1. Introduction: The Bacterial Arms Race
For billions of years, a microscopic war has raged in every niche of the biosphere. Bacteria and phages—the viruses that prey upon them—are locked in a relentless evolutionary arms race, each side developing increasingly sophisticated stratagems to ensure survival. For the modern biotechnology sector, these bacterial defense systems represent a strategic frontier. By decoding how bacteria outmaneuver their viral counterparts, we have previously unlocked revolutionary tools like CRISPR; today, we are on the cusp of discovering an entirely new mode of biological information transfer.
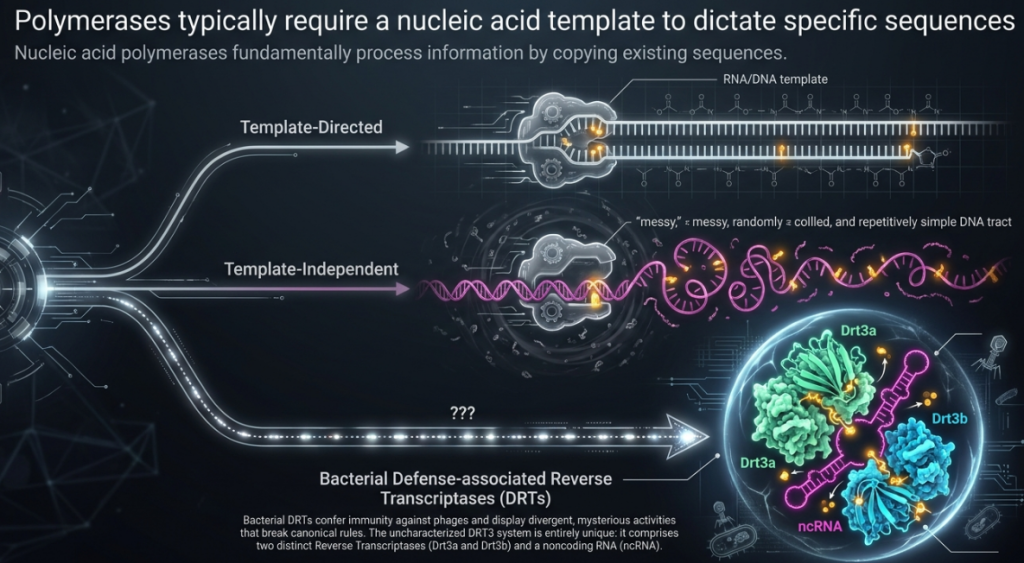
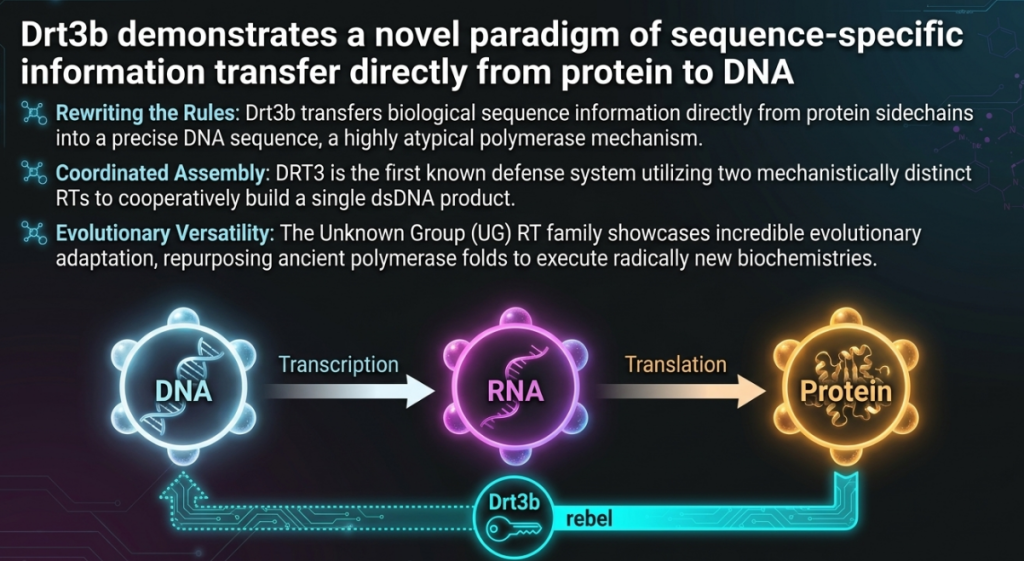
A landmark study by Deng et al., published in Science, has revealed a startling new weapon in this ancient conflict: the DRT3 system. This discovery represents a profound challenge to the “central dogma” of molecular biology—the foundational principle that genetic information flows strictly from DNA to RNA to protein. Specifically, Deng and colleagues demonstrated that the DRT3 system produces defensive DNA through a process of non-canonical information transfer. Instead of copying an existing genetic template, this molecular machine uses the physical shape and chemical properties of a protein to “write” genetic code. This transition from nucleic-acid templating to protein-templated synthesis marks a paradigm shift in our understanding of how life processes and stores information.
