling of entire chromosomes usually fails. In these cases, the “paint” cannot find enough repetitive targets to create a visible signal, leaving the chromosomes largely unlabelled except for concentrated regions like the centromeres.
Valuable failure. Far from being a mere technical setback, this five-million-year “failure” provides a valuable data point for evolutionary dating. If GISH fails to distinguish parental genomes in a known hybrid, scientists can infer that the species is likely older than five million years. This allows researchers to use genome turnover as a biological clock to reconstruct evolutionary timescales for long-term stabilization.
Repetitive Sequences: Unlike FISH, which may target unique genes, GISH relies on the fact that dispersed repetitive sequences evolve much faster than genes, providing the species-specific signatures required for painting.
Viral Stowaways: How Tobacco Built its Architecture
Discovery of viral fossils. Using FISH markers, researchers discovered surprising “stowaway” sequences within the tobacco (Nicotiana tabacum) genome known as Geminiviral Related DNA (GRD). These sequences are not merely random mutations but distinct, tandemly repeated clusters that represent ancient viral genetic material frozen within the plant’s DNA. FISH allowed scientists to map these clusters, revealing two independent insertion events from ancient viruses.
The viral-genetic hybrid machine. These integration events were remarkably complex, particularly in the N. tomentosiformis lineage that gave rise to modern tobacco. One insertion involved a sophisticated recombination between the virus and mobile genetic elements known as helitrons. This created a specialized genetic architecture, transforming a simple viral fossil into a complex machinery that shapes the modern crop’s genome.
Architectural insights. These repetitive remnants provide essential epigenetic and cytological information that dictates how the genome is organized. By identifying these specific genetic loci, science communicators can narrate the molecular sequences that transitioned ancient wild plants into stabilized agricultural staples.
The Tragopogon “Oopsie”: Perfect Numbers, Wrong Parents
Messy evolution in the wild. The study of “young” wild species like Tragopogon mirus, which formed in North America within the last 150 years, reveals that nature does not always demand perfection. These species often exhibit chromosomal irregularities that would be invisible to classical staining. GISH analysis has identified a phenomenon known as “reciprocal aneuploidy,” showcasing the inherent instability of early polyploid evolution.
The balance of error. In several instances, a plant may possess the “perfect” total number of chromosomes (2n=24) but have an uneven parental contribution. For example, a plant might be missing one chromosome from the T. dubius parent but have gained an extra one from the T. porrifolius parent. This resulting -1/+1 state creates a numerically correct but genetically unbalanced complement.
Evolutionary trial and error. While these meiotic “oopsies” may seem like errors, they are actually drivers of genetic diversity. These arrangements represent a mechanism for rapid genomic restructuring, where “errors” are eventually resolved by the stabilizing forces of selection. It is a vivid reminder that evolutionary stabilization often proceeds through a process of messy trial and error rather than rigid precision.
Strategic Value of GISH-Banding and Multicolor GISH
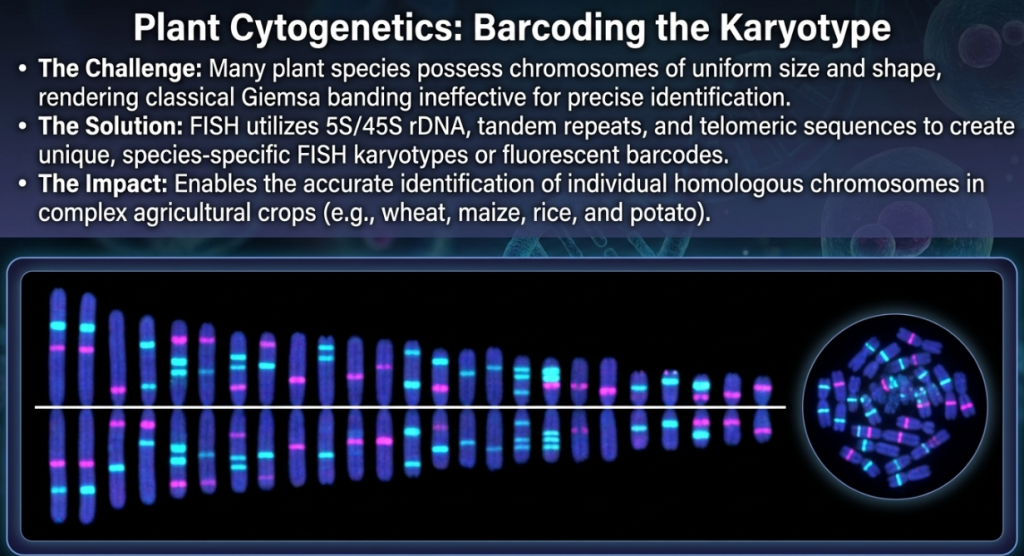
In many plant species, chromosomes are of uniform size and shape, making them indistinguishable by classical means. GISH overcomes this through the identification of species-specific dispersed repetitive sequences.
Multicolor GISH (mcGISH): A promising approach for simultaneously discriminating three or more genomes. A classic application is in hexaploid wheat (Triticum aestivum), where mcGISH can distinguish the A, B, and D genomes in a single spread by using different fluorophores—such as fluorescein (yellow/green) and rhodamine (red)—for different parental probes.
GISH-Banding: This occurs when genome-specific repeats form clusters within heterochromatin blocks. This strategy allows for the construction of a universal reference karyotype (e.g., for rye, Secale cereale) when other banding methods fail to provide sufficient resolution.
cenGISH (Centromeric GISH): In species with small genomes below a critical 1C size of approximately 0.6 pg, probes often preferentially label centromeric satellite repeats. These clusters allow for the definitive identification of parental origin even when the chromosome arms remain unlabeled.
selfGISH: The application of a species’ own gDNA to its chromosomes to observe the non-uniform distribution of repetitive elements, often revealing heterochromatic clusters.
Resolution and Genome Size Constraints
Resolution in GISH is governed by the critical 1C genome size of 0.6 pg. Below this threshold, total genomic labeling typically fails, as seen in certain small-genome Boechera genotypes, where signals may be restricted to pericentromeric regions unless probe concentrations and hybridization times are significantly increased.
4. Comprehensive Technical Execution Protocol
Precision in the experimental timeline—from cell harvest to microscopic observation—is vital for high-resolution imaging and the preservation of spatial relationships within the nucleus.
I. Slide Preparation and Cell Selection
Cells (typically from root tips) must be harvested at the metaphase stage of division. This timing is critical because chromosomes reach their maximum state of condensation during metaphase, providing the clearest targets. Cells are fixed and prepared on microscopic slides, often treated with enzymes like Pectinex to remove the cytoplasm, which can act as a physical barrier to the probe.
II. Probe Preparation and Labeling
Probes are typically 10–100 kb fragments. A standard method for labeling is Nick Translation, utilizing:
- DNAase: To create “nicks” or random cuts in the DNA.
- DNA Polymerase: To replace nucleotides at the nick sites.
- Modified Nucleotides: The polymerase incorporates nucleotides labeled with haptens like Dig-dUTP or Biotin-dUTP.
III. Denaturation and Hybridization
Both the chromosomal DNA and the probe must be denatured to become single-stranded to allow for complementary base pairing. Chromosomal DNA is typically heated to 70-75°C. The hybridization phase is then conducted at a lower temperature—classically 42°C—in a solution containing Formamide to lower the melting temperature and ensure stringent, specific annealing of the probe to the target.
IV. Detection and Visualization
Following hybridization, the slides undergo stringent washes to remove unhybridized probes. The labeled sites are detected using antibodies linked with fluorophores (e.g., Anti-Dig or Avidin-FITC). The results are observed via epifluorescence microscopy, where target signals appear as distinct fluorochrome spots or “painted” segments against DAPI-counterstained (blue) chromosomes.
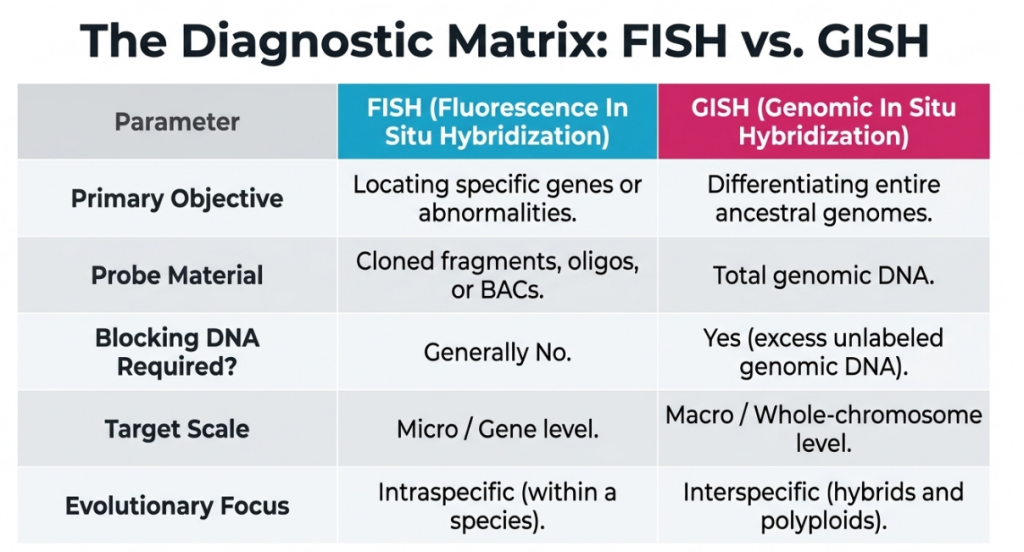
5. Comparative Analysis: FISH vs. GISH
The distinction between FISH and GISH lies in the transformation of raw fluorescence data into evolutionary and breeding insights.
| Feature | FISH | GISH |
|---|---|---|
| Probe Foundation | Specific cloned fragments or oligonucleotides. | Total genomic DNA of a parental species. |
| Primary Objective | Mapping single loci, transgenes, or aberrations. | Discriminating subgenomes and intergenomic interactions. |
| Typical Target | Repetitive arrays (5S/45S rDNA) or unique genes. | Entire chromosomes or alien gene introgression sites. |
| Blocking DNA | Used to reduce non-specific background | Used to reduce the non-specific background |
| Spatial Insight | Pinpoints loci on the chromosome map | Reveals genome territories in the nucleus |
| Visualization | Target gene location/copy number | Intergenomic recombination and genome organization |
GISH serves as an “authentic model” for identifying intergenomic recombination. It allows the visualization of allosyndetic pairing (intergenomic pairing), providing a macroscopic view of how genomes stabilize or restructure following a polyploidization event.
