1. Introduction
A complex interplay between ancient biological processes and modern human intervention defines the strategic landscape of crop evolution. As we navigate this “golden era” of plant genomics—characterized by the synergy of technological breakthroughs in massively parallel sequencing and transformative conceptual advances—we have reached a pivotal moment in our ability to decipher the blueprints of life. These breakthroughs have transitioned our understanding of plant genomes from static maps to dynamic ecosystems, providing the essential insights required to address global food security. By integrating primary genomic drivers, such as whole-genome doubling (WGD) and transposable element dynamics, we can now map the wild-to-crop transition with unprecedented accuracy.

What is Polyploidy?
In the study of plant genetics, polyploidy is defined as the condition of possessing more than two complete genomes (sets of chromosomes) within a single cell. While humans are typically diploid (possessing two sets), polyploidy is a natural and frequent phenomenon in the plant kingdom, acting as a powerful mechanism for adaptation and the birth of new species.
To navigate this field, botanists use a specific shorthand notation to describe chromosome sets:
• X: The complete basic set of chromosomes (the monoploid number).
• 2n: The total number of chromosomes found in a somatic cell (non-reproductive cell).
• n: The haploid set found in gametes (pollen or ovules).
We are currently in a “golden era” of plant biology. High-throughput sequencing has revealed that polyploidy is not a rare accident but a fundamental driver of evolution. Genomic analyses show that 35% to 70% of all angiosperms (flowering plants) have a polyploid ancestry. In fact, modern research suggests that nearly all flowering plants are “paleopolyploid,” meaning they have undergone at least one round of genome doubling in their ancient history.
Understanding how plants reach this state requires looking at two distinct biological pathways: internal doubling and external merging.
2. The Cyclical Engine of Evolution: Whole-Genome Doubling (WGD)
Whole-genome doubling, or polyploidy, serves as the fundamental architect of flowering plant history. Far from a rare anomaly, WGD is a ubiquitous force in angiosperm diversification, acting as an active mode of speciation that provides the raw genetic material for evolutionary innovation. Genomic analyses confirm that all flowering plants are paleopolyploids, harboring nested remnants of multiple doubling events tracing back to the root of the seed plants.

Evolutionary history shows that whole-genome doubling (WGD) is cyclical. Plants go through repeated rounds of doubling followed by a slow return to a simpler state.
Diploidization and the Genomic Vaccine
New polyploids eventually undergo Diploidization, the process where a polyploid begins to behave cytogenetically like a normal diploid. This involves:
• DNA-level changes: Massive loss of redundant DNA and chromosomal rearrangements.
• Small RNA Intervention: During the merging of genomes, Small RNAs (specifically hc-siRNAs) act as a “genomic vaccine.” These 24-nucleotide guardians prevent Transposable Elements (TEs) from running wild and causing destabilizing mutations in the newly formed polyploid.
• Expression-level changes:
◦ Subfunctionalization: Duplicated genes divide the original work.
◦ Neofunctionalization: One copy evolves an entirely new function.
◦ The Gene Balance Hypothesis: Genes that have many protein-protein partners (highly connected in biological networks) are more likely to be retained in duplicate to maintain proper “dosage” balance.
These internal changes have a profound impact on how new species are formed in the wild.
The history of plant evolution follows a “wash, rinse, repeat” cycle of polyploidy operating through three distinct strategic pathways:
• Autopolyploidy (The “Self” Doubling)
Autopolyploidy occurs when genome duplication happens within a single species. Instead of combining with another species, the plant doubles its own existing genetic blueprint.
Mechanism Breakdown: Meiotic Failure
The primary engine of autopolyploidy is meiotic failure. During the formation of gametes (micro- and megasporogenesis), a process called meiotic nuclear restitution may occur. This results in Unreduced Gametes (2n)—pollen or eggs that contain the full somatic chromosome count rather than the usual half. When two of these 2n gametes fuse, the resulting zygote is a tetraploid (4n).
Why it Matters
While autopolyploidy is considered rarer in long-term evolution than allopolyploidy, it is highly significant in agriculture. Autopolyploids often exhibit “gigas” effects—larger cells, thicker leaves, and bigger flowers or fruits.
• Triploid Sugar Beets: These possess larger roots and higher sugar yields per unit area.
• Seedless Watermelons: Created by crossing a tetraploid with a diploid, the resulting triploid is sterile, producing only rudimentary, soft white structures instead of hard seeds.
Artificial Induction
Scientists can intentionally induce autopolyploidy using several methods:
1. Colchicine Treatment: A chemical that inhibits spindle fiber formation, stopping cell division after chromosomes have replicated.
2. Heat Shock: Brief exposure of embryos or zygotes to high temperatures (38–45 °C) to disrupt normal division.
3. Decapitation: Removing the apical meristem of a diploid plant to encourage the growth of polyploid callus tissue.
While internal doubling is powerful, the merger of different species introduces even greater complexity.
• Allopolyploidy (The Hybrid Merger)
Allopolyploidy is the result of hybridization between two different species (interspecific) or genera (intergeneric), followed by a doubling of the chromosomes. This creates a plant with two or more distinct genomes.
The “Triploid Bridge” Explained
A common way allopolyploids form in nature is through a two-step “bridge” process involving an intermediate hybrid:
1. Step One: A reduced gamete (1n) from one species fuses with an unreduced gamete (2n) from another, creating a 3n zygote (a triploid hybrid).
2. Step Two: While triploids are often sterile, this 3n individual can occasionally produce gametes that, when fused with a 1n gamete from a parent species, lead to a stable, fertile tetraploid (4n) state.
Morinaga’s Triangle (The Brassica Triangle)
One of the most famous examples of allopolyploidy is the relationship between common vegetable and oilseed crops, illustrated by the “Brassica Triangle.”
| Hybrid Species (Allopolyploid) | Parent Species A | Parent Species B | Somatic Chromosomes (2n) |
|---|---|---|---|
| B. juncea (Indian Mustard) | B. nigra (n=8) | B. campestris (B. rapa) (n=10) | 2n = 36 |
| B. napus (Oilseed Rape) | B. oleracea (n=9) | B. campestris (B. rapa) (n=10) | 2n = 38 |
| B. carinata (Abyssinian Cabbage) | B. nigra (n=8) | B. oleracea (n=9) | 2n = 34 |
These pathways offer different genetic advantages, which become clear when comparing them side-by-side.
• Segmental Allopolyploidy:
A distinct third type (Stebbins 1947) involving the merger of incompletely distinct genomes, which can lead to complex chromosome pairing (both bivalents and multivalents).
Autopolyploidy vs. Allopolyploidy
The evolutionary success of these two states often depends on how their chromosomes behave during reproduction and the level of genetic diversity they maintain.
| Feature | Autopolyploidy | Allopolyploidy |
|---|---|---|
| Genome Source | Single species (self-duplication) | Two+ different species (merger) |
| Meiotic Pairing | Multivalents: Identical chromosomes struggle to pair in groups. | Bivalents: Homologous chromosomes from the same parent pair neatly. |
| Heterozygosity | Increased, but limited to one species’ alleles. | High: Combines diverse alleles from two different ancestors. |
| Representative Crops | Bread wheat, cotton, tobacco, and upland cotton. | Bivalents: Homologous chromosomes from the same parent pair are neatly. |
The genomic responses to these events reshape the species across two distinct temporal phases:
| Phase | DNA-Level Responses | Expression-Level Responses |
|---|---|---|
| Short-Term | Homoeologous exchange; mutational loss of duplicates; activation of transposable elements (TEs). | Duplicate gene expression bias; subfunctionalization (partitioning ancestral expression domains). |
| Long-Term | Structural rearrangements; chromosome reduction; massive genome downsizing (diploidization). | Neofunctionalization (developing novel expression domains); stable network interactions. |
Central to this process is the Gene Balance Hypothesis. This framework explains Biased Fractionation, the strikingly non-random retention of genes following WGD. Because biological networks require precise stoichiometry during protein complex assembly, genes encoding highly connected proteins (e.g., those in upstream pathways or DNA repair) are retained to maintain balance. Conversely, genes adjacent to repressive TEs are often deemed “expendable” and lost. As Wendel et al. (2016) noted, “much like ancient palimpsests, sequenced genomes metaphorically reveal… the reused manuscript pages from previous authors,” containing the scriptio inferior of their polyploid past.
This structural foundation provides the stage for repetitive elements to further expand the genomic landscape.
Polyploidy as an Evolutionary Engine
Polyploidy is a primary “driving force for plant speciation” because it creates an immediate Reproductive Barrier. Because a tetraploid (4n) crossing with its diploid (2n) parent produces sterile triploids (3n), the new polyploid is instantly isolated from its ancestors.
Student Insight: The Power of Redundancy Why keep extra DNA? Genetic redundancy acts as a “buffer.” In a diploid, a mutation in a vital gene might be fatal. In a polyploid, extra gene copies allow one version to mutate and experiment with new traits (innovation) while the other copy continues to perform the essential life function.