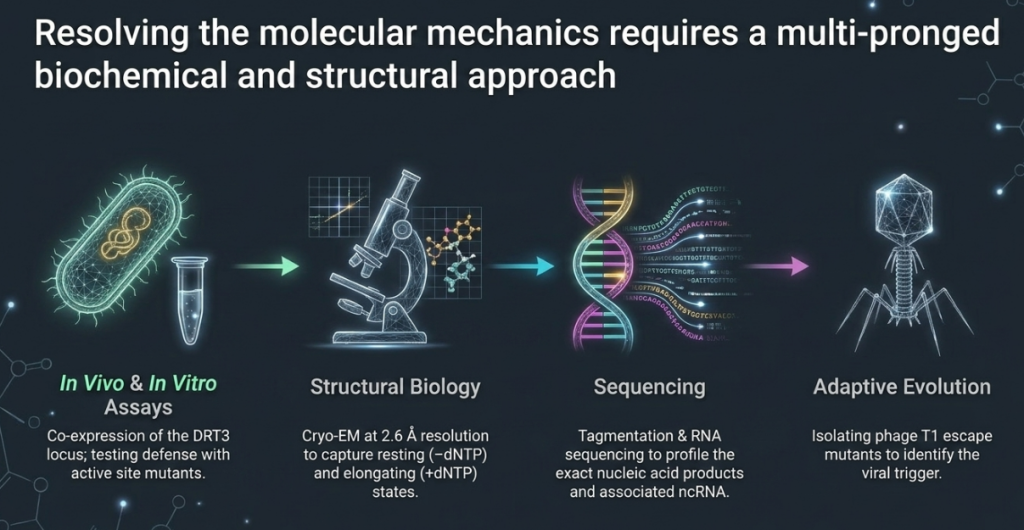
2. Meet the DRT3 Complex: A High-Order Defensive Fortress
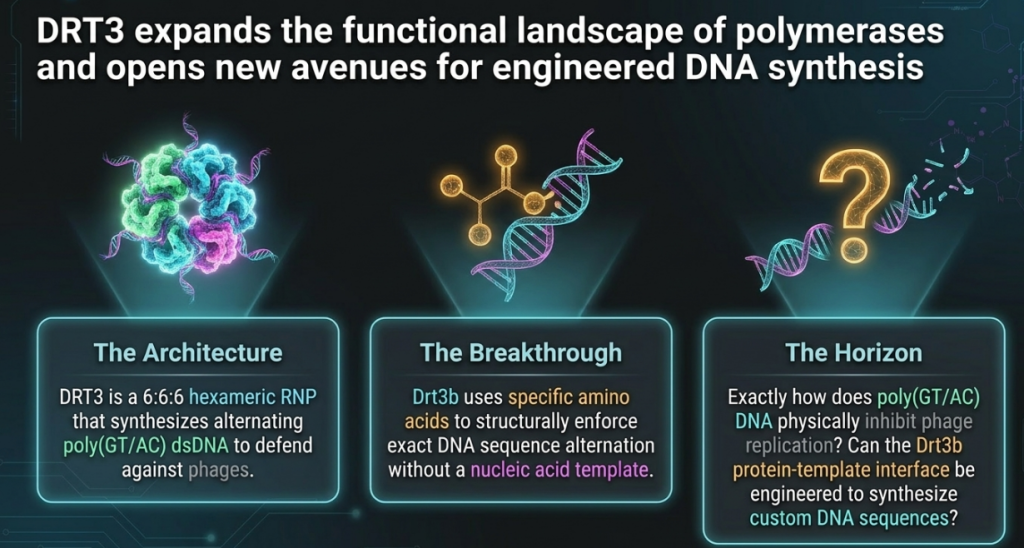
The engine of this breakthrough is a highly structured ribonucleoprotein (RNP) complex. In biological terms, an RNP is a strategic alliance where proteins and RNA operate as a single, coordinated unit. The DRT3 system is not merely a collection of parts; it is a “high-order defensive fortress” characterized by a unique D3-symmetric 6:6:6 stoichiometry. This hexamer of trimers creates a massive, stable molecular machine designed for rapid, large-scale DNA production.
The system relies on three integrated components:
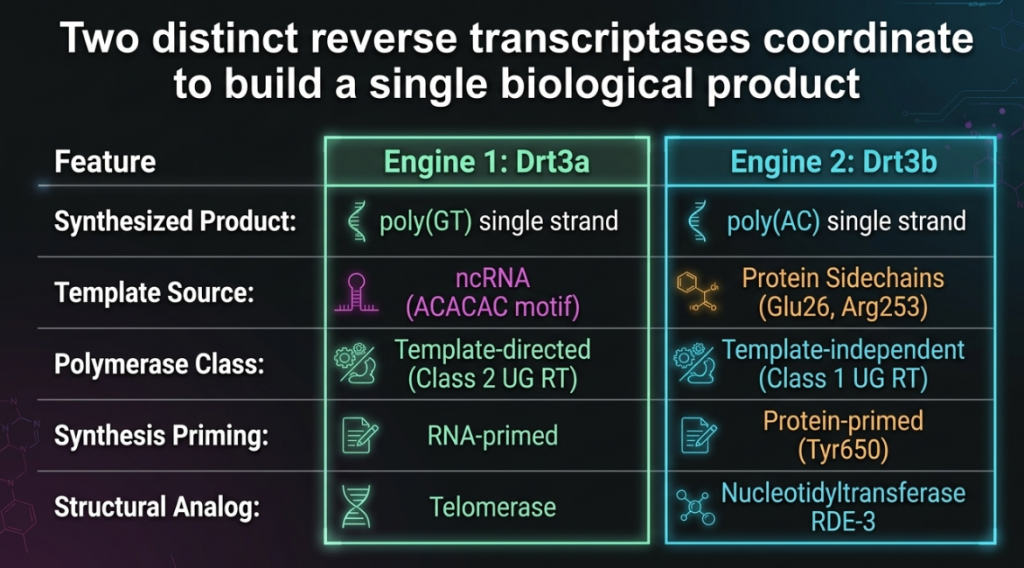
- Drt3a (The RNA-Guided Architect): A class 2 reverse transcriptase that serves as the complex’s “traditionalist.” It follows a genetic blueprint provided by the ncRNA to build one-half of the DNA weapon. It features a unique β-hairpin extension that acts as a mechanical “zipper head” to separate the newly synthesized DNA from its RNA template, facilitating continuous, high-speed synthesis.
- Drt3b (The Template-Independent Innovator): A class 1 reverse transcriptase that breaks the fundamental rules of biology. Drt3b possesses an internal architecture that precludes traditional template binding, instead relying on its own amino acid structure to determine the DNA sequence.