6. Strategic Application in Breeding and Introgression Analysis
In modern agricultural biotechnology, cytogenetic validation serves as the “ground truth” for breeding programs, ensuring the stability and success of wide crosses and wide-hybridization events.
Karyotyping and Physical Mapping
Visualization of multigenic families (5S and 18S-26S rRNA) provides high-resolution landmarks for karyotype analysis in complex species. For instance, in Agave fourcroydes (2n=5x=150), FISH using telomeric repeat sequences is essential for managing the high ploidy levels and detecting aneuploidy within an enormous chromosomal complement.
Interrogation Breeding and the Ph1 Locus
In cereal breeding, GISH is the strategic lever for tracking the transfer of resistance genes. It is particularly used to monitor the Ph1 locus in wheat, which regulates homeologous pairing. By suppressing this locus, researchers can induce allosyndetic recombination, and GISH serves as the primary verification tool for the resulting alien chromatin introgressions.
GISH-Phylogenetics
A sophisticated application is the use of the “DA” (distinguishing ability) parameter. By measuring the ratio of signal intensities between parental genomes, researchers can quantify species relatedness. Notably, log DA values show a 0.98 correlation with DNA sequence-based phylogenetic trees, providing a robust, independent control for molecular systematics.
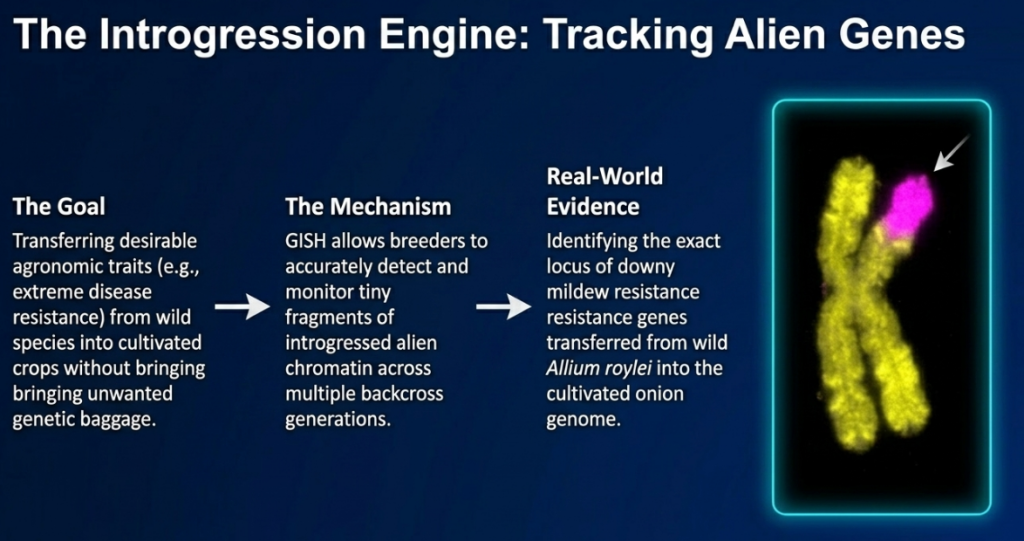
Alien Gene Introgression and Resistance Tracking
GISH is the primary tool for tracking the integration of resistance genes from wild relatives into cultivated lines. A high-impact example is the detection of the Fhb3 gene for resistance to Fusarium head blight, introgressed from wild relatives into wheat. GISH allows us to monitor the integration of these wild chromosomal segments and verify their maintenance through successive backcrosses.
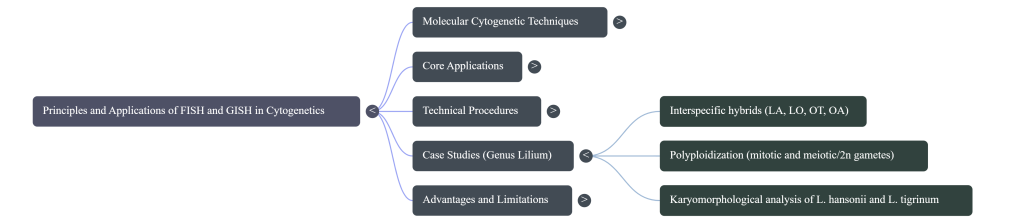
Validation of Interspecific Hybrids and Meiotic Behavior
GISH is essential for distinguishing true hybrids from offspring produced via apomixis. In genera like Lilium, GISH unequivocally identifies true hybrids by visualizing both parental genomes. This allows for the analysis of:
- Karyomorphology: Classifying chromosomes as metacentric, sub-telocentric, or telocentric based on centromere position and shape.
- Recombinant Chromosomes: Identifying homoeologous recombination and translocation breakpoints.
- Meiotic Behavior: Analyzing whether chromosomes form stable bivalents or lead to infertility through univalents or multivalent formation (allosyndetic pairing).
Somaclonal Variation and Transgene Detection
Tissue culture stress often induces genetic instability, or somaclonal variation, characterized by chromosome breakage and DNA transposition. Tracking 5S and 18S-26S rRNA gene distribution is the specific method used to identify these genomic changes. Furthermore, FISH is used to determine the chromosomal position of inserted transgenes. This is vital because the local chromatin structure and physical position of a transgene can induce gene silencing or profoundly impact expression levels.
7. Operational Challenges, Limitations, and Future Horizons
Despite their utility, FISH and GISH face inherent biological and technical constraints.
Operational Advantages
- Single-Cell Analysis: Allows for the study of individual nuclei and the preservation of interphase nuclear organization.
- No Amplification Bias: Unlike PCR-based methods, ISH does not require technology-based amplification, which can lead to false positives or negatives.
- Spatial Integrity: Preserves the spatial relationship of DNA sequences, providing the physical context for sequence data.
Operational Constraints
- High Expertise and Cost: The techniques are labor-intensive, requiring specialized equipment and high levels of expertise in microscopy and cytogenetic counting.
- Detection Limits and Small Genomes: FISH has a high detection limit (approx. 10^6 cells/mL). Furthermore, standard GISH often fails in species with a 1C genome size below 0.6 pg.
- Mitigation Strategies: For small-genome species like Arabidopsis suecica, successful labeling of entire chromosomes can be achieved by using exceptionally high concentrations of hybridizing gDNA and extended hybridization times.
The “Genome Turnover” Limit
A critical limitation of GISH is the evolutionary timeframe. In Nicotiana species, GISH failure occurs in polyploids older than 5 million years. This indicates a near-complete genome turnover and intergenomic homogenization, where parental sequences have diverged or homogenized to the point of being indistinguishable. This suggests GISH is most effective for recently formed allopolyploids.
Next-Generation Integration
The future of cytogenetics involves the integration of Next-Generation Sequencing (NGS). This synergy will facilitate the rapid characterization of repetitive fractions, enabling the design of “synthetic” GISH probes and the physical mapping of genomes where traditional genomic resources are unavailable.
8. Conclusion and Technical Summary
FISH and GISH are the most reliable techniques for the structural and evolutionary analysis of allopolyploids. While FISH provides the precision to map individual transgenes and loci, GISH offers a macroscopic view of parental genome interactions. Together, they form the essential “bridge between cytological and molecular approaches,” providing the strategic foundation for future genetic engineering and species identification in both research and industry.
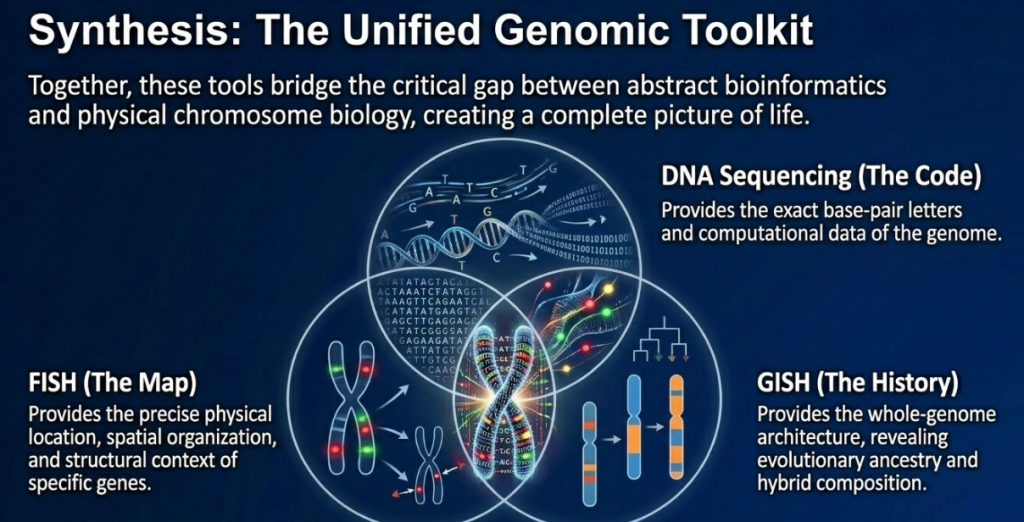
The integration of FISH and GISH with modern Next-Generation Sequencing (NGS) represents the next frontier in agricultural research. While NGS provides the genomic “code,” molecular cytogenetics provides the “context,” anchoring sequence data to physical chromosomal locations. This synergy is essential for understanding intergenomic homogenization and the mechanisms of gene regulation in complex polyploid crops.