Primary Advantages:
1. Heterosis (Hybrid Vigor): Combining different genomes often results in increased vigor, better growth, and higher yields.
2. Genetic Redundancy: Provides a mutation buffer, allowing for evolutionary “experimentation” without risk.
3. Biochemical Flexibility: Polyploids produce multiple versions of enzymes from both parents, potentially allowing them to thrive in stressful or fluctuating environments, such as extreme cold or dry regions.
Reaching the end of our theoretical journey brings us to the tools you need for practical application.
3. Biased Fractionation: The Strategic Legacy of Gene Retention and Loss
Biased fractionation—the non-random retention of genes from one ancestral subgenome over another—is a powerful predictor for gene “expendability” and functional constraint. In allopolyploids, the subgenome with a higher density of TEs often suffers a “repressive effect” on expression, making its genes more prone to loss. This is not a transient state; in Gossypium (cotton), biased fractionation remains evident even 60 million years after the initial WGD event.
The following table distinguishes the fate of genes based on the “Gene Balance Hypothesis,” which posits that dosage-sensitive genes are subject to stronger constraints to maintain stoichiometry.
| Feature | Genes Restored to “Single Copy Status” | Genes “Retained in Duplicate” (Paralogs) |
|---|---|---|
| Stoichiometry & Dosage | High dosage sensitivity (Gene Balance Hypothesis). | Lower dosage sensitivity; tolerant of imbalance. |
| Network Position | Upstream in pathways; high connectivity. | Downstream in pathways; low connectivity. |
| Functional Category | Housekeeping, DNA repair/replication, chloroplast-related. | Regulatory functions, signal transduction, protein complexes. |
| Expression Profile | Broader expression domains; higher levels. | Tissue-specific or specialized expression. |
| Selective Constraint | Extremely high; fundamental for viability. | Prone to neofunctionalization or subfunctionalization. |
| Breeding Utility | Yield stability and fundamental survival traits. | Primary source of novel phenotypic variation and adaptation. |
The Strategic Directive: Knowing which genes are constrained by network connectivity allows breeders to target “Retained in Duplicate” paralogs for trait discovery. In maize, these paralogs are 50% more likely to be associated with functional phenotypic variation than single-copy genes, providing a high-resolution filter for genomic selection.
4. The Genomic Mosaic: Transposable Elements and Size Variation
Understanding genome size variation beyond simple gene counts is a strategic imperative. While genic content remains remarkably constant across species—the 60 Mbp genome of Genlisea aurea contains a similar repertoire to the >150 Gbp genome of Paris japonica—the physical scale of the genome is driven by Transposable Elements (TEs).

TEs act as “agents of rapid adaptation,” as their insertion near regulatory sequences can trigger saltational phenotypic changes. However, their proliferation often represents a “failure of the genomic vaccine.” Small RNA silencing mechanisms (the vaccine) may fail during reproductive stages or genomic mergers, leading to massive TE bursts.
The following table illustrates the genomic complexity of major crops, highlighting the relationship between ploidy history and repeat content:
| Species | Genome Size (Mbp) | Genome Multiples (Ploidy) | % Repeats/Transposons |
|---|---|---|---|
| Asian Rice (Oryza sativa) | ~389 | 32x | 35% |
| Maize (Zea mays) | ~2,500 | 64x | 85% |
| Bread Wheat (Triticum aestivum) | ~17,000 | 96x | 76.6% |
| Soybean (Glycine max) | ~1,100 | 48x | 42% |
| Upland Cotton (G. hirsutum) | ~2,400 | 144x | 67.2% |
| Tomato (S. lycopersicum) | ~900 | 36x | 63.3% |
These repetitive elements are the primary targets of epigenetic remodeling, influencing how humans have historically selected for specific crop traits.
In our genomic ecosystem, the DNA is a mosaic of different inhabitants. We can categorize them into two primary guilds, each playing a different role in the “breath” of the genome.
| Feature | Genic Content (The Founders) | Transposable Elements (The Hitchhikers) |
|---|---|---|
| Identity | The stable blueprints and “permanent residents.” | The “dark matter” or genomic hitchhikers. |
| Stability | Highly conserved; the “scriptio inferior” of life. | Highly dynamic; prone to “genomic explosions.” |
| Function | Encodes essential proteins and trait regulation. | Drives saltational variation and size expansion. |
| Impact on Size | Minimal; gene counts remain remarkably consistent. | Massive; the primary driver of genomic “bloating.” |
The balance between these stable founders and the prolific hitchhikers is maintained through a relentless, episodic process of doubling and purging.
5. The Genetic Architecture of Domestication and Improvement
The wild-to-crop transition is defined by the “domestication syndrome,” a suite of traits—reduced shattering, determinate growth, and increased seed size—that facilitated human reliance on plants.
Strategic analysis reveals distinct evolutionary paths for these transitions:
• Single vs. Multiple Origins: Crops like maize, einkorn wheat, and sunflower trace to single domestication events. In contrast, paddy rice (indica and japonica), common bean, and squash exhibit multiple independent origins. These provide “replicated” natural experiments to study if selection finds the same genetic “paths” to a phenotype.
• The Tempo of Evolution: While the Fertile Crescent transition appeared slow due to seed restocking from the wild, mathematical modeling and Meso-American evidence suggest domestication can occur rapidly—within a few hundred generations—under strong selection.
• Functional Contrast: A vital distinction exists between the genetic drivers of the two phases. Domestication genes typically involve regulatory changes or amino acid substitutions. In contrast, improvement genes (selected post-domestication) are significantly more likely to involve loss-of-function alleles, such as premature stop codons or splice-site defects (e.g., the transition to six-rowed barley).
This complexity is managed by molecular guardians that ensure genomic stability amid such rapid changes.
6. Molecular Guardians: Small RNAs as Regulatory Mediators
Small RNAs are strategic mediators that maintain genomic stability after WGD events and during TE expansion, functioning as a surveillance system to silence invasive elements.
The three major classes of small RNAs are:
1. MicroRNAs (miRNAs): Derived from MIRNA genes and processed by DCL1, these 21-22 nt molecules manage post-transcriptional control of targets, often governing development and stress responses.
2. Secondary siRNAs (phasiRNAs/tasiRNAs): Processed by DCL4/5 into phased configurations. A primary example is tasiARF in TAS3, which acts in trans to regulate target mRNAs.
3. Heterochromatic siRNAs (hc-siRNAs): The 24-nucleotide “guardians of the genome,” derived from Pol IV/V transcripts. They provide multigenerational protection against TEs via RNA-directed DNA methylation (RdDM).
| sRNA Class | Precursor / Dicer Pathway | Primary Function | Evolutionary Impact |
|---|---|---|---|
| miRNAs | Derived from MIRNA genes; processed by DCL1. | Post-transcriptional silencing; regulates development/stress. | High retention post-WGD; dosage-sensitive control of duplicates. |
| phasiRNAs | Derived from mRNAs; requires DCL4/5 and RDR6. | Functions as trans-acting siRNAs (tasiRNAs) against mRNAs. | Targets TEs during reproduction; slower to pseudogenize than coding genes. |
| hc-siRNAs | Transcribed by Pol IV/V; processed by DCL3. | Directs RNA-directed DNA methylation (RdDM). | Multi-generational TE defense; stabilizes newly merged genomes. |
Following WGD, MIRNA and PHAS genes often have a longer “half-life” than protein-coding genes. Their functional relevance is tied to small hairpin structures rather than long coding sequences; thus, they have fewer positions where a mutation is catastrophic, allowing them to persist and regulate duplicated gene dosages over long evolutionary timescales.
7. Future Frontiers: Pangenomes and Global Food Security
Agricultural strategy is shifting from “reference genomes” to “pangenomes.” A single reference sequence fails to capture the “presence-absence variation” and structural diversity found within a species. Projects like the 3,000 Rice Genomes Project show that massive proportions of genetic variation are unmappable to a single reference.
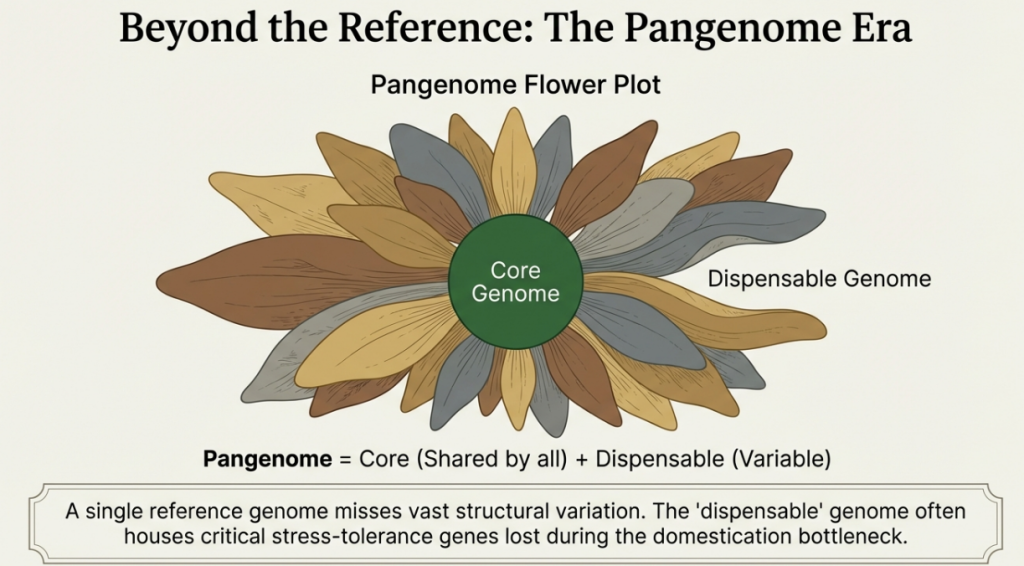
Capturing this diversity is essential for:
• Utilizing Germplasm Banks: Institutions like IRRI and CIMMYT harbor “untapped reservoirs” in wild relatives adapted to varied environments.