- ncRNA (The Internal Blueprint): A non-coding RNA that serves as both the structural scaffold for the hexameric complex and the specific sequence template for Drt3a.
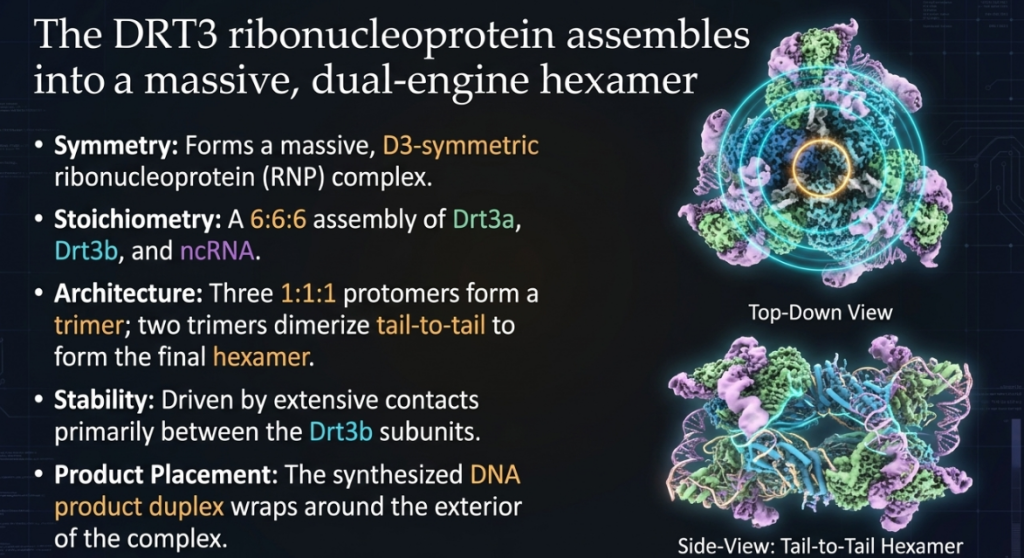
The co-evolution of these components across 20 bacterial phyla suggests that this “coordinated dance” of DNA synthesis is an ancient and robust defense mechanism, honed over eons to create a nearly impenetrable biological barrier.
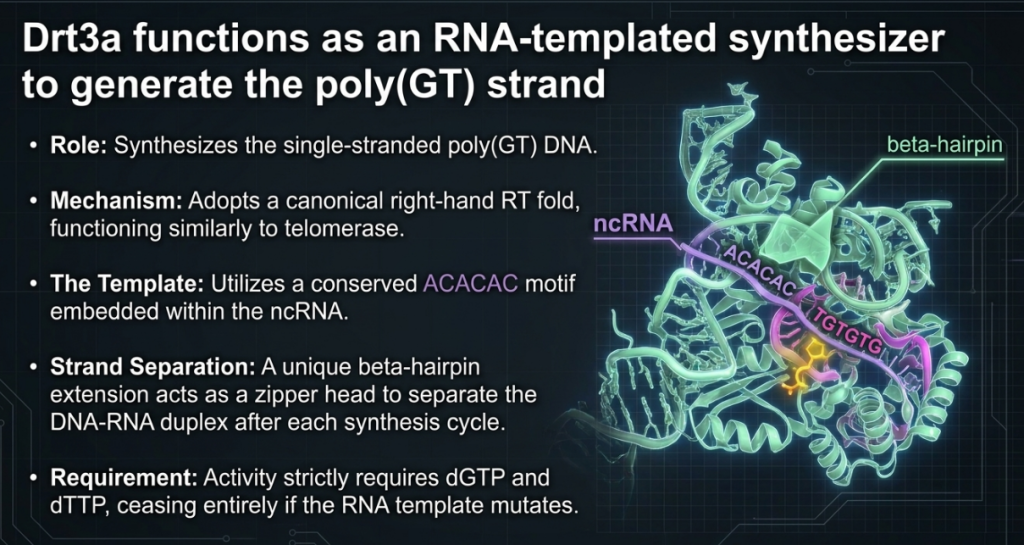

3. Key Findings: How Bacteria “Write” New DNA
The most profound discovery of the Deng et al. study is that the DRT3 system shifts the role of nucleic acid polymerases from “copiers” to “creators.” To use a strategic analogy: while most polymerases function like a photocopier (Drt3a), requiring an external master sheet to produce a copy, Drt3b functions like a biological 3D printer. It does not “read” an external instruction; rather, its own internal architecture and specific “settings” dictate the final output.