As we continue to refine these tools, the ability to precisely manipulate and validate genome architecture will be paramount. FISH and GISH remain indispensable in the global effort to secure sustainability and crop resilience, ensuring that the innovations of chromosome engineering are successfully translated from the laboratory to the field.
Image Summary
Questions/Answers
1. What are the primary differences between FISH and GISH?
The primary differences between Fluorescence In Situ Hybridization (FISH) and Genomic In Situ Hybridization (GISH) revolve around the type of DNA used as a probe, the specific methodological requirements, and their intended research or diagnostic objectives. While GISH is a specialized modification of the FISH technique, they serve distinct roles in molecular cytogenetics.
1. Probe Composition and Target
- FISH utilizes small, known DNA sequences as probes to locate and visualize specific genes or repetitive sequences on a chromosome. These probes can range from short 10–100 kb fragments to modern synthetic oligonucleotides (oligos). FISH targets locus-specific regions, such as ribosomal DNA (rDNA) gene families (5S and 18S-26S), centromere-specific repeats, or individual genes.
- GISH uses the total genomic DNA (gDNA) of a species as the probe. Rather than looking for a specific gene, GISH targets the entire set of chromosomes from a particular parent or ancestor to differentiate them from other genomes in a hybrid specimen.
2. Methodological Requirements (Blocking DNA)
A defining technical difference is the use of blocking DNA in GISH, a step that distinguishes it from standard FISH analysis.
- In GISH, labeled total genomic DNA from one species is hybridized alongside a large excess (often 20- to 100-fold) of unlabeled “blocking” DNA from a related species. This blocking agent binds to repetitive sequences shared between the two species, preventing the labeled probe from cross-hybridizing and allowing only the species-specific sequences to be visualized.
- While FISH protocols may use certain blocking agents (like Cot-1 DNA) to suppress general repetitive sequences, it typically does not require the specific parental genomic blocking used in GISH to distinguish whole genomes.
3. Primary Objectives and Applications
- FISH is primarily used for physical mapping, identifying individual chromosomes within a genome (karyotyping), and detecting chromosomal aberrations such as deletions, translocations, and inversions. It is widely applied in medical science for genetic counseling, diagnosing cancers (oncology), and identifying infectious microbes.
- GISH is used for genome differentiation in interspecific hybrids and allopolyploids. Its main goals include determining the parental origin of chromatin, identifying alien gene introgressions from wild relatives into crops, and studying evolutionary relationships and genome stability in polyploid species.
4. Field of Dominance
- FISH is a paradigm-changing tool used extensively across human medicine and general biological research to relate DNA sequence information to chromosome biology.
- GISH has found its most significant niche in plant and horticultural sciences, particularly in breeding programs and the study of crop evolution, due to the high frequency of polyploidy and hybridization in the plant kingdom.
2. How do these techniques help in plant breeding and hybridization?
Molecular cytogenetic techniques, specifically Fluorescence In Situ Hybridization (FISH) and Genomic In Situ Hybridization (GISH), provide powerful GPS-like tools for plant breeders to visualize and manipulate the genetic blueprint of crops. These techniques help overcome traditional breeding challenges by allowing researchers to see directly where specific DNA sequences or entire sets of parental chromosomes are located within a cell.
The primary ways these techniques assist in plant breeding and hybridization include:
1. Verification and Confirmation of Hybrids
Breeding often involves crossing distantly related species to combine desirable traits. FISH and GISH are essential for hybridity status confirmation—proving that a cross was successful.
- True Hybrid Status: GISH can unequivocally identify if a plant is a true hybrid or if it developed through apomixis (reproduction without fertilization), in which case the offspring would only contain maternal genetic material.
- Early Selection: Breeders can identify hybrid status even during the vegetative phase, significantly accelerating the breeding process by avoiding years of waiting for plants with long juvenile periods to flower.
- Paternity Determination: GISH can distinguish and identify donor parental genomes in complex interspecific hybrids, such as those in Rhododendron, Clivia, and Lilium.
2. Monitoring Alien Gene Introgression
In introgression breeding, useful genes (such as those for disease resistance or drought tolerance) are transferred from wild relatives into cultivated crops.
- Tracking Alien Chromatin: GISH acts as a visual monitor, allowing researchers to see the amount and location of alien chromatin (DNA from the wild relative) that has been successfully integrated into the crop’s genome.
- Detecting Translocations: These techniques help identify translocation breakpoints, where a piece of one chromosome has attached to another. For example, GISH has been used in wheat to detect rye segments carrying resistance to leaf, stem, and stripe rust.
- Removing Linkage Drag: By precisely mapping desirable genes, breeders can work to eliminate “linkage drag”—the unwanted wild genetic material that often comes along with a beneficial gene.
3. Analysis of Genomic Constitution and Ploidy
Many major crops, such as wheat, cotton, and sugarcane, are polyploids (having multiple sets of chromosomes).
- Genome Characterization: FISH and GISH help determine the genomic composition and origin of polyploid species, identifying which ancestral species contributed which chromosomes.
- Determining Ploidy Levels: These techniques effectively distinguish different ploidy levels (e.g., diploid, triploid, tetraploid) and can detect aneuploidy—the presence of an abnormal number of chromosomes.
- Confirming Stability: GISH contributes to analyzing the karyotypic stability of new breeding lines, helping breeders select the most genetically stable genotypes for further development.
4. Studying Meiotic Behavior and Recombination
Understanding how chromosomes pair and exchange genetic material during cell division (meiosis) is crucial for fertility and trait stability in hybrids.
- Chromosome Pairing: GISH allows for the visualization of homologous (identical) and homoeologous (partially similar) chromosome pairing during meiosis.
- Recombination Patterns: Researchers can analyze the patterns and frequency of genetic recombination (crossovers) between different genomes.
- Predicting Fertility: By observing irregular meiotic behavior (like lagging chromosomes or bridges), these techniques can help explain reduced fertility in certain hybrids and guide strategies to restore it.
5. Physical Mapping and Gene Localization
FISH is the primary tool for physical mapping—determining the exact physical location of a gene or DNA sequence on a chromosome.
- Integrating Maps: FISH mapping helps integrate genetic linkage maps (based on recombination) with physical chromosomal maps, which is vital for accurate gene positioning and sequencing efforts.
- Identifying Gene Clusters: Techniques like Fiber-FISH provide high resolution to analyze the organization of multigene clusters and repetitive DNA sequences.
- Transgene Detection: FISH is used to identify the specific integration sites of transgenes in genetically engineered plants, helping researchers understand how their position affects gene expression.
6. High-Precision Breeding with Oligo-FISH
The recent development of oligo-FISH, which uses synthetic oligonucleotide probes, has further refined these capabilities.
- Chromosome Barcoding: Ample sets of short probes can create a unique “barcode” for every chromosome in a species, allowing for the identification of an entire set of individual chromosomes in a single experiment.
- Precise Introgression Tracking: This technique increases the precision of detecting small introgressed segments and chromosomal rearrangements never before recordable.
- Linking Traits to Collinearity: Recent studies in wheat have used oligo-FISH to link specific changes in chromosome collinearity with superior resistance to stripe rust.
3. What are the common limitations or challenges in using GISH?
The common limitations and challenges in using Genomic In Situ Hybridization (GISH) primarily revolve around the genetic relatedness of the species being studied, the size of their genomes, the age of the hybridization events, and various technical complexities.
Genetic Relatedness and Cross-Hybridization
The most significant challenge in GISH is the high degree of sequence homology between closely related parental species.
- Non-specific Hybridization: If parental genomes are very similar, the labeled genomic probe may bind to both sets of chromosomes, making them impossible to distinguish.
- Blocking DNA Issues: To combat cross-hybridization, unlabeled “blocking” DNA from one parent is used to saturate common sequences, but determining the exact ratio of probe to blocking DNA is difficult and must often be optimized empirically for each species.
- Stringency Requirements: Closely related genomes often require increased stringency conditions—such as higher temperatures, higher formamide concentrations, or longer wash durations—which can sometimes weaken the target signal to the point of being undetectable.
Genome Size and Labeling Constraints
The physical size of the genome directly impacts the success of the GISH technique.
- Small Genome Failure: There is a critical 1C genome size limit of approximately 0.6 pg; below this threshold, GISH labeling of entire chromosomes typically fails.
- Regional Labeling: In species with very small genomes, probes may preferentially label only the pericentromeric regions or repetitive clusters (like ribosomal DNA), leaving the rest of the chromosome arms uncolored.
- Large Repetitive Fractions: Conversely, in plants with very large genomes like wheat, the high percentage of repetitive DNA can create significant background noise, complicating the localization of unique segments.
