1. Introduction
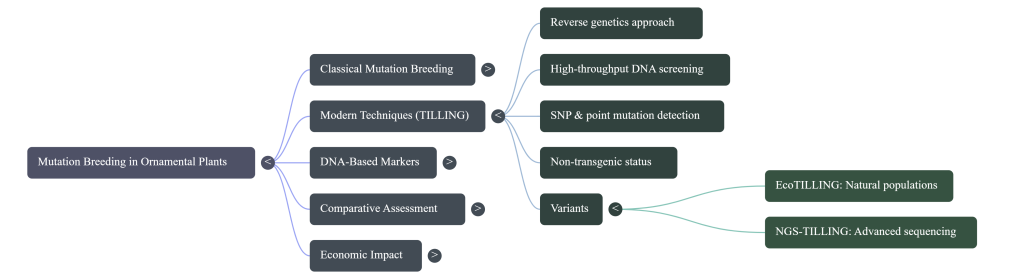
Targeting Induced Local Lesions IN Genomes (TILLING) represents a sophisticated evolution in plant genomics, acting as a critical bridge between stochastic induced mutations and precise high-throughput screening. As a robust reverse genetics platform, TILLING allows researchers to identify specific DNA lesions within genes of interest, effectively bypassing the limitations of traditional phenotypic selection. The strategic importance of this technology is underscored by the global release of approximately 3,377 mutant varieties, of which 728 are ornamental—with Chrysanthemum alone accounting for roughly 288 to 293 reported mutant varieties. Understanding the technical framework of TILLING is essential for appreciating its role in current agricultural advancements and its capacity to meet evolving commercial demands.

| Feature | Forward Genetics | Reverse Genetics (TILLING) |
|---|---|---|
| Starting Point | Visible traits (phenotypes). | DNA sequences (genotypes). |
| Process | Screen for plants with physical changes, then find the gene. | Induce mutations, then screen DNA for specific changes before traits appear. |
| Main Advantage | Identifies obvious physical traits. | Identifies mutations in genes that cannot be surely selected via visible phenotypes. |
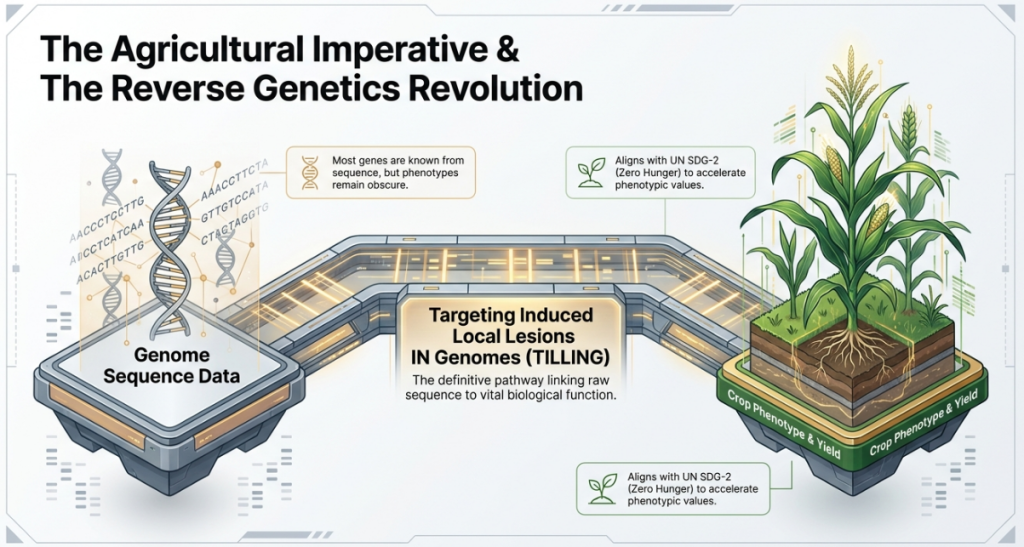
2. The Mechanics of TILLING: From Mutagenesis to DNA Screening
TILLING transforms the inherently random nature of chemical mutagenesis into a targeted discovery tool. By shifting focus from a “phenotype-first” to a “DNA-first” approach, breeders can identify mutations at the molecular level before they manifest in the plant. This is particularly vital in vegetatively propagated ornamentals where mutations often appear as chimeras; TILLING, combined with direct in vitro regeneration, provides a standardized pathway for isolating chimeric tissues into stable, solid mutants.