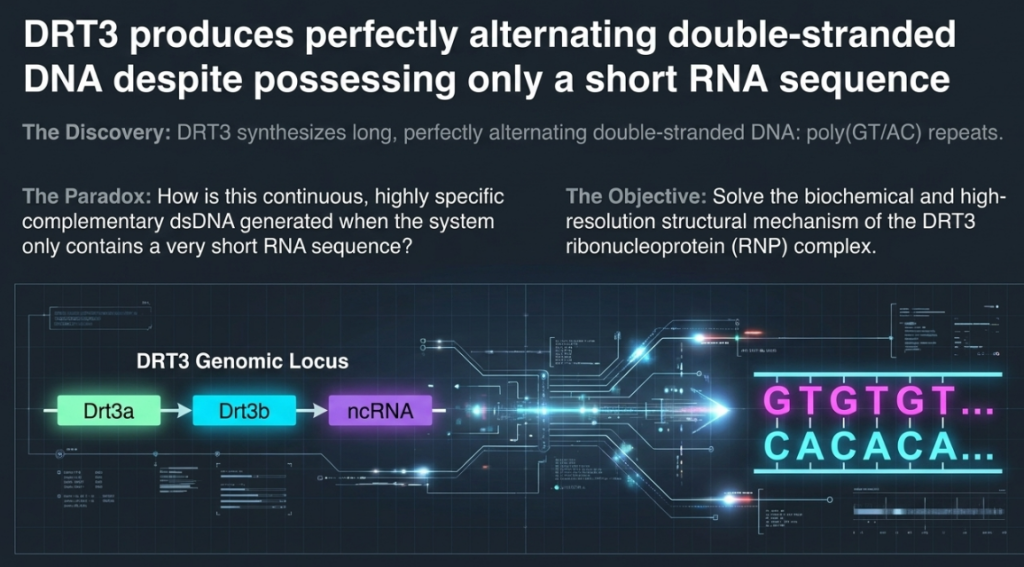
The system executes a perfectly synchronized, two-pronged synthesis that results in a double-stranded DNA “net” of alternating poly(GT/AC) repeats:
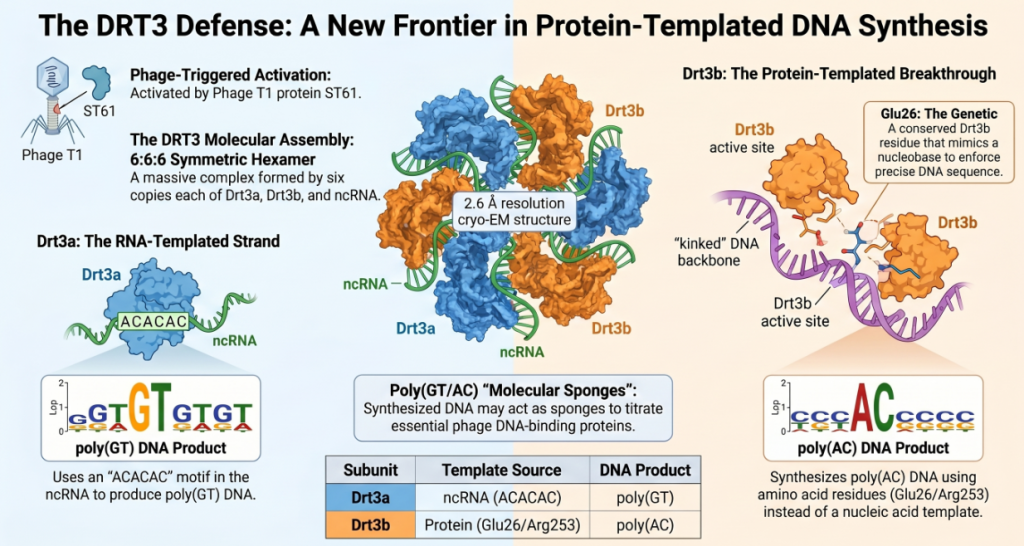
- The GT Strand: Drt3a uses a conserved “ACACAC” motif within the ncRNA to weave a repeating pattern of GT DNA. This is a highly efficient process, mirroring the mechanism used by eukaryotic telomerase.