Evolutionary Time and “Genome Turnover”
GISH is most effective on recent hybrids and allopolyploids.
- Ineffectiveness in Old Species: Studies in Nicotiana demonstrate that GISH failure represents a near-complete genome turnover associated with long-term diploidization.
- The 5-Million-Year Rule: For many natural allopolyploids, GISH is only successful if the polyploidy event occurred less than 5 million years ago; beyond this age, distinguishing parental genomes becomes impossible due to sequence homogenization and DNA loss.
Technical and Operational Hurdles
Beyond biological limits, GISH is constrained by laboratory-level difficulties.
- Labor and Cost: The method is described as slow, labor-intensive, and expensive compared to molecular markers, requiring specialized expertise for protocol development.
- Sample Quality: Results are highly dependent on the quality of chromosome spreads; dense cytoplasm can act as a physical barrier that prevents probes from reaching their targets, and over-condensed chromosomes can lead to overlapping signals.
- Resolution Limits: While powerful for whole-genome differentiation, GISH is less accurate for identifying very small recombinant segments or single genes compared to sequencing-based methods.
- Misinterpretation: Variations in signal strength can lead to the misidentification of chromosomes; for instance, a weak signal on a single-species chromosome can be misinterpreted as a translocation from a different genome if not carefully controlled.
4. What is blocking DNA and why is it used?
Blocking DNA is a critical component used primarily in Genomic In Situ Hybridization (GISH) and specialized Fluorescence In Situ Hybridization (FISH) protocols to ensure the accuracy and specificity of DNA labeling. It consists of unlabeled, fragmented DNA that is added to the hybridization mixture alongside the labeled probe.
What is Blocking DNA?
- Composition: It is typically derived from the total genomic DNA (gDNA) of a species related to the target, such as the second parent in a hybrid cross. Other common blocking agents include Cot-1 DNA (enriched for repetitive sequences) or sheared salmon sperm DNA.
- Physical State: The DNA is fragmented into very short pieces, usually ranging from 100 to 300 base pairs in length.
- Concentration: It is used in large excess compared to the labeled probe, typically at a ratio of 20-fold to 100-fold.
Why is it used?
The primary purpose of blocking DNA is to prevent non-specific hybridization and “cross-talk” between related genomes. Its specific functions include:
- Saturating Shared Sequences: Closely related species often share a high degree of sequence homology, particularly within repetitive DNA elements. During hybridization, the unlabeled blocking DNA binds to these common sequences first, “blocking” them from the labeled probe.
- Enhancing Species-Specificity: By saturating the shared sequences, the labeled probe is forced to hybridize only with the unique, species-specific sequences of the target genome. This allows researchers to see a clear distinction between parental sets of chromosomes in a hybrid, which would otherwise look identical.
- Increasing Signal Contrast: Blocking DNA reduces “background noise” or unwanted fluorescent signals on non-target chromosomes. This is essential for identifying alien gene introgressions, where a tiny fragment of DNA from one species is integrated into the genome of another.
- Managing High Homology: In cases where parental genomes are nearly identical, blocking DNA is used under increased stringency conditions (higher temperatures or chemical concentrations) to ensure that only the most perfectly matched DNA sequences are visualized.
5. How does multicolor GISH (McGISH) work in wheat?
Multicolor Genomic In Situ Hybridization (McGISH) is an advanced cytogenetic technique used to simultaneously discriminate and visualize multiple genomes within complex polyploid species like wheat. In hexaploid wheat (Triticum aestivum), which contains three distinct genomes (A, B, and D), McGISH works by using different combinations of labeled and unlabeled total genomic DNA (gDNA) from ancestral progenitor species.
The specific process for distinguishing the three wheat genomes involves the following components:
- Probes: Researchers typically use biotinylated total gDNA from the A-genome progenitor (Triticum urartu) and digoxigenin-labeled total gDNA from the D-genome progenitor (Aegilops tauschii).
- Blocking DNA: Non-labeled total gDNA from a possible B-genome progenitor, such as Aegilops speltoides, is added to the hybridization mixture in excess. This unlabeled DNA acts as a competitor, binding to repetitive sequences shared across all three genomes to prevent cross-hybridization and ensure the labeled probes only bind to their specific target chromosomes.
- Visualization: Through the use of different fluorochromes (such as fluorescein and rhodamine), all three genomes can be detected at once. In a successful experiment, the A-genome chromosomes fluoresce yellow, the B-genome chromosomes appear blue, and the D-genome chromosomes fluoresce red.
McGISH is particularly valuable in wheat research for identifying alien gene introgressions and tracking chromosomal rearrangements. For example, it has been used to visualize intergeneric translocations in wheat-rye lines and to analyze the genomic constitution of trigeneric hybrids involving wheat and wheatgrass relatives. This technique allows for the detection of alien chromatin even when it has been integrated into the wheat background as small segments. Furthermore, McGISH has been employed to study the evolutionary origin of polyploid wheat by characterizing the amount of pairing and homology between the various sub-genomes. Recent advancements have also integrated oligo-FISH, which uses synthetic oligonucleotide probes to provide even higher precision in labeling individual wheat chromosomes and sub-genomes.
6. Can GISH determine the age of a polyploid species?
While Genomic In Situ Hybridization (GISH) cannot provide a precise chronological date like carbon dating, it is used as a powerful tool in GISH-phylogenetics to estimate the relative evolutionary age and reconstruct the time scales of polyploidization.
This application relies on the biological phenomenon of genome turnover and intergenomic homogenization. The ability of GISH to determine age is based on the following principles:
- The 5-Million-Year Threshold: Research in natural Nicotiana allopolyploids has shown that GISH is generally only effective for distinguishing parental genomes if the polyploidy event occurred less than 5 million years ago.
- Genome Turnover: In polyploid species of this age or older, distinguishing parental genomes using GISH often becomes impossible. This failure indicates a near-complete genome turnover associated with long-term genome diploidization, where parental DNA is lost, retroelements become active, and intergenomic sequences homogenize.
- Reconstructing Time Scales: By comparing hybridization patterns in allopolyploids of different known ages—including synthetic allopolyploids that mimic natural species—researchers can reconstruct the evolutionary time scales for when these genomic changes occurred.
- Quantitative Measurement of Divergence: The intensity and pattern of GISH signals can provide a quantitative measure of chromosome divergence. As the evolutionary distance between species increases, the ability of a genomic probe to distinguish parental sets of chromosomes diminishes.
In summary, GISH helps determine the age of a polyploid species by acting as a visual indicator of how much the parental genomes have merged or “turned over” since the original hybridization event.
7. What is apomixis and why is GISH used to detect it?
Apomixis is a biological phenomenon in plants where reproduction occurs without fertilization. Essentially, it is a form of asexual crossing that results in offspring that are typically genetically identical to the maternal parent.
Genomic In Situ Hybridization (GISH) is used to detect apomixis primarily for the following reasons:
- Hybridity Status Confirmation: In plant breeding, researchers often attempt to cross distantly related species to create new hybrids. However, if the maternal plant is apomictic, the resulting seeds might actually be clonal copies of the mother rather than successful crosses. GISH is the most effective tool to unequivocally identify whether a plant is a “true hybrid” (containing DNA from both parents) or an apomictic offspring (containing only maternal DNA).
- Visualizing Genomic Differences: Because GISH can distinguish between entire parental genomes, it can clearly show the presence or absence of the paternal genome in a single cell. If a cross was successful, GISH will reveal two distinct sets of chromosomes; if apomixis occurred, only one parental genome will be visible.
- Detection of Chromosomal Aberrations: In certain species complexes, such as Boechera holboellii, diploid apomicts are known to display large-scale chromosome substitutions and aberrant chromosomes. GISH allows researchers to visualize these structural novelties and identify the specific parental origin of individual chromosomes within these complex groups.
- Studying Polyploidy Origins: Apomixis is often linked to polyploidy. GISH helps researchers understand the relationship between these two phenomena by providing insights into the chromosome number and the constitution of functional gametes in apomictic lineages.
8. How can these techniques help remove ‘linkage drag’ in crops?
Linkage drag refers to the unwanted wild genetic material that is unintentionally transferred along with a beneficial gene during introgression breeding. Because these undesirable traits are physically linked to the target gene on the same chromosome, they are often difficult to separate using traditional breeding methods alone.
Molecular cytogenetic techniques like FISH and GISH help remove this drag by providing a visual “GPS” for a plant’s genome, allowing breeders to see exactly where and how much alien DNA has been integrated. They assist in the process through several specific functions:
1. Identifying Translocation Breakpoints
These techniques act as a visual monitor to identify translocation breakpoints—the precise points where a segment of alien DNA has attached to the crop’s chromosome. By seeing these boundaries, researchers can determine if a desired gene is being transferred in isolation or if it is “dragging” along a large, unwanted flanking segment from the wild relative.
2. Measuring Introgressed Segment Size
GISH allows breeders to quantify the amount of alien chromatin successfully integrated. This is critical because larger segments are more likely to contain multiple genes, some of which may be harmful or reduce the crop’s agricultural quality. For example, in wheat research, GISH is used to detect rye segments carrying disease resistance while helping breeders monitor the reduction of surrounding wild genetic material.
3. Monitoring Recombination Events
Removing linkage drag requires a crossover event (recombination) that breaks the bond between the target gene and the undesirable traits. GISH and FISH enable breeders to:
- Visualize recombinations: Researchers can see which offspring have undergone meiotic crossovers that resulted in a shortening of the alien segment while still retaining the target gene.