• Genomic Archaeology: Projects like Seeds of Discovery (CIMMYT) use “genomic archaeology” to identify traits for nutrient efficiency and stress tolerance.
Addressing the “9.7 billion people question” requires unravelling these genomic ecosystems. Strategic integration of these data into genomic selection breeding programs is the prerequisite for the next Green Revolution.

The Strategic Imperative for Pangenomic Reference Sets
Single reference genomes are “snapshots frozen in time” that fail to capture the “dispensable” genome—the presence-absence variation (PAV) and lineage-specific TE activity that often harbor the most critical adaptation traits.
• Pangenomic Scale: The “3000 Rice Genomes Project” identified 18.9 million SNPs, yet a large portion of resequencing data remains unmappable because it does not exist in the single Oryza reference.
• Assembly Challenges: We must invest in third-generation, long-read sequencing (PacBio, Oxford Nanopore) to overcome the “vexing” repetitive architecture of allopolyploids like wheat (~15 Gbp).
• Origins and Archaeology: We must utilize Genomic In Situ Hybridization (GISH) to clarify the subgenomic origins of complex polyploids like peanut and coffee.
• Genotyping Gene Bank Collections: Strategic initiatives like the Seeds of Discovery project must be expanded to tap into the reservoirs of adaptive traits found in domesticated crop relatives.

Implementation: Integrating Paleogenomics into Modern Breeding Programs
To bridge the gap between genomic archaeology and the field, we must operationalize the following workflow:
1. Targeting Paralogs: Prioritize neofunctionalized genes for selection, focusing on recent WGD descendants (e.g., maize paralogs) which are statistically more likely to drive phenotypic diversity.
2. Managing Heterosis: Leverage “enzyme multiplicity” and intergenomic heterozygosity. In B. napus, allopolyploid-derived diversity is a direct driver of increased oil production.
3. Small RNA Modulation: Monitor the dosage and stability of MIRNA and PHAS loci post-WGD. Utilize the extended half-life of miRNAs to achieve stable expression targets.
4. Overcoming Hybridization Barriers: Use ploidy manipulation and embryo culture to restore fertility. Success depends on matching genomic ratios; specifically, achieving the required 2:1 maternal-to-paternal endosperm ratio to prevent seed abortion in wide hybrids.


8. Conclusion
The architecture of modern crop genomes is a dynamic product of cyclical polyploidy, transposable element expansion, and small RNA regulation. This complex evolutionary history underlies the phenotypes required for human survival and agricultural progress. Integrating massive “omics” datasets is no longer a scientific pursuit but a strategic necessity. To secure global food systems, we must prioritize the genotyping of gene bank collections and the application of genomic selection, bridging the gap between evolutionary archaeology and sustainable field applications.
Master Glossary of Terminological Insights
• Homoeologous Exchange: The swapping of DNA segments between subgenomes in an allopolyploid, which can be reciprocal or non-reciprocal.
• Endoreduplication: A process where DNA replicates without cell division, often occurring in specific tissues like seeds or leaves to increase gene dosage and metabolic output.
• Aneuploidy: A condition where the chromosome number is not an exact multiple of the basic set (e.g., 2n+1 or 2n-1).
• Transposable Elements (TEs): Mobile DNA “jumping genes” whose proliferation and removal account for the vast majority of variation in plant genome size.
• Segmental Allopolyploidy: A state where the merged genomes are not completely distinct, leading to a mix of bivalent and multivalent chromosome pairing.
Image Summary





Questions/Answers
1. What defines the domestication syndrome in major cereal crops?
The domestication syndrome in major cereal crops refers to a suite of morphological and physiological traits that distinguish these cultivated plants from their wild ancestors. These traits were selected by early humans, either consciously or unconsciously, to make the plants more suitable for agriculture, harvesting, and consumption.

In major cereals like wheat, maize, rice, and barley, the defining characteristics of this syndrome include:
1. Loss of Natural Seed Dispersal (Non-shattering)
The most critical trait of the domestication syndrome in cereals is non-shattering, where the plant retains its seeds on the head (rachis) after ripening instead of dispersing them naturally.
• Innovations: This allows for a single, efficient harvest once the crop is ripe.
• Mechanism: In wild cereals, the rachis is brittle and disarticulates easily to spread seeds; in domesticated forms, a mutation results in a non-brittle rachis.
2. Increased Grain/Seed Size
Domesticated cereals typically feature larger and heavier grains compared to their wild progenitors.
• Impact: This increases the overall yield and nutritional value per plant.
• Archaeological Evidence: While grain size often shows a continuum between wild and domestic gene pools, a general increase is a hallmark of the long-term domestication process.
3. Loss of Seed Dormancy (Free Germination)
Wild seeds often have strong dormancy mechanisms to ensure survival across seasons; domesticated cereals have reduced or absent seed dormancy.
• Agricultural Benefit: This ensures uniform and rapid germination immediately after sowing, which is essential for synchronized crop management.
4. Changes in Plant Architecture and Growth Habit
Domesticated cereals exhibit a more compact growth habit compared to the prostrate or highly branched growth of their wild relatives.
• Apical Dominance: A primary example is the reduction in axillary branching (tillering), which leads to a more erect plant.
• Density: These changes allow crops to be grown at much higher densities in cultivated fields.
5. Threshability and Casing
Selection favored traits that made the grain easier to process for food.
• Free-threshing: In crops like bread wheat, selection led to “naked” grains that easily separate from the glumes (chaff) during threshing.
• Maize: Domesticated maize features naked kernels, whereas its ancestor teosinte has seeds enclosed in a hard, enduring tegument.
6. Alterations in Phenology (Flowering and Ripening)
Domestication often involves shifts in the timing of reproduction to adapt to human-constructed environments or new latitudes.
• Uniformity: Traits include uniform seed ripening and synchronized flowering.
• Environmental Adaptation: Changes in photoperiod sensitivity and vernalization requirements allowed tropical cereals like maize and rice to spread into temperate regions with different day lengths and temperatures.
7. Changes in Secondary Metabolites
Domesticated cereals often show a reduction in bitterness, toxicity, or chemical defenses that characterize their wild relatives. This makes the harvested portions more palatable and safe for human consumption.
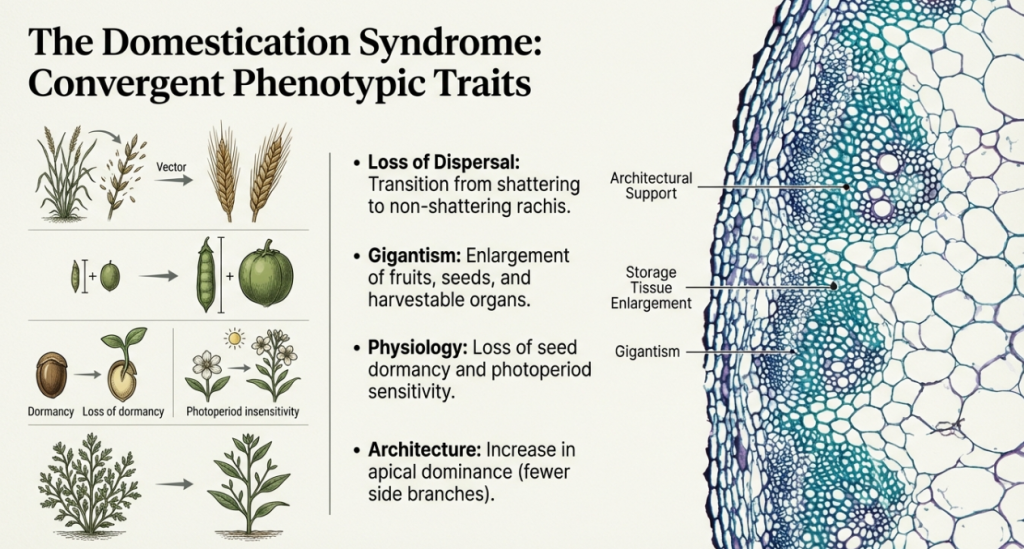
Genetic Basis of the Syndrome
The dramatic morphological changes of the domestication syndrome are often controlled by a relatively small number of genes with major phenotypic effects. Many of these traits are conditioned by recessive, loss-of-function alleles. Furthermore, the genes controlling these characteristics are frequently clustered or linked within the genome, which facilitated the rapid selection and fixation of these traits by early farmers.