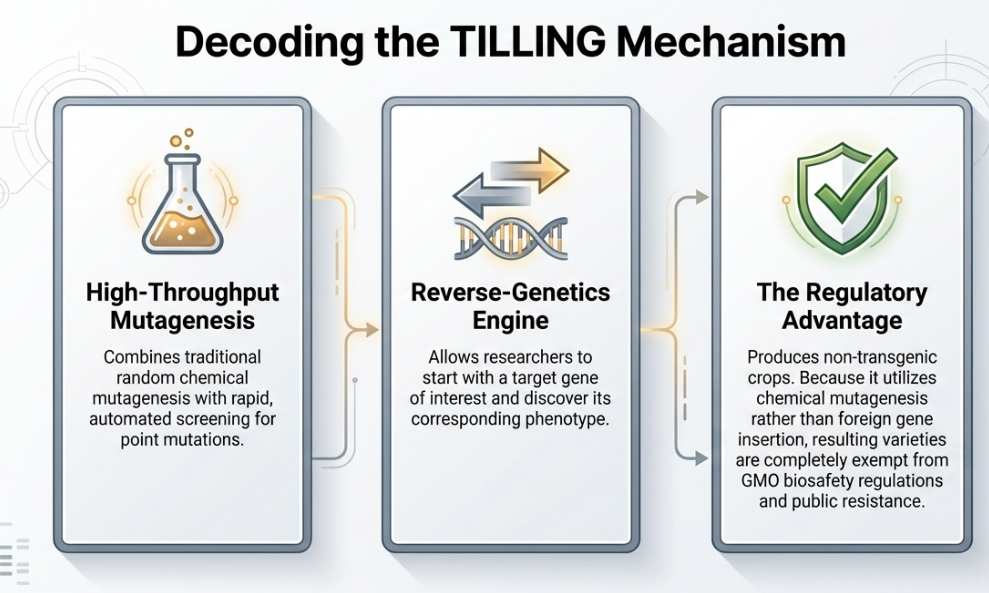
The multi-step process of TILLING, as refined in modern molecular breeding, includes:
Step 1: The Spark of Change (Mutagenesis)
The journey begins with Mutagenesis, the intentional creation of genetic variation. While various radiations can be used, TILLING primarily utilizes a chemical mutagen called Ethyl Methane Sulfonate (EMS).
- The Molecular Glitch: EMS is highly effective at inducing point mutations. Specifically, it creates “transition mutations,” where the DNA base pairs G/C are swapped for A/T.
- The Balancing Act: A critical step for any molecular biologist is Dose Optimization. We aim for the LD50 (Lethal Dose 50%), a concentration of EMS strong enough to saturate the genome with mutations without killing more than half of the population.
Scientist’s Note: Seeds are the preferred “propagule” (starting material) for TILLING. They are durable, easy to treat in mass quantities, and serve as the foundation for a permanent genetic library.
Once these seeds are treated, they carry the potential for thousands of unique genetic variations, ready to be grown into a living catalog.
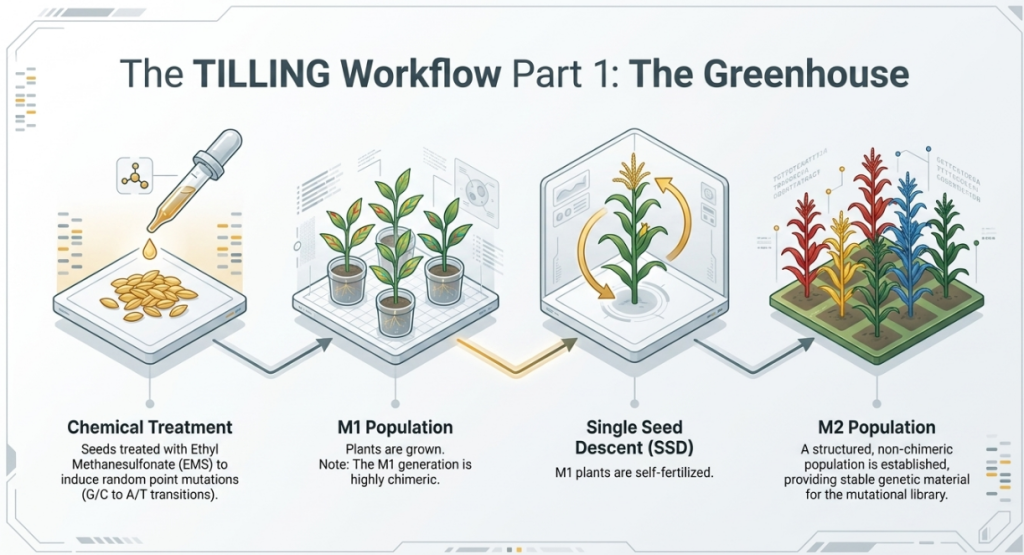
Step 2: Building the Genetic Library (M1 and M2 Generations)
Building a TILLING library is a multi-generational commitment. We don’t just study the first plants that grow; we develop a structured population:
- EMS Treatment: Seeds are soaked in the mutagen.