- Target specific selections: By identifying plants with the smallest possible alien fragments that still provide the desired trait, breeders can effectively “clean” the crop’s genetic background.
4. High-Precision Tracking with Oligo-FISH
The development of oligo-FISH has significantly increased the precision of this process. Because it uses many short, synthetic probes, it can create a “barcode” for every chromosome, allowing for the detection of very small introgressed segments that were previously impossible to record. This allows for a more surgical approach to removing linkage drag by following chromosomal fragments across generations and backcrosses with high accuracy.
5. Practical Examples
- Onion: In breeding bulb onions for downy mildew resistance, GISH was used to monitor the introgression of the resistance gene from Allium roylei. It allowed researchers to precisely map a lethal factor (undesirable trait) that was closely linked to the resistance gene, enabling them to select recombinants that had lost the lethal segment but kept the resistance.
- Wheat: GISH is used to ensure that genes from wild relatives (like those coding for glutenin subunits) are transferred with minimal flanking segments, ensuring the wild DNA does not interfere with the grain’s quality.
- Lettuce: Historically, aphid resistance was linked to an undesirable plant phenotype; linkage drag was removed by using markers to identify specific crossovers in large populations, a process that is now visually enhanced by these cytogenetic tools.
9. Tell me more about oligo-FISH and its ‘barcoding’ capabilities.
Oligo-FISH (Oligonucleotide Fluorescence In Situ Hybridization) represents a recent advancement in molecular cytogenetics that uses thousands of short, synthetic, single-stranded DNA sequences (oligos) as probes. This technique is considered a paradigm shift from traditional methods that relied on cloned DNA fragments, such as bacterial artificial chromosomes (BACs), which often suffered from low sensitivity and high background noise in complex genomes.
The Concept of Chromosome Barcoding
The most significant innovation of oligo-FISH is its barcoding capability. This allows researchers to uniquely identify every chromosome within a genome in a single experiment.
- Mechanism: Researchers computationally design pools of oligos (typically 40–60 nucleotides long) targeted at specific “spots” or regions along a chromosome.
- Banding Pattern: By using different fluorochromes for various spots, a unique banding pattern—or barcode—is created for each individual chromosome.
- Precision: To ensure a clear signal, “spots” are typically separated by 5–10 Mb, and each spot contains roughly 1,000 distinct oligos. This provides a level of resolution and precision never before achieved in plant cytogenetics.
Applications in Plant Breeding and Research
Oligo-FISH barcoding has transformed the study of complex plant genomes, particularly those that are polyploid (containing multiple sets of chromosomes).
- Potato and Related Species: An oligo-FISH barcode system was developed for the 12 chromosomes of potato (Solanum tuberosum). Remarkably, the same barcode can be used to identify homoeologous chromosomes in related species like tomato and eggplant, revealing how these genomes have rearranged over millions of years of evolution.
- Maize and Rice: Barcoding systems have been established for all 10 chromosomes of maize and all 12 of rice, facilitating the detection of translocations and other structural variations.
- Sugarcane: Because modern sugarcane cultivars are extremely complex polyploids (2n = 100–120), traditional techniques struggle to distinguish their genomic composition. Oligo-barcoding has allowed researchers to finally “decrypt” this architecture, revealing highly variable patterns of genome contribution from ancestral species.
- Switchgrass: Researchers have used barcoding to design probes that distinguish between the highly similar subgenomes (8a and 8b) of this allotetraploid species.
Advantages and Efficiency
- Universal Applicability: In theory, oligo-FISH probes can be designed for any species with a sequenced genome.
- Infinite Resource: Once an oligo pool is synthesized, it can be amplified via PCR, making it a virtually inexhaustible and cost-effective probe resource.
- Non-Model Species: Oligo probes designed for a model species can often be used to study genetically related non-model plants whose genomes have not yet been sequenced.
Limitations
While powerful, the technique has constraints. It requires prior genome sequence information and advanced bioinformatic expertise to design the probes and filter out repetitive elements. In species with massive genomes dominated by repetitive DNA (like wheat), it can be challenging to find enough unique “single-copy” regions to create a punctuated barcoding signal.
10. Why does GISH usually fail in species with very small genomes?
The failure of Genomic In Situ Hybridization (GISH) in species with very small genomes is primarily due to the low quantity and limited distribution of dispersed repetitive DNA sequences.
The sources highlight several specific reasons for this limitation:
1. Dependence on Repetitive DNA
The efficacy of GISH is largely based on the detection of genome-specific dispersed repetitive sequences. Because repetitive DNA constitutes the major source of variation in plant genome sizes, species with small genomes inherently contain fewer of these sequences. When these repeats are sparse, there is not enough labeled probe binding across the chromosomes to create a visible, uniform signal.
2. Preferential Labeling of Specific Regions
In very small genomes, GISH labeling is often restricted to repeat-rich regions, such as:
- (Peri)centromeric regions
- Nucleolar Organizing Regions (NORs)
- Centromeric satellite repeats
This leaves the chromosome arms mostly unlabeled, preventing the “painting” of the entire chromosome complement.
3. The Critical Genome Size Threshold
Researchers have proposed a critical 1C genome size of approximately 0.6 pg. Below this threshold, GISH labeling of entire chromosomes usually fails because the dispersed repeats are not abundant enough to overcome background noise or to provide a contiguous signal.
4. Technical Workarounds
While GISH often fails under standard conditions for small genomes, it is not always impossible. Success has been achieved in some small-genome species, such as Arabidopsis suecica, by utilizing exceptionally high concentrations of labeled probe DNA and significantly longer hybridization times to facilitate the binding of lower-copy sequences.
11. What causes the ‘genome turnover’ that limits GISH in older species?
Genome turnover refers to a collection of biological processes that cause the parental genomes in an allopolyploid species to gradually merge and become indistinguishable over evolutionary time. This phenomenon is the primary reason why Genomic In Situ Hybridization (GISH), which relies on identifying species-specific dispersed repetitive sequences, usually fails in species where the polyploidy event occurred more than 5 million years ago.
According to the sources, the specific causes of this turnover include:
- Intergenomic Homogenization: This is a major driver of turnover where repetitive DNA sequences shared between the subgenomes are “averaged out” or made similar across the entire chromosome complement. As these sequences homogenize, the labeled genomic probes lose their ability to selectively bind to one parental set of chromosomes over the other.
- Retroelement Activity and Amplification: Retrotransposons can become active following a hybridization event (often referred to as “genome shock”). These elements can multiply and spread throughout both subgenomes, effectively diluting the unique repetitive signature of each parent with new, shared sequences.
- Parental DNA Loss: Over time, specific segments of DNA that were unique to one of the ancestral progenitors may be physically lost from the hybrid’s genome. This reduction in parental-specific markers directly diminishes the signal intensity of GISH probes.
- Genome Diploidization: Turnover is often viewed as a near-complete replacement of sequences associated with long-term genome diploidization, the structural and functional process by which a polyploid eventually starts to behave like a diploid species.
- Sequence Replacement vs. Mutation: Research on older Nicotiana species suggests that this failure is caused by the actual replacement of DNA sequences rather than simple point mutations accumulating over time.
In summary, as a polyploid ages, these processes collectively “turn over” the genome’s repetitive fraction, replacing parent-specific elements with a homogenized set of sequences that renders GISH ineffective for genome differentiation.
12. What is the 1C genome size limit for GISH?
The proposed critical 1C genome size limit for GISH is approximately 0.6 pg. Below this threshold, the technique typically fails to label entire chromosomes because there is an insufficient quantity of genome-specific dispersed repetitive sequences to create a visible, uniform signal.
Key Details Regarding the Limit
- Repetitive DNA Dependence: GISH relies primarily on detecting repetitive DNA sequences, which are the main source of variation in plant genome sizes. Species with very small genomes often lack enough of these sequences for the labeled probe to “paint” the entire chromosome complement effectively.
- Regional Labeling: In species that fall below this 0.6 pg threshold, labeling is often restricted to specific, repeat-rich regions such as (peri)centromeric regions, Nucleolar Organizing Regions (NORs), or centromeric satellite repeats, leaving the rest of the chromosome arms uncolored.
- Exceptions and Technical Adjustments: While the 0.6 pg limit is a general rule, successful whole-chromosome labeling in small-genome species can sometimes be achieved by using exceptionally high concentrations of probe DNA and significantly longer hybridization times. This was demonstrated in Arabidopsis suecica, which has a very small genome size.
13. How do researchers use GISH to track alien gene introgression?
Researchers use Genomic In Situ Hybridization (GISH) as a powerful visual “GPS” to track the transfer of genetic material from wild relatives into cultivated crops, a process known as alien gene introgression. This technique provides a direct method for identifying the parental origin of chromatin, measuring the size of integrated segments, and locating precise chromosomal breakpoints.
The Mechanism for Tracking Introgression
To track introgression, GISH utilizes the total genomic DNA of the donor species as a labeled probe.
- Genome Differentiation: Labeled DNA from the alien species is hybridized with the hybrid plant’s chromosomes in the presence of an excess of unlabeled “blocking” DNA from the recipient crop.
- Specific Labeling: The blocking DNA saturates repetitive sequences shared by both species, forcing the labeled probe to bind only to the species-specific dispersed repetitive sequences of the alien chromatin.
- Visual Confirmation: Under a fluorescence microscope, the introgressed alien chromosomes or segments appear in a distinct color against the counterstained background of the recipient crop’s genome.