- The AC Strand (The Rule-Breaker): Drt3b synthesizes the complementary “AC” strand in the total absence of a nucleic acid template. It initiates this process through Tyr650-mediated priming, which acts as the critical “start switch.”
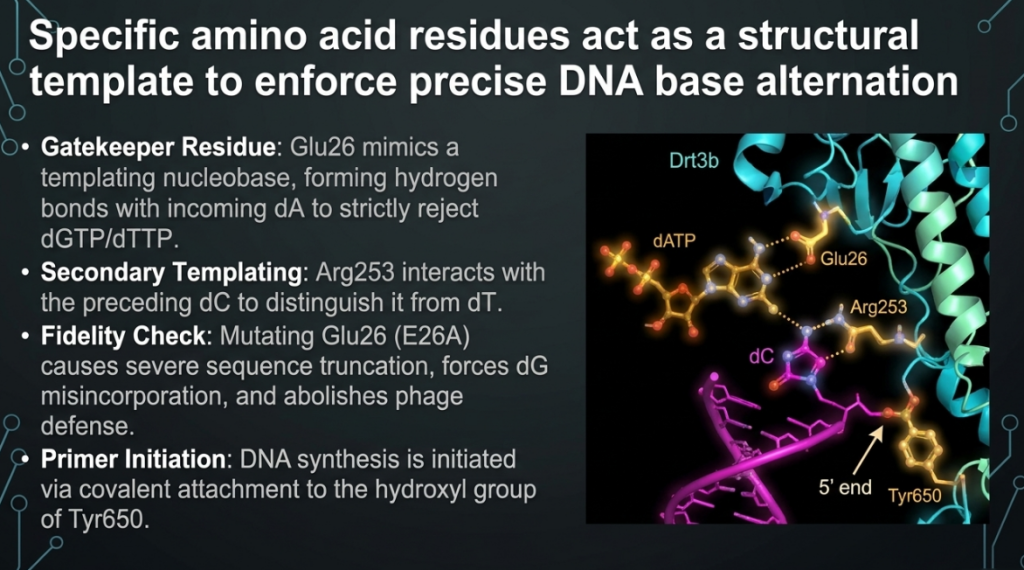
- The Protein Template: Instead of Watson-Crick base pairing, Drt3b uses a “residue duo” of Glu26 and Arg253 to select the correct bases. Glu26 forms hydrogen bonds to select adenine, while Arg253 provides a cation-π interaction that stabilizes the incoming base.