- The M1 Generation: These seeds are planted. The resulting plants are allowed to self-pollinate.
- The M2 Generation: The seeds from M1 plants are harvested and grown to form the M2 population.
The “Permanent Resource” Concept: The M2 generation is the heart of the TILLING platform. Because we archive both the Genomic DNA (for screening) and a Seed Bank (for future planting), the library becomes a “living catalog.” Unlike other experiments that are one-and-done, a TILLING library is a permanent source of mutations that can be screened repeatedly for different genes of interest over many years.
With our library archived, the focus shifts from the greenhouse to the high-throughput laboratory.
Step 3: High-Throughput DNA Screening (The Detective Work)
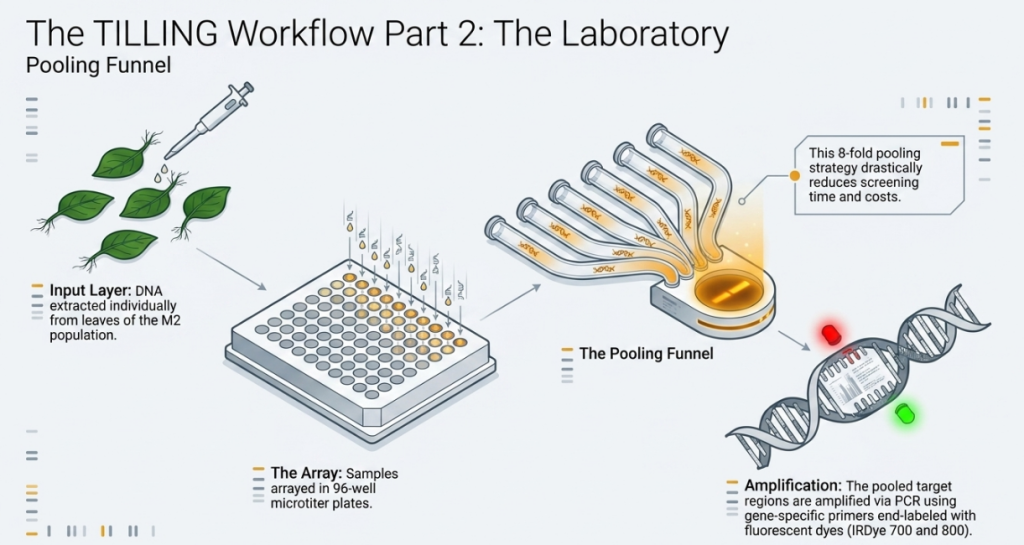
Finding a specific point mutation in a library of thousands is a massive task. To make this feasible, we use a “pooling” strategy that balances resolution with efficiency.
- The Pooling Logic: Rather than testing every plant individually, we pool DNA from 4 to 8 individuals. This significantly increases “screening throughput”—the resolution magnitude of our search—while drastically reducing the labor and cost of the screening process.
Tech Profile: The Screening Tools
- PCR Technology: Used to amplify and magnify the specific target gene or genomic region we wish to study.
- Dye-labeled Primers: These act as molecular beacons, allowing us to track and visualize the target DNA during the magnification process.
- Pooled DNA: This allows us to screen large populations quickly to identify which “pool” shelters a mutation.
Once a pool is flagged, we move to the most delicate stage: identifying the physical mismatch in the DNA.
Step 4: Finding the “Mismatches” (Endonuclease Cleavage)
At this stage, we are looking for heteroduplexes—locations where the DNA strands do not match perfectly because of the mutation.
Heteroduplex Formation: We take the magnified DNA, heat it to separate the strands, and then cool it slowly. As the strands re-pair, a mutated strand will attempt to bind with a normal (wild-type) strand. Because they don’t match perfectly, they form a “bubble” or a mismatch.