Applications in Breeding Programs
GISH allows researchers to monitor the successful integration of desirable traits through several critical functions:
- Identifying Translocation Breakpoints: Researchers use GISH to see exactly where an alien segment has attached to a host chromosome, which is vital for understanding how the new genetic material is organized.
- Monitoring Successive Generations: Breeders use GISH to follow introgressed segments through successive backcross generations to ensure the desired trait is stable and remains in the lineage.
- Measuring Introgressed Segment Size: It provides a way to quantify the amount of alien DNA present. This is critical for removing linkage drag, the unwanted wild genetic material that often accompanies beneficial genes.
- Verification of Recombination: GISH allows for the visualization of meiotic recombination events. Researchers can select offspring that have undergone crossovers resulting in the shortest possible alien segments that still retain the target resistance gene.
Practical Examples in Crop Research
- Wheat: GISH has been extensively used to track rust resistance genes introgressed from rye and various wheatgrass relatives like Thinopyrum intermedium.
- Onion: In breeding for downy mildew resistance, GISH was used to select introgression lines where fragments of Allium roylei were successfully integrated into the long arm of Allium cepa chromosome 3.
- Cotton: Researchers use increased stringency GISH and multicolor GISH (McGISH) to identify alien chromosomes and segments from wild species in the complex tetraploid cotton background.
- Sugarbeet and Forage Grasses: These techniques have been applied to map resistance genes and characterize the genomic constitution of hybrids like Festulolium.
Advanced Tracking with McGISH and Oligo-GISH
Modern refinements have significantly increased tracking precision:
- Multicolor GISH (McGISH): This permits the simultaneous discrimination of multiple genomes. In hexaploid wheat, it can distinguish the A, B, and D genomes at once while also identifying any alien segments.
- Oligo-GISH: By using pools of synthetic oligonucleotides as probes, researchers can achieve even higher resolution. This allows for the detection of very small introgressed segments that were previously impossible to record with traditional genomic probes.
14. Explain the concept of genome turnover in allopolyploid species.
Genome turnover is a long-term evolutionary phenomenon in allopolyploid species where the distinct genetic signatures of ancestral parental genomes gradually merge, eventually making them indistinguishable from one another. This process is considered a near-complete replacement of sequences associated with long-term genome diploidization, the structural and functional transition of a polyploid back toward behaving like a diploid species.
According to the sources, the concept of genome turnover involves several key biological mechanisms and evolutionary constraints:
1. Primary Drivers of Turnover
Genome turnover is not caused by simple point mutations but rather by the actual replacement and homogenization of DNA sequences. The main processes include:
- Intergenomic Homogenization: Repetitive DNA sequences shared between subgenomes are “averaged out” or made uniform across the entire chromosome set.
- Retroelement Activity: Retrotransposons may become active and amplify following the “genomic shock” of hybridization, spreading new shared sequences across both subgenomes and diluting parental-specific signatures.
- Parental DNA Loss: Over evolutionary time, specific DNA segments unique to one of the original ancestors are physically lost from the hybrid’s genome.
2. The 5-Million-Year Threshold
The effects of genome turnover are most clearly observed through the limitations they place on cytogenetic techniques like Genomic In Situ Hybridization (GISH).
- GISH relies on identifying species-specific dispersed repetitive sequences to tell parental genomes apart.
- Research on Nicotiana species demonstrates that GISH is generally only effective for distinguishing parental genomes if the polyploidy event occurred less than 5 million years ago.
- In polyploid species older than this threshold, the sequences have “turned over” to such an extent that genomic probes cross-hybridize with all chromosomes, rendering the technique ineffective for differentiation.
3. Progressive Stages of Turnover
By comparing allopolyploids of different ages, researchers can reconstruct the time scale of this turnover:
- Recent Allopolyploids: Display strong, clear GISH patterns that easily distinguish parental chromosomes.
- Intermediate Stages (approx. 1 million years): Show increased cross-hybridization, particularly in specific regions like the subtelomeres.
- Advanced Stages (approx. 4.5 million years and older): Probes may fail to distinguish genome sets entirely, often labeling only highly conserved sites like ribosomal DNA (rDNA).
In summary, genome turnover represents the evolutionary “blending” of two distinct lineages into a single, homogenized genomic unit over millions of years.
15. What is the 5-million-year rule in GISH?
The 5-million-year rule is an empirical observation in molecular cytogenetics stating that Genomic In Situ Hybridization (GISH) is generally only effective for distinguishing parental genomes in natural allopolyploids if the hybridization event occurred less than 5 million years ago.
In polyploid species of this age or older, the ability of genomic probes to selectively bind to one parental set of chromosomes over the other typically fails.
Causes for the Rule
The failure of GISH in older species is attributed to a biological process known as “genome turnover” associated with long-term genome diploidization. This process involves several phenomena that cause parental genomes to merge and become indistinguishable:
- Intergenomic Homogenization: Shared repetitive DNA sequences between subgenomes are “averaged out” or made similar across the entire chromosome complement, reducing the species-specific signature GISH relies upon.
- Retroelement Activity: Following a hybridization event, retrotransposons can become active and spread throughout both subgenomes, effectively diluting the unique repetitive signature of each parent.
- Parental DNA Loss: Unique DNA segments from ancestral progenitors are physically lost over evolutionary time.
- Sequence Replacement: Research suggests this is caused by the actual replacement of DNA sequences rather than just the accumulation of point mutations.
Scientific Evidence from Nicotiana
This rule was established through studies on the evolution of polyploid species in the genus Nicotiana. Researchers observed a clear correlation between the age of the species and GISH effectiveness:
- Recent Allopolyploids: Synthetic hybrids and natural species like N. tabacum (less than 200,000 years old) show strong GISH patterns that clearly distinguish parental chromosomes.
- Intermediate Species: In N. quadrivalvis (approx. 1 million years old), distinction is still possible, but considerable cross-hybridization is observed in subtelomeric regions.
- Older Species: In species like N. nesophila (approx. 4.5 million years old), parental probes are no longer able to distinguish between chromosome sets.
By observing these patterns, researchers use GISH as a tool in GISH-phylogenetics to reconstruct evolutionary time scales and estimate when “genome turnover” occurs in different plant lineages.
16. How can GISH identify alien gene introgressions in wheat?
Genomic In Situ Hybridization (GISH) identifies alien gene introgressions in wheat by acting as a visual “GPS” that differentiates between the host wheat genome and DNA segments introduced from wild relatives. Because wheat is a complex allopolyploid with a long history of hybridization, GISH has become a model technique for tracking the integration of foreign genetic material.
The process works through several key mechanisms and applications:
1. The Methodological Mechanism
- Genome Differentiation: GISH utilizes the total genomic DNA of the alien donor species (such as rye or wheatgrass) as a labeled probe.
- The Role of Blocking DNA: To ensure specificity, a large excess of unlabeled wheat DNA is used as “blocking DNA”. This unlabeled DNA hybridizes to repetitive sequences shared between wheat and the alien species, “blocking” the labeled probe from binding to the wheat chromosomes and forcing it to only highlight the alien-specific sequences.
- Visual Identification: Under a fluorescence microscope, the introgressed alien chromosomes or segments appear in a distinct color, allowing researchers to see their exact physical location within the wheat genome.
2. Identifying Translocations and Breakpoints
GISH is the primary tool for locating translocation breakpoints, where a piece of an alien chromosome has physically attached to a wheat chromosome.
- Rye Introgressions: It is extensively used to detect rye (Secale cereale) chromatin, such as the widely utilized 1BL/1RS translocation, which carries multiple disease resistance genes.
- Wheatgrass Introgressions: Researchers use GISH to track segments from Thinopyrum intermedium (intermediate wheatgrass) and Agropyron elongatum, which provide resistance to Barley Yellow Dwarf Virus (BYDV) and various rusts.
- Precision Mapping: By identifying these breakpoints, breeders can determine if a desired gene is being transferred in isolation or if it is “dragging” along unwanted wild genetic material, known as linkage drag.
3. Advanced Multi-Color and Oligo Techniques
- Multicolor GISH (McGISH): This advanced variation can simultaneously discriminate multiple genomes. In hexaploid wheat, McGISH can distinguish the A, B, and D subgenomes at once while also identifying any alien segments integrated into them.
- Oligo-GISH: The recent shift toward using synthetic oligonucleotide (oligo) probes has further increased precision. Oligo-FISH can create unique “barcodes” for every wheat chromosome, allowing for the detection of very small introgressed segments that were previously invisible to traditional genomic probes.
4. Practical Breeding Applications
- Tracking Resistance Genes: GISH has successfully identified the chromosomal locations of genes for resistance to leaf, stem, and stripe rust, as well as Fusarium head blight.
- Monitoring Generations: Breeders use GISH to follow these alien segments through successive backcross generations to ensure the desired trait is stable and that the amount of alien chromatin is being reduced to the smallest possible size around the target gene.
- Verifying Hybrids: It is used to prove that wide crosses were successful and to distinguish true hybrids from plants that may have developed through other means.
17. How does McGISH distinguish between the three wheat genomes?
Multicolor Genomic In Situ Hybridization (McGISH) distinguishes the three sub-genomes (A, B, and D) of hexaploid wheat (Triticum aestivum) by using a combination of differentially labeled and unlabeled total genomic DNA (gDNA) from ancestral progenitor species.
The process works through the following specific components and stages:
1. Probe Composition
To visualize the distinct genomes simultaneously, researchers utilize specific labeled probes for the A and D genomes:
- A-Genome Probe: Labeled total genomic DNA from the diploid progenitor Triticum urartu is used. This is commonly biotinylated for detection.