2. How does the non-shattering trait impact modern cereal harvesting?
The non-shattering trait, a hallmark of domesticated cereal crops, fundamentally enables modern, efficient, and mechanized harvesting by preventing the natural dispersal of seeds.
The impact of this trait on modern harvesting includes:
• Single, Synchronized Harvest: In wild ancestors, seeds disperse (shatter) as soon as they ripen to ensure survival; however, domesticated non-shattering cereals retain their seeds on the head (rachis). This allows farmers to wait until the entire field is ripe and perform a single harvest rather than multiple manual gatherings.
• Mechanical Efficiency: Non-shattering is a prerequisite for mechanical harvesting. Because the grains are held firmly on the plant, they can be collected by machinery without the seeds falling to the ground and being lost during the process. In crops where this trait is not yet fully fixed, such as certain lentils, modern agriculture must use additional techniques like chemical desiccation to minimize grain loss during mechanical harvest.
• Grain Quality and Yield: The trait prevents the “shattering” that characterizes weedy relatives, which otherwise degrades the quality of the harvested grain and reduces overall yield. Without this trait, the efficiency of modern large-scale agriculture would be impossible, as illustrated by cases where high-shattering weedy populations forced farmers to abandon cultivation entirely.
• Post-Harvest Processing: By retaining the seeds on the plant axis (rachis), the non-shattering trait allows for cleaner collection, which is often paired with other domestication traits like free-threshing, allowing the grain to be easily separated from the protective husks after mechanized collection.
In summary, the transition from a brittle rachis in wild cereals to a non-brittle rachis in domesticated varieties is the critical innovation that facilitates modern, high-intensity grain collection.
3. Explain the role of polyploidy in the evolution of crops.
Polyploidy, or whole-genome doubling (WGD), is the condition of possessing more than two complete sets of chromosomes in a cell. It is far more prevalent in the evolutionary history of plants than previously recognized; in fact, all flowering plants are paleopolyploids, meaning their genomes derive from one or more ancient episodes of genome doubling followed by a process of returning to a diploid state. Polyploidy has played a transformative role in crop evolution by providing the genetic material for diversification, adaptation, and the selection of agronomically favorable traits.
Enrichment in Domesticated Crops
Polyploidization is significantly enriched in crops compared to their wild relatives. Approximately 30% of cultivated crops are neo-polyploids, possessing multiple chromosome sets that are still independent and recognizable. This prevalence suggests that polyploid genomes offer unique advantages that made them prime targets for early human selection.
Key Roles in Crop Evolution
• Enlargement and Vigor (Gigantism): Polyploidy often leads to larger cells and organs, which results in the increased seed and fruit size favored during domestication. For example, induced polyploidy in kiwi has been used to dramatically increase fruit size and alter shape.
• Improved Stress Tolerance: Polyploids frequently exhibit enhanced tolerance to abiotic and biotic stresses, such as drought, salt, and cold, allowing them to thrive in environments that challenge their diploid ancestors. In Hordeum maritimum (sea barley), autotetraploids adapt better to drought-prone mountains than diploids due to a higher number of differentially expressed stress genes.
• Genetic Redundancy and Innovation: The extra sets of genes in a polyploid genome provide a cushion of genetic redundancy. While one copy of a gene maintains its original function, redundant copies are free to mutate and evolve new functions (neo-functionalization) or partition ancestral functions into distinct roles (sub-functionalization).
• Reproductive Isolation: Polyploidization provides a mechanism for rapid sympatric speciation. New polyploids are often reproductively isolated from their diploid ancestors, allowing for the quick fixation of desirable traits by preventing gene flow from wild populations.
• Production of Seedless Varieties: Polyploidy facilitates the development of sterile triploid cultivars (3x), such as those found in commercial bananas and watermelons, which are prized for being seedless.
Molecular and Genomic Dynamics
• Sub-genome Dominance: In allopolyploids (derived from different species), one sub-genome often becomes dominant, experiencing less gene loss and exhibiting higher gene expression than the others. In maize, the dominant sub-genome explains more trait variations than the recessive one.
• Biased Fractionation: Following WGD, the genome undergoes fractionation, where duplicated genes are lost over time. This loss is often non-random; genes restored to a single copy are frequently those involved in essential “housekeeping” or DNA repair functions.
• Asymmetric Selection: Different sub-genomes can contribute to different types of traits. In allotetraploid cotton, the A sub-genome was primarily selected for fiber improvement, while the D sub-genome contributed genes for stress tolerance.
Examples in Major Crops
• Wheat: Hexaploid bread wheat (Triticum aestivum) carries three sub-genomes (AABBDD). The Q locus, a major domestication gene controlling free-threshing, requires the coordinated regulation of paralogs from these three sub-genomes—a complexity not found in diploid wheat.
• Brassica: A whole-genome triplication (WGT) event in the Brassiceae tribe drove the extreme morphological diversification seen in crops like cabbage, broccoli, and cauliflower.
• Sugarcane: Modern sugarcane cultivars are highly complex allopolyploids and aneuploids, often resulting from backcrossing Saccharum officinarum with the wild S. spontaneum to transfer disease resistance.
• Potato: A polyploid series (ranging from 2x to 6x) has allowed cultivated and wild potatoes to adapt to a vast range of altitudes and climates across the Americas.
4. How were secondary metabolites like flavor and toxicity selected during domestication?
Selection for secondary metabolites, such as those governing flavor and toxicity, is considered the most universal domestication-related trait across hundreds of crop species. During the transition from wild plants to cultivated crops, early humans exerted intense pressure to reduce chemical defenses and increase the palatability of the parts they intended to consume.
Conscious Selection for Palatability
Domesticated crops were primarily selected through conscious human choice to enhance taste and nutritional quality.
• Direct Selection: Humans targeted the reduction of bitter, unpalatable, or toxic defense compounds directly within the harvested organs (such as fruits, seeds, or tubers).
• Examples of Bitterness Removal: The loss of bitterness was a crucial domestication trait for crops like almonds, cucumbers, melons, and squash. In many of these cases, the transition from a bitter wild form to a non-bitter domesticated form is simply inherited, meaning it is controlled by only one or two genes.
• Toxicity Reduction: Selection led to the significant reduction of harmful substances, such as glycoalkaloids in potatoes, glucosinolates in cabbage, and cyanogenic glycosides in cassava.
Genetic Mechanisms
The shift in chemical profiles typically resulted from specific types of genetic mutations:
• Loss-of-Function Mutations: This is the most common mechanism. Mutations effectively “blocked” biochemical pathways that produced toxic metabolites, making the plant safe for human consumption.
• Regulatory Changes: Many flavor and color traits are controlled by regulatory genes, such as transcription factors, rather than structural enzymes. For example, the waxy (glutinous) phenotype in various grains arose through loss-of-function mutations at the Waxy locus, which altered starch composition to suit cultural preferences for sticky textures.
• Transposable Elements: In some cases, the insertion of “jumping genes” (transposable elements) triggered changes in secondary metabolism, such as the accumulation of anthocyanins in blood oranges or color changes in grapes.
Trade-offs and Unintended Consequences
Selection for flavor and yield often had side effects on the plant’s natural resilience:
• Growth vs. Defense Trade-off: Selecting for increased yield and larger harvested organs often led to a reallocation of metabolic resources away from chemical defenses toward growth.
• Increased Vulnerability: As a result of reduced chemical and morphological defenses, domesticated plants are often less stress-resistant and more susceptible to herbivory and pests than their wild ancestors.
• Induced vs. Constitutive Defense: While direct selection often reduced “constitutive” (always present) defenses, some plants retained “inducible” defenses that only activate when the plant is under attack, which carries a lower yield penalty.
Selection During Diversification
While the initial removal of toxicity occurred during early domestication (Stage 1), the diversification of flavors and colors often happened later as crops spread to new cultures. Traits like aroma in rice or specific pigment variations in fruits were selected and maintained based on cultural preferences and specific local uses. For instance, preference for sticky grains led to the parallel evolution of waxy mutants in unrelated species like rice, sorghum, and foxtail millet across East Asia.
5. Does the domestication syndrome differ between annual and perennial crops?
Yes, the domestication syndrome differs significantly between annual and perennial crops in terms of the number of traits affected, the rate of evolutionary change, and the specific physiological modifications that occurred.
1. Number and Complexity of Traits
Perennial crops, particularly fruit and nut trees, typically exhibit significantly fewer domestication syndrome traits than annual seed crops. While annuals like cereals often undergo a wholesale transformation across a suite of traits (such as non-shattering, loss of dormancy, and compact growth), many perennials show fewer phenotypic shifts. This difference is partly due to the fact that perennials have longer generation times, meaning they have undergone fewer sexual generations since the onset of domestication to accrue and fix these changes.