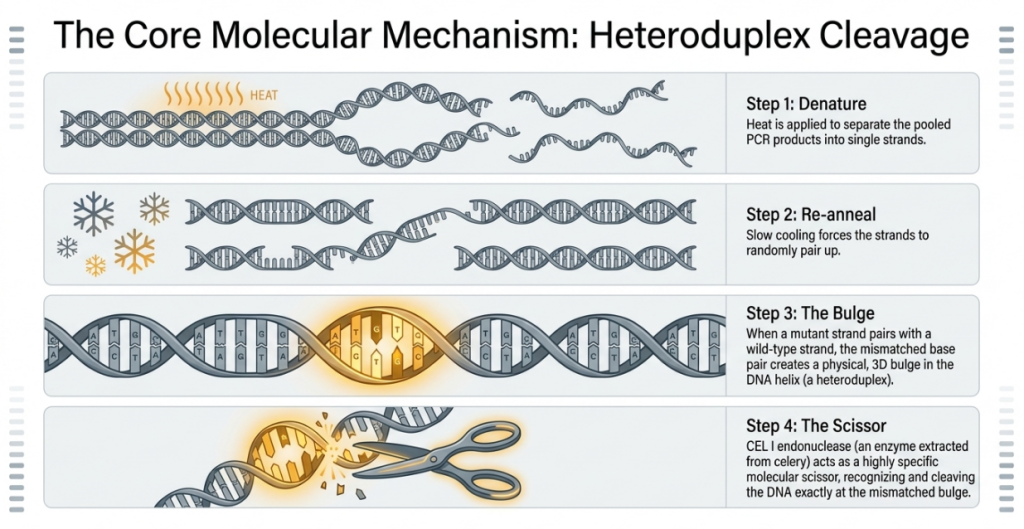
Discovery Step: Endonuclease Cleavage To find these bubbles, we introduce a specialized enzyme called an endonuclease. In a process called the mismatch cleavage assay, the enzyme acts like molecular scissors, cutting the DNA specifically at the site of the mismatch. By measuring the size of the resulting DNA fragments, we can verify exactly where the mutation is located and which plant in our pool carries it.
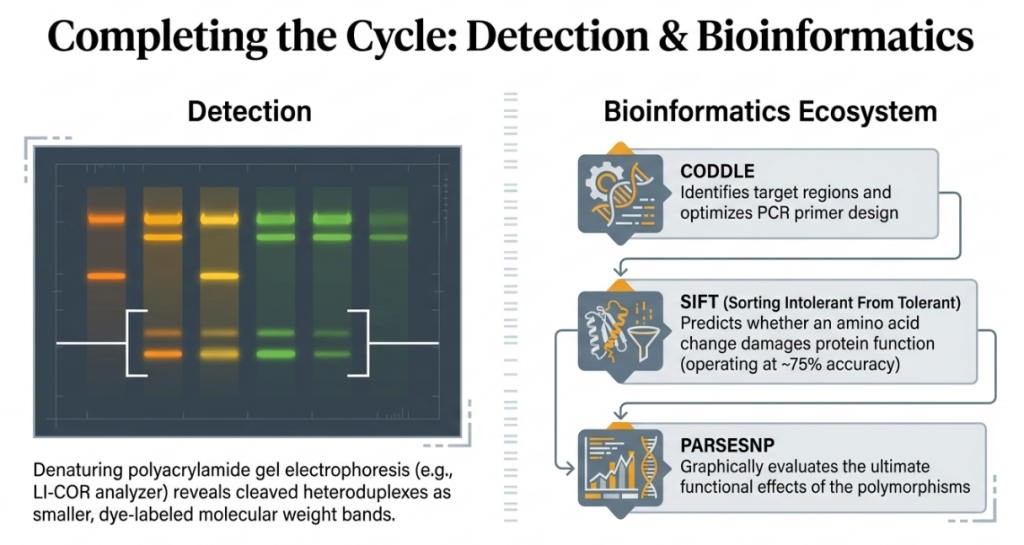
Once the plant is identified, we return to the seed bank to grow the specific M2 line and evaluate its new traits.
Summary:
- Mutagen Treatment: Seeds or propagules are typically treated with Ethyl Methane Sulfonate (EMS). EMS is the preferred chemical mutagen because it reliably induces point mutations—specifically G/C to A/T transitions—rather than large-scale chromosomal rearrangements.
- Population Development: Treated materials (M1 generation) are grown and self-pollinated to produce M2 seeds. In vegetatively propagated species, researchers meticulously track generations as M1v1, M1v2, and M1v3 to ensure mutational stability.
- DNA Assembly and Pooling: Genomic DNA is extracted and pooled (typically in groups of 4 to 8 individuals). This high-throughput pooling is essential for maximizing screening efficiency and reducing costs.
- Mutation Identification: Specific dye-labeled primers amplify target gene regions. Using a mismatch cleavage assay (often employing an endonuclease like CEL I) or advanced platforms like EMAIL (capillary electrophoresis), researchers detect polymorphisms. EcoTILLING is frequently utilized as an extension to identify natural allelic variations in wild populations.