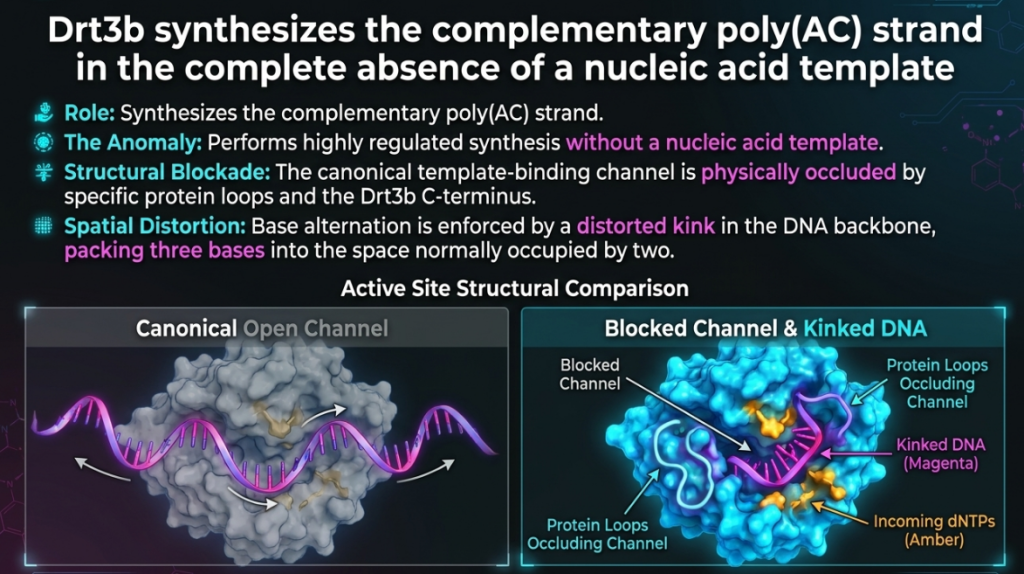
- The Kinked Backbone: To “feel” these bases without a template, Drt3b forces the DNA into a distorted, “kinked” geometry, compressing three bases into the physical space usually occupied by two. This geometric distortion is the key to protein-templated synthesis.
Finally, these two independently synthesized strands anneal to form the final double-stranded weapon.
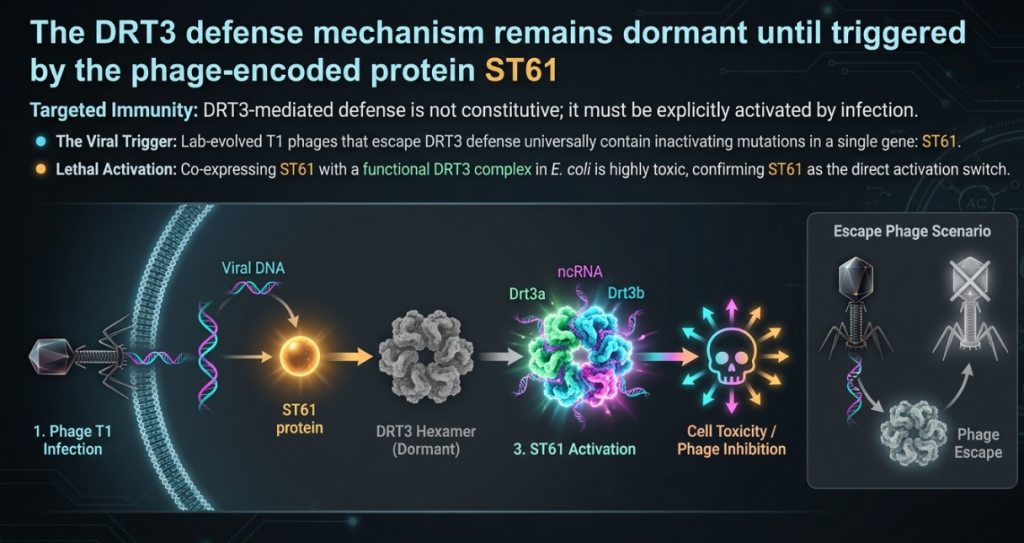

4. The Decoy Strategy: Strategic Implications of the “Molecular Sponge”
Why would a bacterium invest such significant energy into manufacturing miles of repetitive DNA? The researchers propose “The Decoy Strategy.” By flooding the cellular environment with repetitive poly(GT/AC) “nets,” the bacteria create a molecular sponge that effectively “bankrupts” the virus. These repetitive tracts are thought to sequester essential viral DNA-binding proteins, titrating them away from the viral genome and preventing replication through resource exhaustion.