2. Rate of Domestication
The rate at which a species transitions from its wild form to a domesticated state is significantly slower in perennials/trees than in annuals. Annual crops are often estimated to have been domesticated over a shorter period, whereas the process for perennials has been more protracted. However, once certain technical innovations like scion grafting were developed, the domestication of many tree crops accelerated in later “waves”.
3. Specific Trait Divergence
The traits emphasized in the domestication syndrome vary based on the life cycle:
• Seed Dispersal and Dormancy: These are critical traits for annual seed crops (e.g., non-shattering grains), but they are often irrelevant or not selected for in perennial fruit and nut trees. For instance, no changes in seed shattering are observed in crops from humid regions like Near Oceania, where perennials are more prevalent.
• Yield and Organ Size: While both groups show increased yield in the edible portions, perennials focus more on fruit morphology, size, and color diversification.
• Secondary Metabolites: Changes in secondary metabolites (reducing toxicity or bitterness and improving flavor) are considered the most universal domestication trait across all species, affecting both annuals and perennials.
4. Reproductive and Genetic Mechanisms
Annual and perennial crops differ fundamentally in their reproductive evolution:
• Reproductive Strategy: Domestication of perennials is heavily associated with a shift toward vegetative (clonal) propagation (e.g., cuttings or grafting), which allows for the immediate fixation and maintenance of desirable phenotypes without the genetic reshuffling of sexual reproduction.
• Ploidy Levels: Changes in ploidy (whole-genome doubling) are much more common in perennial crops than in annuals. Approximately 78% of the crops in a large dataset that showed ploidy changes under domestication were perennials. In contrast, ploidy changes in sexually reproducing annual crops like bread wheat are considered exceptional.
• Genetic Bottlenecks: Perennials typically experienced less severe genetic bottlenecks during domestication compared to annuals. This is attributed to their longer generation times and a history of repeated introgression with wild relatives.
5. Summary Table of Key Differences
| Feature | Annual Crops (e.g., Cereals) | Perennial Crops (e.g., Fruit Trees) |
|---|---|---|
| Number of Syndrome Traits | Many (High complexity) | Fewer (Lower complexity) |
| Pristine Traits | Non-shattering, loss of dormancy | Edible organ size, flavor, color |
| Domestication Rate | Relatively faster | Slower/Protracted |
| Ploidy Changes | Rare/Exceptional | Common (78% of cases) |
| Propagation Method | Mostly sexual | Often shifted to vegetative/clonal |
| Genetic Diversity Loss | High (Severe bottlenecks) | Moderate (Less severe) |
6. How do non-shattering traits differ across various cereal species?
Non-shattering—the loss of natural seed dispersal—is a foundational trait of the domestication syndrome in cereals, though the genetic mechanisms and selective pressures that produced it vary significantly across species. While many cereals share orthologous (homologous) genes for this trait, others evolved non-shattering through unique genetic pathways or maintained shattering due to specific harvesting cultures.
The following table and sections summarize how non-shattering traits differ across various cereal species:
Comparison of Non-Shattering Traits by Species
| Cereal Species | Primary Genetic Mechanism | Morphological Feature Affected | Evolutionary Pattern |
|---|---|---|---|
| Sorghum | SH1 gene | Rachis (head) | Three unique haplotypes suggest at least three separate domestication origins. |
| Rice (Asian) | Sh4 and qSH1 | Abscission zone at the base of the grain | A single amino acid substitution in Sh4 is the primary driver. |
| Barley | Btr1 and Btr2 | Rachis (spike axis) | Diphyletic origin; two different mutations in western vs. eastern accessions. |
| Wheat | Br1, Br2, Br3 (Btr orthologs) and Q locus | Rachis toughness and glume tenacity | Stepwise recruitment; Btr fixed in emmer, Q and Tg (tenacious glumes) later in bread wheat. |
| Maize | ZmSh1 and tga1 | Rachis (cob) and glumes | tga1 specifically allows for “naked kernels” by reducing the seed casing. |
Key Differences in Mechanisms
• Convergent Evolution at the Shattering1 Locus: A striking example of parallel evolution is found in the Shattering1 (Sh1) gene. Orthologs of this gene control shattering in sorghum, rice, and maize. This suggests that while these species were domesticated independently, human selection often targeted the same genetic pathways to achieve seed retention.
• Brittle Rachis vs. Free-Threshing (Wheat/Barley): In the Triticeae (wheat and barley), the primary trait is a non-brittle rachis, which prevents the spike from falling apart at maturity. However, wheat also possesses a second layer of selection for free-threshing (controlled by the Q and Tg loci), which allows the grain to be easily separated from the protective glumes (chaff) during processing.
• Glume Architecture (Maize): Maize is unique because its non-shattering is tied to the teosinte glume architecture 1 (tga1) gene. In teosinte, seeds are enclosed in a hard, stony casing that shatters; in domesticated maize, a single amino acid substitution in tga1 results in “naked kernels” that remain firmly attached to the cob.
• Cultural Exceptions (African Rice): Harvesting techniques can influence whether non-shattering is selected. Unlike Asian rice, African rice (Oryza glaberrima) was traditionally harvested by swinging a basket to catch seeds as they fell. Because this method favored the natural dispersal of grain, the non-shattering trait was never fixed in many African rice populations.
De-Domestication and Weed Evolution
Non-shattering is often the first trait lost when a crop becomes feral (the process of de-domestication). Weedy relatives, such as weedy rice in China and the US or feral rye in North America, have re-evolved shattering mechanisms. This allows them to disperse their seeds naturally in agricultural fields, making them highly effective mimics that are difficult to control during commercial harvesting.
7. How does pan-genomics identify alleles lost in modern breeding?
Pan-genomics identifies alleles lost in modern breeding by providing a comprehensive species-wide genomic space that captures the diversity across cultivars, landraces, and wild relatives, rather than relying on a single reference genome. Because a single reference genome represents only a fraction of a species’ genetic variability, pan-genomics allows researchers to uncover alleles that were inadvertently removed during intense selection for specific agricultural traits.
The identification process typically involves the following mechanisms:
1. Mapping the Accessory Genome
Pan-genomics categorizes genes into two main groups:
• Core Genome: Genes present in all individuals of a species.
• Accessory (Dispensable) Genome: Variable genes found only in a subset of the species, further divided into “shell genes” (present in many accessions) and “cloud genes” (found in only one or two taxa). Lost alleles associated with traits like disease resistance or abiotic stress tolerance are often located within this accessory genome, particularly in the gene pools of landraces and wild ancestors that have not undergone the “bottleneck” of modern industrial breeding.
2. Detecting Large Structural Variations
Modern breeding analysis often focuses only on Single Nucleotide Polymorphisms (SNPs). However, pan-genomics utilizes de novo genome assemblies and graph-based approaches to detect larger, more complex variations that are frequently lost, including:
• Presence-Absence Variations (PAVs): Entire genes or sequences that are missing in modern cultivars but present in ancestors.
• Copy Number Variations (CNVs): Reductions in the number of gene copies that might have occurred during domestication.
• Large Structural Variants (SVs): Chromosomal rearrangements, inversions, and translocations that are often “blind” to short-read resequencing against a single reference.
3. Comparative Evolutionary Scanning
By scanning the pan-genome across the wild-to-cultivated transition, researchers can pinpoint exactly which genomic regions have been depleted of diversity:
• Soybean Example: A pan-genomic study of 1,110 soybean accessions revealed that 1.5% of genes were lost during domestication, specifically those related to defense and salt responses, while genes for flowering time and seed composition were maintained or increased.
• Sorghum and Millet: Pan-genomic analysis of sorghum identified 79 new drought-related genes that were missing from the standard reference genome. Similarly, pearl millet pan-genomics identified over 400,000 structural variants and new genes for heat tolerance.
• Cotton: Research indicates that the cotton pan-genome specifically “retrieves” lost sequences and genes that were selected against during the evolution of elite allopolyploid varieties.
4. Overcoming Ascertainment Bias
Traditional genotyping arrays were designed primarily from sequences of domesticated accessions, creating a bias against detecting wild alleles. Pan-genomics eliminates this by using reference-free alignments or graph-based models that allow any variant discovered in any individual to be included as a node, making it possible to visualize and track alleles that are unique to wild populations.
8. How did selection reduce glycoalkaloids in potatoes and cassava?
The reduction of glycoalkaloids in potatoes and cyanogenic glycosides in cassava was achieved through conscious human selection specifically targeting the palatability and safety of the harvested organs. This selection for reduced toxicity is considered the most universal domestication-related trait across hundreds of crop species.
The process involved several key factors:
• Selection for Palatability: Early humans targeted the reduction of bitter, unpalatable, or toxic defense compounds directly within the organs they intended to consume, such as potato tubers and cassava roots.
• Simple Inheritance: In many root and tuber crops, the transition from a toxic wild form to a safe domesticated form is simply inherited, meaning it is often controlled by only one or two genes with major phenotypic effects.
• Genetic Mechanisms: The reduction in these metabolites typically resulted from loss-of-function mutations. These genetic changes (such as nonsense mutations or premature truncations) effectively “blocked” the biochemical pathways responsible for synthesizing the harmful substances.