- D-Genome Probe: Labeled total genomic DNA from the progenitor Aegilops tauschii (also referred to as Ae. squarrosa) is used. This is typically labeled with digoxigenin.
2. The Role of Blocking DNA
The B-genome is distinguished through a competitive process using blocking DNA. Unlabeled total gDNA from a possible B-genome progenitor, such as Aegilops speltoides, is added to the hybridization mixture in a large excess. This unlabeled DNA binds to the repetitive sequences shared across all three genomes, “blocking” the labeled A and D probes from cross-hybridizing and forcing them to bind only to their target, species-specific sequences.
3. Visualization and Color Differentiation
Through the use of specific fluorochromes (such as fluorescein and rhodamine), each of the three genomes is assigned a unique color for microscopic identification:
- A-genome chromosomes fluoresce yellow.
- B-genome chromosomes appear blue.
- D-genome chromosomes fluoresce red.
4. Research and Breeding Applications
By simultaneously painting all 21 pairs of wheat chromosomes in three distinct colors, McGISH allows researchers to:
- Identify intergenomic translocations where a segment of one wheat genome has moved to another.
- Track alien gene introgressions from wild relatives like rye or wheatgrass, even when integrated as very small segments.
- Monitor genome stability in hybrid lines and synthetic amphiploids, detecting the loss or gain of specific chromosomes.
- Investigate evolutionary relationships between polyploid wheat and its diploid ancestors by analyzing meiotic pairing behavior.
18. Why does genome turnover make GISH ineffective in older species?
Genome turnover makes Genomic In Situ Hybridization (GISH) ineffective in older species because it causes the distinct genetic signatures of parental genomes to merge and become indistinguishable over evolutionary time.
The primary reasons for this loss of effectiveness include:
Reliance on Specific Repetitive Sequences
GISH’s efficacy is largely dependent on the presence of genome-specific dispersed repetitive sequences. These sequences typically evolve faster than functional genes, allowing researchers to differentiate between genomes in relatively recent hybrids. However, genome turnover involves biological processes that gradually eliminate these unique markers.
Mechanisms of Turnover
As an allopolyploid species ages, several phenomena associated with long-term genome diploidization occur:
- Intergenomic Homogenization: Shared repetitive DNA sequences between subgenomes are “averaged out” or made uniform across the entire chromosome complement. This reduces the species-specific signature that genomic probes rely on to distinguish parental sets of chromosomes.
- Retroelement Activity: Following hybridization, retrotransposons can become active and amplify, spreading new, shared sequences across both subgenomes. This process dilutes the unique parental signatures with a homogenized set of elements.
- Parental DNA Loss: Over millions of years, specific DNA segments that were unique to one of the original ancestors may be physically lost from the hybrid’s genome.
- Sequence Replacement: Research suggests that this failure is caused by the actual replacement of DNA sequences rather than just the gradual accumulation of point mutations.
The 5-Million-Year Threshold
In natural allopolyploids, such as those in the genus Nicotiana, GISH is generally only effective if the hybridization event occurred less than 5 million years ago. In species older than this threshold, the genomes have “turned over” to such a degree that parental probes may fail entirely or only label highly conserved sites, such as ribosomal DNA (rDNA), rather than identifying whole chromosome sets.
19. How do blocking DNA ratios vary by species?
Blocking DNA ratios in Genomic In Situ Hybridization (GISH) vary significantly depending on the genetic relatedness of the species involved, with closer relatives requiring a higher concentration of blocking DNA to suppress shared sequences. The ratio of labeled probe to unlabeled blocking DNA typically ranges from 1:10 to 1:200.
General Ratios by Relatedness
- Highly Homologous Species: For hybrids derived from closely related species with high sequence similarity, blocking DNA is typically added at a 20- to 50-fold excess of the probe DNA.
- Optimization Needs: Because shared sequences are often concentrated in repetitive DNA elements, the exact ratio must often be optimized empirically for each specific species combination to ensure the labeled probe only binds to species-specific sequences.
Specific Species Examples
- Cotton (Gossypium): Due to high sequence homology among closely related cotton species, standard conditions often fail. Research found that an optimized 100:1 ratio (blocking DNA to probe) was effective for discriminating genomes in interspecific hybrids like G. hirsutum × G. sturtianum. Other protocols for cotton mention a range from 1:60 to 1:200.
- Wheat and Rye (Triticeae): In common wheat and rye hybrids or triticale, a 1:10 ratio (probe to blocking agent) is frequently used to successfully label rye chromosomes while blocking wheat DNA.
- Wheat and Thinopyrum intermedium: Derivatives of these species have been analyzed using a 1:40 ratio (12.5 ng probe to 500 ng blocking DNA).
- Begonia Hybrids: Clear distinctions in ‘Elatior’-begonia hybrids were achieved using blocking DNA at a concentration 30 times higher than the probe DNA.
- Brassica Species: In Brassica napus, a 10-fold excess of B. oleracea blocking DNA successfully distinguished the parental genomes when B. rapa was the probe. Interestingly, the reciprocal combination failed; unlabeled B. rapa DNA could not block shared sequences even at a 100-fold excess over the B. oleracea probe.
Situations Requiring No Blocking DNA
Blocking DNA is not always necessary if the parental species are sufficiently divergent.
- Diverged Genomes: In cases like Phalaenopsis orchids or certain Silene species, GISH can efficiently identify parental chromosome sets without any blocking DNA because their repetitive elements have diverged significantly over evolutionary time.
- Outgroup Blocking: Sometimes, instead of parental DNA, an excess of unrelated DNA, such as sheared salmon sperm DNA (used at a 50-fold excess) or herring sperm DNA, is used to block non-specific binding.
20. What is genome turnover?
Genome turnover is a collection of evolutionary biological processes that cause the ancestral parental genomes in an allopolyploid species to gradually merge and become indistinguishable over time,. This phenomenon is a primary reason why Genomic In Situ Hybridization (GISH), which relies on identifying species-specific dispersed repetitive sequences, eventually fails to differentiate between sub-genomes in older species,.
According to the sources, the concept of genome turnover involves several key mechanisms and evolutionary patterns:
1. Primary Drivers of Turnover
Genome turnover is not simply the result of point mutations accumulating over time, but rather the actual replacement and homogenization of DNA sequences. The main processes include:
- Intergenomic Homogenization: Repetitive DNA sequences shared between the sub-genomes are “averaged out” or made uniform across the entire chromosome complement,.
- Retroelement Activity: Retrotransposons can become active following the “genome shock” of a hybridization event, multiplying and spreading throughout both sub-genomes,. This dilutes the unique repetitive signature of each parent with new, shared sequences.
- Parental DNA Loss: Over evolutionary time, specific segments of DNA unique to one of the ancestral progenitors are physically lost from the hybrid’s genome,.
2. The 5-Million-Year Threshold
The effects of genome turnover are most clearly observed through studies of natural allopolyploids, particularly in the genus Nicotiana,.
- Effective Limit: GISH is generally only effective for distinguishing parental genomes if the polyploidy event occurred less than 5 million years ago,.
- Turnover Stages: By comparing species of different ages, researchers found that while recent allopolyploids show strong, clear GISH patterns, species approximately 4.5 million years or older often fail to show any genomic distinction,.
- GISH-Phylogenetics: Because of this predictable decline in signal, researchers use GISH to estimate the relative evolutionary age and reconstruct the time scales of polyploidization,.
3. Relation to Genome Diploidization
Genome turnover is considered a near-complete replacement of sequences associated with long-term genome diploidization,. This is the structural and functional transition of a polyploid species back toward behaving like a diploid species,. As the sub-genomes diploidize, they become so homogenized that genomic probes can no longer selectively bind to one set of parental chromosomes over the other,.
21. What are GISH and FISH’s limits?
The limitations of Fluorescence In Situ Hybridization (FISH) and Genomic In Situ Hybridization (GISH) primarily involve technical constraints related to genetic relatedness, the physical size of the target genome, and the resolution of the imaging.
Limitations of GISH
- Genetic Relatedness and Cross-Hybridization: The most significant challenge is the high degree of sequence homology between closely related species. If parental genomes are too similar, the labeled probe binds non-specifically to all chromosomes, requiring high stringency conditions and precisely optimized ratios of blocking DNA that can be difficult to achieve.
- The “5-Million-Year Rule”: GISH is generally ineffective for polyploid species where the hybridization event occurred more than 5 million years ago. Over time, processes like “genome turnover,” intergenomic homogenization, and parental DNA loss merge the ancestral genomes until they are no longer distinct enough to be differentiated by genomic probes.
- Genome Size Threshold: There is a critical 1C genome size limit of approximately 0.6 pg. Below this threshold, GISH typically fails to label entire chromosomes because there are insufficient dispersed repetitive sequences to create a visible signal, often resulting in labeling only at the centromeres.
- Technical Intensity: The method is described as slow, labor-intensive, and expensive compared to molecular markers. Furthermore, weak signals in certain regions can lead to the misinterpretation of intact chromosomes as translocations.
Limitations of FISH
- Analytical Resolution: The resolution of FISH is heavily dependent on the condensation of the target chromosomes. On highly condensed metaphase chromosomes, the resolution is limited to 5–10 Mb. While pachytene chromosomes (100 kb) and DNA fibers (1–2 kb) offer higher resolution, they are more difficult to prepare and analyze.
- Probe Size Constraints: Standard FISH struggles to detect very small DNA sequences; the reliable lower limit is typically 1–3 kb. Probes smaller than 1 kb are often inconsistent, have high background noise, and yield low detection frequencies in cells.