The precision of this defense is managed by ST61, a protein found in phages like T1. ST61 acts as the “phage-encoded trigger” that activates the DRT3 complex. Once triggered, the system becomes constitutively toxic, ensuring a definitive “kill-switch” response to infection.
For the biotech industry, the implications are vast. We are looking at a programmable, triggerable DNA synthesis engine. Beyond understanding bacterial immunity, this technology could be repurposed for:
- Synthetic Biology: Engineering custom “kill-switches” or decoy systems.
- Genomic Data Storage: Using protein-templated enzymes to “write” data without RNA templates.
- Antiviral Therapy: Developing “molecular sponges” that neutralize human viruses by exhausting their replication machinery.
5. Conclusion: Redefining the Possible
The discovery of the DRT3 system by Deng et al. is a milestone that redefines the limits of molecular biology. It demonstrates that the “central dogma” is not a rigid law, but a set of common practices that life can override through sheer mechanical ingenuity. By proving that a protein can serve as a sequence-specific template for DNA, this research fundamentally expands the biological toolkit for information processing.
As we look toward the future of biotechnology and genomic research, the DRT3 system stands as a reminder that the most revolutionary “innovations” are often those that have been perfected over billions of years in the microscopic trenches of the bacterial arms race. This isn’t just a discovery of a new enzyme; it is the opening of a new chapter in our ability to write the code of life.
Image Summary
Question/Answer
1. Why is protein-templated DNA synthesis considered a unique biological discovery?
Protein-templated DNA synthesis is considered a unique biological discovery because it represents a distinct mode of nucleotide polymerization that defies the traditional classification of nucleic acid polymerases.
The uniqueness of this mechanism, specifically within the Drt3b subunit of the DRT3 system, is characterized by several key factors:
- Absence of a Nucleic Acid Template: Most polymerases are either template-directed (copying an existing DNA or RNA sequence) or template-independent (producing random or low-complexity homopolymers). Drt3b, however, synthesizes a specific, alternating poly(AC) strand de novo without any nucleic acid template.
- Protein Residues as “Templating Bases”: Instead of using Watson-Crick base pairing with a DNA or RNA strand, Drt3b uses conserved active site residues to dictate the sequence. Specifically, the side chain of Glu26 projects into the binding pocket and mimics a templating nucleobase, forming hydrogen bonds to select for dATP.
- Enforced Sequence Specificity: While other template-independent polymerases like AbiK generate random DNA, Drt3b uses a network of base-specific and conformation-stabilizing interactions to enforce precise base alternation. Another residue, Arg253, is hypothesized to act as a secondary template to facilitate dCTP incorporation.
- Structural Occlusion of Traditional Templates: The structure of Drt3b is specialized to physically block the entry of a nucleic acid template. Its own C-terminus and unique internal loops occupy the space where a template strand would normally reside, forcing the enzyme to rely entirely on its internal protein structure for sequence information.
- Synthesis of Long, Specific Polymers: While very limited forms of protein-templated incorporation exist in other enzymes (like Rev1 adding a single cytosine or CCA-adding enzymes forming short motifs), Drt3b is “highly atypical” because it generates long, sequence-specific nucleic acids.
These findings significantly expand the known functional landscape of polymerases by revealing that biological information transfer can occur directly from a protein structure to a DNA sequence, bypassing the need for a nucleic acid blueprint for that specific strand.
Reference
Deng, P., Lee, H., Armijo, C., Wang, H., & Gao, A. (2026). Protein-templated synthesis of dinucleotide repeat DNA by an antiphage reverse transcriptase. Science (New York, N.Y.), eaed1656. https://doi.org/10.1126/science.aed1656