• Targeted Organ Selection: Selection pressure was exerted specifically on the harvested portions of the plant rather than the vegetative tissues. This allowed the plants to reduce toxins where humans ate them while sometimes retaining chemical defenses in other parts of the plant to protect against herbivory.
• Inducible vs. Constitutive Defense: While “constitutive” (always present) defenses were often reduced during selection, some crops retained “inducible” defenses that only activate when the plant is under attack, which carries a lower yield penalty for the farmer.
• Modern Mutagenesis: In modern potato breeding, researchers have also used induced mutagenesis (such as gamma-ray irradiation) followed by high-performance screening to isolate specific mutant lines with significantly lower glycoalkaloid content for industrial use.
While both crops underwent selection for reduced toxicity, the sources note that cassava still contains cyanogenic glycosides that require specific processing techniques to ensure they are safe for consumption.
9. Are there yield trade-offs when plants lose chemical defenses?
Yes, there are significant yield trade-offs associated with the loss or maintenance of chemical defenses in plants. This relationship is often described as the growth vs. defense trade-off, where a plant must allocate limited metabolic resources between either growing (producing biomass/yield) or protecting itself from pests and diseases.
The impact of these trade-offs manifests in several ways:
1. Resource Allocation Shifts
The central theory is that reallocating metabolic resources from defense to growth allows domesticated crops to achieve much higher yields than their wild ancestors.
• Domestication selection: Early humans intentionally selected for the reduction of bitter, toxic, or unpalatable secondary metabolites (like glycoalkaloids in potatoes or glucosinolates in cabbage) to improve flavor and safety.
• Yield gains: By removing the “metabolic cost” of producing these expensive defensive compounds, the plant can direct that energy toward larger seeds, fruits, or tubers.
2. The “Yield Penalty” for Resistance
Conversely, when modern breeders attempt to reintroduce resistance traits from wild relatives into crops, it often results in a yield penalty.
• Constitutive resistance: Defenses that are always present (constitutive) are particularly costly because they shift resources toward defense mechanisms even when no threat is present, which can reduce overall productivity in ideal, stress-free environments.
• Linkage drag: Yield trade-offs also occur during breeding because resistance genes from wild relatives are often linked to undesirable traits (linkage drag). Carrying these additional genomic segments can inadvertently reduce the yield of the resulting cultivar.
3. Vulnerability and Human Dependency
A major trade-off of losing these chemical defenses is that domesticated plants have reduced fitness under natural conditions.
• Increased susceptibility: Because they have lost natural chemical and morphological defenses, modern crops are often more susceptible to herbivory and pathogens.
• Human intervention: This loss of self-defense makes crops dependent on human management, such as the use of pesticides and controlled environments, to maintain their high yield potential.
4. Mitigation Strategies (Inducible Defenses)
Not all defensive traits carry the same yield penalty.
• Inducible resistance: Some plants utilize defenses that only activate when the plant is under attack. Because these mechanisms are not expressed under ideal conditions, they typically carry a lower yield penalty compared to constitutive defenses.
• Exception examples: Some studies have shown that it is possible to uncouple growth and defense. For instance, a study on domesticated cabbage found reduced glucosinolates with no corresponding trade-off in leaf area (growth), suggesting that the relationship is not always strictly linear.
In summary, while losing chemical defenses allowed for the dramatic yield increases seen in modern agriculture, it created a trade-off of increased vulnerability and a continuing challenge for breeders who must balance high productivity with necessary resilience.
10. How do transcription factors regulate flavor and color mutations?
Transcription factors regulate flavor and color mutations by controlling the expression of genes within metabolic pathways, such as those responsible for pigment or secondary metabolite synthesis. While some mutations in these pathways target structural enzymes directly, many universal domestication and diversification traits—particularly those involving pigmentation—arise from selection on regulatory genes that coordinate the activity of multiple genes.
The sources highlight several specific mechanisms by which transcription factors influence these traits:
1. Regulation of Pigmentation (Anthocyanins)
Transcription factors, particularly from the MYB and bHLH families, are critical regulators of anthocyanin production, which determines the color of fruits and grains.
• Grapes: The transition from red to white berries in cultivated grapes resulted from mutations in the VvMybA gene family. Specifically, a mutation in VvMYBA2 followed by a retrotransposon insertion in the promoter of VvMYBA1 disrupted the regulatory signal for anthocyanin synthesis, leading to non-pigmented fruit.
• Blood Oranges: The deep red color of blood oranges is caused by the cold-induced transcriptional activation of Ruby, a Myb regulatory gene. This activation is triggered by a transposable element (TE) insertion in the gene’s regulatory region.
• Rice Bran Color: The transition from red to white bran in rice is controlled by the Rc locus, which encodes a basic helix-loop-helix (bHLH) transcription factor. A 14-base pair deletion in the protein-coding region of this gene disrupts its function, leading to the loss of red pigment in most domesticated varieties.
2. Regulation of Fruit Ripening and Flavor
Flavor is often linked to the concentration of secondary metabolites and sugars, which can be modulated by transcription factors during the ripening process.
• Tomato Ripening: A WD40 transcription factor (SlWD40) has been identified as a coordinator of fruit ripening in tomatoes. Ripening involves a complex metabolic shift that affects the fruit’s final flavor and aroma profile.
• Secondary Metabolite Shifts: Domestication frequently involves reducing bitter or toxic secondary metabolites for better flavor. In many cases, these shifts are governed by a relatively small number of genes with major effects, often encoding transcriptional regulators that can “block” or “activate” entire defense pathways.
3. Mutational Mechanisms in Regulatory Genes
The regulation of these traits often changes through specific types of mutations in transcription factors:
• Cis-regulatory Mutations: Mutations in the non-coding regions (like promoters) can alter when and where a transcription factor is expressed, thereby changing the plant’s chemical profile without losing the gene’s function entirely.
• Loss-of-Function Mutations: These are the most common type of mutation in crop evolution genes. For example, premature stop codons or frameshifts in transcription factor genes can completely eliminate a specific color or bitter flavor.
• Transposable Element Insertions: Approximately 15% of characterized domestication and diversification genes feature TE insertions that serve as the causative mutation, often by disrupting or enhancing the regulation of transcription factors like tb1 in maize or Ruby in oranges.
11. How did early farmers manage seed germination for perennials?
Early farmers managed the germination of perennials primarily by bypassing sexual reproduction through seeds and instead utilizing vegetative (clonal) propagation. This shift allowed farmers to immediately fix and maintain desirable plant traits that would otherwise be lost or reshuffled through sexual seed production.
The management of perennial crops occurred in two distinct technological waves:
• Initial Wave (Cuttings and Suckers): Starting approximately 6,000 years ago, farmers began using simple vegetative techniques like cuttings or suckers to propagate species such as the olive.
• Second Wave (Scion Grafting): Between 3,000 and 2,000 years ago, the discovery and dissemination of scion grafting (e.g., for carob) allowed for the more sophisticated management and spread of perennial varieties.
Impact on Germination Traits
Because of the reliance on cloning rather than seeds, the “domestication syndrome” for perennials differs from annuals:
• Irrelevance of Seed Dormancy: Traits such as the loss of seed dormancy, which are critical for annual grain crops to ensure uniform germination after sowing, were often not selected for or were irrelevant in perennial fruit and nut trees.
• Fixation of Polyploidy: Vegetative propagation allowed farmers to maintain polyploid perennials (which often have larger fruits) without the decreased reproductive output typically associated with genome duplication in plants that reproduce sexually.
• Fewer Syndrome Traits: Perennials generally exhibit significantly fewer domestication syndrome traits than annuals because they have undergone fewer sexual generations since the onset of their cultivation.
In summary, rather than managing the germination of seeds, early farmers of perennials focused on maintaining superior individual genotypes through physical division and grafting, effectively removing the need for a seed-based germination stage in their agricultural cycles.
12. How do weedy rice and feral rye re-evolve shattering?
The re-evolution of shattering in weedy rice and feral rye occurs through a process known as de-domestication, where the traits accumulated during human selection (like non-shattering) are evolutionarily lost. The mechanisms vary depending on the ancestry of the specific weedy lineage.
Weedy Rice (Oryza sativa f. spontanea)
Weedy rice is a polyphyletic set of lineages, meaning it has evolved multiple times through different pathways. The re-emergence of shattering happens through:
• Hybridization with Wild Relatives (Exoferality): In many cases, such as “strawhull” weedy rice in the United States, the lineage is a hybrid between a cultivated rice variety (indica) and the wild ancestor O. rufipogon. In these instances, the wild relative provides the functional alleles for shattering through gene flow.
• Direct Descent from Cultivars (Endoferality): Some weedy rice populations, like “blackhull” weedy rice in the US and certain populations in China, descended directly from domesticated rice without wild hybridization. In these cases, shattering re-evolves through mutation or epistatic recombination events that restore the dispersal mechanism.
• Intervarietal Hybridization (Exo-endoferality): In regions like Bhutan, weedy rice evolved from hybrids between two different domesticated varieties (indica x japonica). These crosses can create a “burst of variation” that allows for the selection and fixation of shattering phenotypes.