- Repetitive DNA in Large Genomes: In species with massive genomes (like wheat), large-insert probes such as BACs often contain extensive repetitive elements that cause non-specific cross-hybridization across the entire genome, masking the signal of the target unique sequence.
- Commercial and Operational Hurdles: Many probes are not commercially available, requiring research labs to develop “home-brew” probes, which is labor-intensive. The technique also requires specialized expertise and expensive equipment, such as confocal microscopes for 3D analysis.
Shared Limitations and Challenges
- Sample Quality: Results are highly sensitive to the quality of chromosome spreads; dense cytoplasm can block probes, and over-condensation can lead to overlapping, unreadable signals.
- Need for Prior Information: Newer, high-precision methods like oligo-FISH require an existing sequenced and well-assembled reference genome to design the synthetic probes, which limits their use in non-model tropical crops.
- Diagnostic Risks: While highly sensitive, both techniques carry a risk of false positive or negative results, particularly in medical prenatal diagnostics, meaning clinical decisions should never be based on these results alone.
- Rate-Limiting Data Analysis: Manual counting of signals remains the standard for many applications, creating a significant post-experimental bottleneck in high-throughput research.
22. Why does GISH fail after 5 million years?
Genomic In Situ Hybridization (GISH) fails in allopolyploid species older than 5 million years due to a biological phenomenon known as “genome turnover” associated with long-term genome diploidization. This process causes the ancestral parental genomes to gradually merge and become indistinguishable at the molecular level, stripping away the unique signatures that GISH relies on for differentiation.
The sources identify several specific biological mechanisms that drive this failure:
- Intergenomic Homogenization: Repetitive DNA sequences shared between subgenomes are “averaged out” or made uniform across the entire chromosome complement. As these sequences become identical across all chromosomes, the labeled genomic probes lose their ability to selectively bind to one parental set over the other.
- Retroelement Activity: Following the “genomic shock” of a hybridization event, retrotransposons often become active and amplify. These elements spread throughout both subgenomes, effectively diluting the unique repetitive signature of each parent with a new, shared set of sequences.
- Actual Sequence Replacement: Research on the genus Nicotiana suggests that GISH failure is caused by the physical replacement of DNA sequences associated with long-term diploidization, rather than the simple accumulation of point mutations.
- Parental DNA Loss: Over evolutionary time, specific segments of DNA that were unique to one of the ancestral progenitors are physically lost from the hybrid’s genome, reducing the number of targets available for the probe.
The “5-million-year rule” was established by observing these stages in natural Nicotiana allopolyploids of different ages:
Older species (> 4.5 million years): Parental probes fail to distinguish chromosome sets entirely, often only labeling highly conserved sites like ribosomal DNA (rDNA).
Recent species (< 200,000 years): Display strong, clear GISH patterns that easily distinguish parental chromosomes.
Intermediate species (~1 million years): Still show distinction, but exhibit significant cross-hybridization, particularly in the subtelomeric regions.
References
Basit, A., & Lim, K. B. (2024). Systematic approach of polyploidy as an evolutionary genetic and genomic phenomenon in horticultural crops. In Plant Science (Vol. 348). Elsevier Ireland Ltd. https://doi.org/10.1016/j.plantsci.2024.112236
Basit, A., & Lim, K. B. (2025). Recent approaches towards characterization, genetic, and genomic perspectives of genus Lilium. In Genetic Resources and Crop Evolution (Vol. 72, Number 1, pp. 1–28). Springer Nature. https://doi.org/10.1007/s10722-024-01969-6
Chester, M., Leitch, A. R., Soltis, P. S., & Soltis, D. E. (2010). Review of the application of modern cytogenetic methods (FISH/GISH) to the study of reticulation (Polyploidy/Hybridisation). In Genes (Vol. 1, Number 2, pp. 166–192). https://doi.org/10.3390/genes1020166
Cui, C., Shu, W., & Li, P. (2016). Fluorescence in situ hybridization: Cell-based genetic diagnostic and research applications. In Frontiers in Cell and Developmental Biology (Vol. 4, Number SEP). Frontiers Media S.A. https://doi.org/10.3389/fcell.2016.00089
Guan, B., Wang, K., Zhou, B. L., Guo, W. Z., & Zhang, T. Z. (2008). Establishment of a multi-color genomic in situ hybridization technique to simultaneously discriminate the three interspecific hybrid genomes in Gossypium. Journal of Integrative Plant Biology, 50(3), 345–351. https://doi.org/10.1111/j.1744-7909.2007.00623.x
Harun, A., Liu, H., Song, S., Asghar, S., Wen, X., Fang, Z., & Chen, C. (2023). Oligonucleotide Fluorescence In Situ Hybridization: An Efficient Chromosome Painting Method in Plants. In Plants (Vol. 12, Number 15). Multidisciplinary Digital Publishing Institute (MDPI). https://doi.org/10.3390/plants12152816
Huber, D., Voith von Voithenberg, L., & Kaigala, G. V. (2018). Fluorescence in situ hybridization (FISH): History, limitations and what to expect from micro-scale FISH? In Micro and Nano Engineering (Vol. 1, pp. 15–24). Elsevier B.V. https://doi.org/10.1016/j.mne.2018.10.006
Jamja, T., Akhtar, S. F., Bora, S., Tabing, R., Boruah, P., Saatu, M., Uike, A. R., & Rao, E. G. (2024). Molecular and cytogenetic techniques (FISH and GISH) for rapid hybrid confirmation to hasten the breeding pace of ornamental geophytes. Journal of Applied and Natural Science, 16(2), 768–776. https://doi.org/10.31018/jans.v16i2.5544
Jensen, E. (2014). Technical review: In situ hybridization. Anatomical Record, 297(8), 1349–1353. https://doi.org/10.1002/ar.22944
Jiang, J. (2019). Fluorescence in situ hybridization in plants: recent developments and future applications. Chromosome Research, 27(3), 153–165. https://doi.org/10.1007/s10577-019-09607-z
Jiang, J., & Gill, B. S. (2006). Current status and the future of fluorescence in situ hybridization (FISH) in plant genome research. In Genome (Vol. 49, Number 9, pp. 1057–1068). https://doi.org/10.1139/G06-076
Khrustaleva, L., Mardini, M., Kudryavtseva, N., Alizhanova, R., Romanov, D., Sokolov, P., & Monakhos, G. (2019). The power of genomic in situ hybridization (GISH) in interspecific breeding of bulb onion (allium cepa l.) resistant to downy mildew (peronospora destructor [berk.] casp.). Plants, 8(2). https://doi.org/10.3390/plants8020036
Markova, M., Michu, E., Vyskot, B., Janousek, B., & Zluvova, J. (2007). An interspecific hybrid as a tool to study phylogenetic relationships in plants using the GISH technique. Chromosome Research, 15(8), 1051–1059. https://doi.org/10.1007/s10577-007-1180-8
Mekonen, A. A., & Ali, A. (2023). A Review on Principles of FISH and GISH and its Role in Cytogenetic Study (Vol. 1, Number 4).
Ohmido, N., Fukui, K., & Kinoshita, T. (2010). Recent advances in rice genome and chromosome structure research by fluorescence in situ hybridization (FISH). In Proceedings of the Japan Academy Series B: Physical and Biological Sciences (Vol. 86, Number 2, pp. 103–106). https://doi.org/10.2183/pjab.86.103
Patussi, S., Vasconcelos, S., Balvedi Poersch, L., Rafaela, A., & Christina, A. (2013). Genomic in situ Hybridization in Triticeae: A Methodological Approach. In Plant Breeding from Laboratories to Fields. InTech. https://doi.org/10.5772/52928
Raina, S., & Rani, V. (2001). GISH technology in plant genome research. Methods in Cell Science, 23, 83–104.
Ramzan, F., Younis, A., & Lim, K. B. (2017). Application of Genomic in Situ Hybridization in Horticultural Science. In International Journal of Genomics (Vol. 2017). Hindawi Limited. https://doi.org/10.1155/2017/7561909
Ratan, Z. A., Zaman, S. Bin, Mehta, V., Haidere, M. F., Runa, N. J., & Akter, N. (2017). Application of Fluorescence In Situ Hybridization (FISH) Technique for the Detection of Genetic Aberration in Medical Science. Cureus. https://doi.org/10.7759/cureus.1325
Stakelienė, V., & Pašakinskienė, I. (2022). Fluorescent in situ hybridisation in plants: new ways to link DNA sequence resources and chromosome loci – a review. Zemdirbyste, 109(2), 171–178. https://doi.org/10.13080/z-a.2022.109.022
Tang, S., Li, Z., Jia, X., & Larkin, P. (2000). Genomic in situ hybridization (GISH) analyses of Thinopyrum intermedium, its partial amphiploid Zhong 5, and disease-resistant derivatives in wheat. Theor Appl Genet, 100, 344–352.
Volpi, E. V., & Bridger, J. M. (2008). FISH glossary: An overview of the fluorescence in situ hybridization technique. In BioTechniques (Vol. 45, Number 4, pp. 385–409). https://doi.org/10.2144/000112811
Younis, A., Ramzan, F., Hwang, Y. J., & Lim, K. B. (2015). FISH and GISH: molecular cytogenetic tools and their applications in ornamental plants. In Plant Cell Reports (Vol. 34, Number 9, pp. 1477–1488). Springer Verlag. https://doi.org/10.1007/s00299-015-1828-3