Feral Rye (Secale cereale)
Unlike rice, western North American feral rye is a monophyletic lineage that evolved directly from one or more cereal rye cultivars. Its re-evolution of shattering is defined by:
• Endoferal Pathway: Genetic analysis has failed to detect ancestry from wild perennial rye (S. strictum) or other wild species.
• Genetic Restoration: The transition from the non-shattering cultivated form to the shattering feral form likely occurred via mutation or epistatic recombination.
• Rapid Evolution: This reversal occurred very quickly—approximately 60 generations after the original observations of volunteer populations.
Ecological Context and Survival
The re-evolution of shattering is critical for the survival of these plants as weeds. Shattering allows seeds to disperse naturally in the field before or during the harvest of the main crop. In both species, the shattering trait is often paired with morphological mimicry, making the weeds difficult to distinguish from the crop during hand-weeding, which further protects the re-evolved dispersal units until they can ripen and shatter.
13. How does pan-genomic research overcome bias toward domesticated varieties?
Pan-genomic research overcomes bias toward domesticated varieties by moving beyond the limitations of a single reference genome to capture the full species-wide genomic space, including diversity found in landraces and wild relatives.
Traditional genomic tools often suffer from ascertainment bias because they were designed primarily using sequences from domesticated accessions, which makes them less effective at detecting unique alleles present in wild populations. Pan-genomics addresses this through several key strategies:
1. Comprehensive Inclusion of Diversity
Instead of relying on an “elite” cultivar as the sole representative of a species, pan-genomes are constructed from a diverse panel of genotypes, including wild ancestors, landraces, and ancestral species. For example, the “Super Pan-genome of rice” includes 251 high-quality assemblies covering domesticated Asian and African rice along with their respective wild ancestors.
2. Reducing Reference Bias with Graph-Based Models
Traditional methods align new sequences to a single linear reference genome, often causing divergent reads from wild relatives to be discarded or misaligned. The graph-based approach in pan-genomics overcomes this by:
• Adding variants as nodes: Any discovered variant is added as a node at its specific genomic location.
• Accommodating complex variation: It can represent structural variants, inversions, and large-scale rearrangements that a single reference genome cannot capture.
• Accurate Mapping: Reads are realigned to the entire graph, leading to more accurate mapping and the identification of unique genomic regions.
3. De Novo Assembly and Independent Analysis
Pan-genomic research often employs de novo assembly of multiple individuals rather than just mapping reads back to a single reference.
• Retaining divergent information: This approach allows for the retention of divergent sequences and “novel gene space” that would otherwise be missed.
• Uncovering lost alleles: It enables researchers to identify valuable genes associated with disease resistance or stress tolerance that were inadvertently lost during the intensive bottlenecks of domestication and modern breeding.
4. Overcoming Technical Blind Spots
By providing multiple reference genomes, pan-genomics allows for the systematic study of structural variation (like presence-absence variations and copy number variations) across different genotypes. This provides a much more accurate reflection of genome-wide diversity and helps precisely detect wild-to-crop introgressions.
14. How does pan-genomic research overcome bias toward domesticated varieties?
Pan-genomic research overcomes the historical bias toward domesticated varieties by moving beyond the limitations of a single, often “elite,” reference genome to capture the full species-wide genomic space. Traditional genomic tools often suffer from ascertainment bias because they were designed primarily using sequences from domesticated accessions, which makes them less effective at detecting unique alleles present in wild populations.
Pan-genomics addresses these technical and conceptual biases through the following strategies:
1. Comprehensive Inclusion of Genetic Diversity
Instead of relying on a single cultivar as the representative of a species, pan-genomes are constructed from a diverse panel that includes wild ancestors, landraces, and ancestral species.
• Super Pan-genomes: Researchers are now building “super pan-genomes” that represent entire genera. For instance, the super pan-genome of rice includes high-quality assemblies for 251 accessions covering both Asian and African domesticated rice and their wild relatives.
• Accessory Genome: By including a wide array of genotypes, pan-genomics identifies the accessory (dispensable) genome—genes present only in a subset of the species—which often harbors valuable traits lost during the “bottlenecks” of domestication.
2. Technical Shifts: De Novo Assembly and Graph Models
Pan-genomics utilizes advanced assembly and alignment methods to capture variation that traditional linear references miss:
• Retention of Divergent Information: Using de novo assembly of multiple individuals allows researchers to retain divergent reads containing important variant information that would otherwise be discarded if mapped to a single reference.
• Graph-Based Pan-genomes: This approach represents variants as nodes at their specific genomic locations. This allows the genome to accommodate complex variation, such as large-scale rearrangements, inversions, and translocations, providing a more accurate mapping for wild relatives.
• Reference-Free Alignments: Resources like the Rice Super Pan-genome Database host reference-free whole-genome multiple sequence alignments, eliminating the bias inherent in choosing one variety as the “standard”.
3. Recovery of “Lost” Alleles and Structural Variation
Pan-genomics acts as a tool for genomic archaeology, uncovering genetic material inadvertently removed during intense agricultural selection.
• Presence-Absence Variations (PAVs): Traditional GWAS often focus only on Single Nucleotide Polymorphisms (SNPs). Pan-genomics identifies PAVs—entire sequences or genes present in wild ancestors but missing in modern cultivars.
• Empirical Examples:
◦ Soybean: A study of 1,110 accessions revealed that 1.5% of genes were lost during domestication, specifically those related to defense and salt responses.
◦ Sorghum: Iterative assembly added 174.5 Mb of new sequence (a 20% increase over the reference), including 79 newly identified drought-related genes.
◦ Zea: A pan-Zea genome map identified that approximately 44% of genes were dispensable across the genus, highlighting the vast reservoir of variation outside the primary B73 reference.
4. Overcoming Technical Blind Spots
By providing a species-wide atlas, pan-genomics enables more precise detection of wild-to-crop introgressions. It overcomes the “blindness” of traditional resequencing to large structural variants (SVs), which account for a larger proportion of sequence differences than SNPs in some species, such as tomato. This comprehensive view allows breeders to reintroduce lost alleles for disease resistance, abiotic stress tolerance, and nutritional quality into modern elite lines.
15. How did weedy mimics evolve to evade hand-weeding by farmers?
Weedy mimics evolved to evade hand-weeding through a process known as Vavilovian mimicry, driven by the intense selective pressure of visually-based human weeding over thousands of years.
The Mechanism of Selection
The evolution of crop mimicry is primarily a result of unconscious selection by farmers.
• Visual Screening: During the vegetative growth phase, farmers manually remove plants from their fields that do not look like the intended crop.
• Differential Survival: Weeds that happen to possess mutations making them morphologically similar to the crop—such as having a similar erect growth habit, leaf color, or leaf shape—are more likely to be overlooked and spared.
• Reproductive Advantage: Because these mimics are not removed, they survive to maturity, allowing them to reproduce and disperse their seeds (often through shattering) back into the field for the next season.
Genetic and Morphological Basis
The mimicry is often so effective that the weed and crop are vegetatively indistinguishable, even when they belong to entirely different genera.
• Plant Architecture: In the case of barnyard grass (Echinochloa crus-galli var. oryzicola) mimicking rice, researchers have identified 87 putative plant architecture-related genes that have been under selection to facilitate this mimicry.
• Convergent Traits: While crops were selected for compact growth to facilitate high-density planting, weedy mimics evolved the same compact, non-branching stature to blend in.
• Detection Window: Many mimics, including weedy rice, weedy beet, weedy rye, and semi-wild wheat, are nearly impossible to distinguish from the crop until they reach the flowering stage, by which time they may have already successfully competed for resources or be too late to easily remove.
Examples of Successful Mimics
• Barnyard Grass and Rice: In Japanese rice fields, barnyard grass individuals that most closely resemble cultivated rice are significantly less likely to be hand-weeded. Multivariate analysis of 15 quantitative characters shows that these mimics are not significantly different from rice morphologically during their vegetative phase.
• Weedy Rice: As the same species as cultivated rice (Oryza sativa), weedy rice is a specialized crop mimic with very similar physiology, making it extremely difficult to control through both manual weeding and chemical means.
• Feral Rye: In the western United States, weedy rye became such a successful mimic and competitor in cultivated rye fields that many farmers were forced to abandon efforts to grow the crop for human consumption entirely.
16. How does de-domestication lead to the re-emergence of wild traits?
De-domestication, also known as ferality, is the evolutionary process by which a domesticated plant loses traits accumulated under human selection and re-acquires characteristics that allow it to survive, reproduce, and disperse in the wild or as a weed. This transition typically occurs when crops escape cultivation and are subjected to natural selection in non-managed environments.
The re-emergence of wild traits through de-domestication occurs via several genetic pathways and manifests in specific morphological and physiological changes:
1. Key Wild Traits that Re-emerge
The most defining wild traits that reappear during de-domestication are those that were suppressed by early farmers to facilitate harvesting and consumption:
• Re-evolution of Shattering (Seed Dispersal): This is the most common wild trait to re-emerge. While crops were selected for a non-brittle rachis to keep seeds on the plant for harvest, de-domesticated lineages (like weedy rice and feral rye) re-evolve shattering mechanisms to naturally disperse their seeds into the soil.
• Increased Seed Dormancy: Domesticated annuals were selected for uniform and immediate germination. De-domestication often leads to a return of strong seed dormancy, a “bet-hedging” strategy that allows seeds to survive in the soil across unfavorable seasons until the right conditions for growth occur.
• Defensive Structures: Some plants re-evolve morphological defenses. For example, artichoke thistle in California, derived from domesticated artichoke, re-developed spininess and more deeply dissected leaves.
• Reduced Organ Size: Crops selected for “gigantism” (large seeds or fruits) often revert to smaller, more numerous reproductive organs. Feral rye has evolved smaller seeds and leaves compared to its cultivated ancestors.
• Changes in Life Cycle: Weed beets in Europe have shifted from a biennial habit (typical of the crop) to an annual habit, accompanied by the re-emergence of a woody root.
2. Pathways to De-Domestication
Lineages can follow two primary evolutionary paths to acquire wild traits:
• Endoferality (Direct Descent): The pest lineage descends directly from the crop without crossing with wild relatives. Wild traits re-emerge through spontaneous mutations in domestication genes or epistatic recombination events that restore ancestral functions. Western North American feral rye is a monophyletic endoferal lineage that evolved directly from rye cultivars.
• Exoferality (Hybridization): The crop crosses with a wild relative or a different cultivar, reintroducing functional wild alleles. For instance, “strawhull” weedy rice in the U.S. resulted from hybridization between cultivated rice and the wild ancestor O. rufipogon, which provided the alleles for shattering.
• Exo-endoferality: A specific case (e.g., in Bhutanese weedy rice) where hybrids between different domesticated varieties (like indica and japonica) create a “burst of variation” that natural selection then uses to fix feral phenotypes.
3. Genetic Mechanisms
De-domestication can be driven by a small number of allele changes.
• Restoration of Function: Functional alleles for shattering can be re-introduced through gene flow from wild relatives or through mutations that reverse a crop’s loss-of-function domestication mutation.
• Regulatory Shifts: Changes in flowering time, such as the delayed flowering observed in feral rye, can act as a reproductive isolating mechanism to prevent “maladaptive” gene flow from the nearby crop back into the weedy lineage, allowing the wild traits to fix rapidly.
• Rapid Evolution: De-domestication can be remarkably fast. Feral rye exhibited significant phenotypic divergence from its ancestor in roughly 60 generations.
4. Mimicry as a Survival Strategy
Intriguingly, while re-evolving wild dispersal traits, many feral plants retain certain domesticated traits that make them successful weeds. Weedy rice, rye, and beet often remain morphologically indistinguishable from the crop during their vegetative phase. This crop mimicry protects them from being removed by hand-weeding until they flower and can disperse their re-evolved wild seeds.
References
Abbo, S., Lev-Yadun, S., & Gopher, A. (2012). Plant Domestication and Crop Evolution in the Near East: On Events and Processes. Critical Reviews in Plant Sciences, 31(3), 241–257. https://doi.org/10.1080/07352689.2011.645428
Abbo, S., Pinhasi van-Oss, R., Gopher, A., Saranga, Y., Ofner, I., & Peleg, Z. (2014). Plant domestication versus crop evolution: A conceptual framework for cereals and grain legumes. In Trends in Plant Science (Vol. 19, Number 6, pp. 351–360). Elsevier Ltd. https://doi.org/10.1016/j.tplants.2013.12.002
Alam, O., & Purugganan, M. D. (2024). Domestication and the evolution of crops: variable syndromes, complex genetic architectures, and ecological entanglements. Plant Cell, 36(5), 1227–1241. https://doi.org/10.1093/plcell/koae013
Burger, J. C., Chapman, M. A., & Burke, J. M. (2008). Molecular insights into the evolution of crop plants. In American Journal of Botany (Vol. 95, Number 2, pp. 113–122). https://doi.org/10.3732/ajb.95.2.113
Burke, J. M., Burger, J. C., & Chapman, M. A. (2007). Crop evolution: from genetics to genomics. In Current Opinion in Genetics and Development (Vol. 17, Number 6, pp. 525–532). https://doi.org/10.1016/j.gde.2007.09.003
Butt, H., Shan-e-Ali Zaidi, S., Hassan, N., & Mahfouz, M. (2020). CRISPR-Based Directed Evolution for Crop Improvement. Trends in Biotechnology, 38, 236–240. https://doi.org/10.1016/j.tibtech
Cheeseman, J. M. (2015). The evolution of halophytes, glycophytes and crops, and its implications for food security under saline conditions. New Phytologist, 206(2), 557–570. https://doi.org/10.1111/nph.13217
Dar, J. A., Beigh, Z. A., & Wani, A. A. (2017). Polyploidy: Evolution and crop improvement. In Chromosome Structure and Aberrations (pp. 201–218). Springer India. https://doi.org/10.1007/978-81-322-3673-3_10
Ellstrand, N. C., Heredia, S. M., Leak-Garcia, J. A., Heraty, J. M., Burger, J. C., Yao, L., Nohzadeh-Malakshah, S., & Ridley, C. E. (2010). Crops gone wild: Evolution of weeds and invasives from domesticated ancestors. Evolutionary Applications, 3(5–6), 494–504. https://doi.org/10.1111/j.1752-4571.2010.00140.x
Frary, A., & DOĞANLAR, S. (2003). Comparative Genetics of Crop Plant Domestication and Evolution. Turkish Journal of Agriculture and Forestry, 27(2).
Gao, L., Kantar, M. B., Moxley, D., Ortiz-Barrientos, D., & Rieseberg, L. H. (2023). Crop adaptation to climate change: An evolutionary perspective. In Molecular Plant (Vol. 16, Number 10, pp. 1518–1546). Cell Press. https://doi.org/10.1016/j.molp.2023.07.011
Gui, S., Martinez-Rivas, F. J., Wen, W., Meng, M., Yan, J., Usadel, B., & Fernie, A. R. (2023). Going broad and deep: sequencing-driven insights into plant physiology, evolution, and crop domestication. Plant Journal, 113(3), 446–459. https://doi.org/10.1111/tpj.16070
Haas, M., Schreiber, M., & Mascher, M. (2019). Domestication and crop evolution of wheat and barley: Genes, genomics, and future directions. In Journal of Integrative Plant Biology (Vol. 61, Number 3, pp. 204–225). Blackwell Publishing Ltd. https://doi.org/10.1111/jipb.12737
Meyer, R. S., Duval, A. E., & Jensen, H. R. (2012). Patterns and processes in crop domestication: An historical review and quantitative analysis of 203 global food crops. In New Phytologist (Vol. 196, Number 1, pp. 29–48). https://doi.org/10.1111/j.1469-8137.2012.04253.x
Meyer, R. S., & Purugganan, M. D. (2013). Evolution of crop species: Genetics of domestication and diversification. In Nature Reviews Genetics (Vol. 14, Number 12, pp. 840–852). https://doi.org/10.1038/nrg3605
Naithani, S., Deng, C. H., Sahu, S. K., & Jaiswal, P. (2023). Exploring Pan-Genomes: An Overview of Resources and Tools for Unraveling Structure, Function, and Evolution of Crop Genes and Genomes. In Biomolecules (Vol. 13, Number 9). Multidisciplinary Digital Publishing Institute (MDPI). https://doi.org/10.3390/biom13091403
Olsen, K. M., & Wendel, J. F. (2013). Crop plants as models for understanding plant adaptation and diversification. In Frontiers in Plant Science (Vol. 4, Number AUG). Frontiers Research Foundation. https://doi.org/10.3389/fpls.2013.00290
Pathirana, R. (2021). Mutations in plant evolution, crop domestication and breeding. Tropical Agricultural Research and Extension, 24(3), 124. https://doi.org/10.4038/tare.v24i3.5551
Schreiber, M., Stein, N., & Mascher, M. (2018). Genomic approaches for studying crop evolution. In Genome Biology (Vol. 19, Number 1). BioMed Central Ltd. https://doi.org/10.1186/s13059-018-1528-8
Vigouroux, Y., Barnaud, A., Scarcelli, N., & Thuillet, A. C. (2011). Biodiversity, evolution and adaptation of cultivated crops. Comptes Rendus – Biologies, 334(5–6), 450–457. https://doi.org/10.1016/j.crvi.2011.03.003
Wendel, J. F., Jackson, S. A., Meyers, B. C., & Wing, R. A. (2016). Evolution of plant genome architecture. In Genome Biology (Vol. 17, Number 1). BioMed Central Ltd. https://doi.org/10.1186/s13059-016-0908-1
Zhang, K., Wang, X., & Cheng, F. (2019). Plant Polyploidy: Origin, Evolution, and Its Influence on Crop Domestication. Horticultural Plant Journal, 5(6), 231–239. https://doi.org/10.1016/j.hpj.2019.11.003