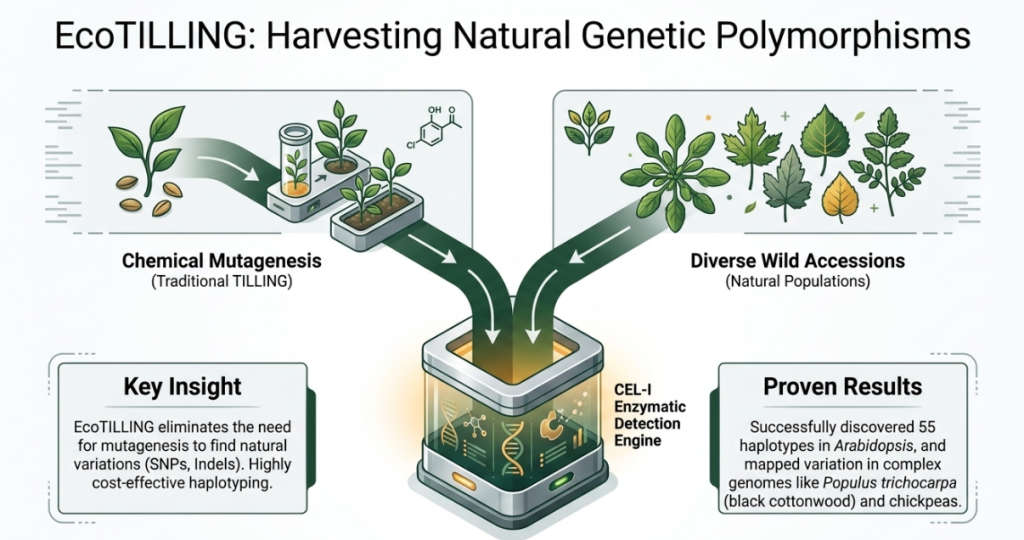
The “So What?” Layer
The transition from phenotypic screening to DNA-level analysis fundamentally alters the trajectory of variety development. By identifying “variants of interest” at the seedling stage or via stored genomic libraries, scientists can predict traits like flower color or fruit quality years before the plant matures. Furthermore, expert-level parameters such as pollen grain morphology (analyzed via Scanning Electron Microscopy or SEM) and pollen sterility serve as critical “supplementary parameters” for determining the radio-sensitivity of the material, allowing for more precise dosage optimization in early generations.
3. Comparative Assessment: TILLING vs. Classical Mutagenesis
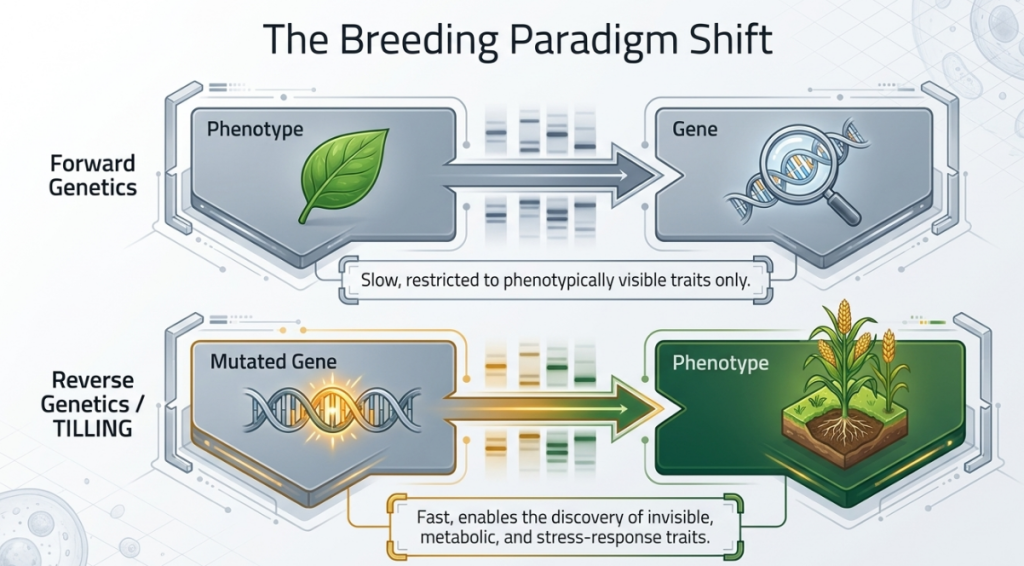
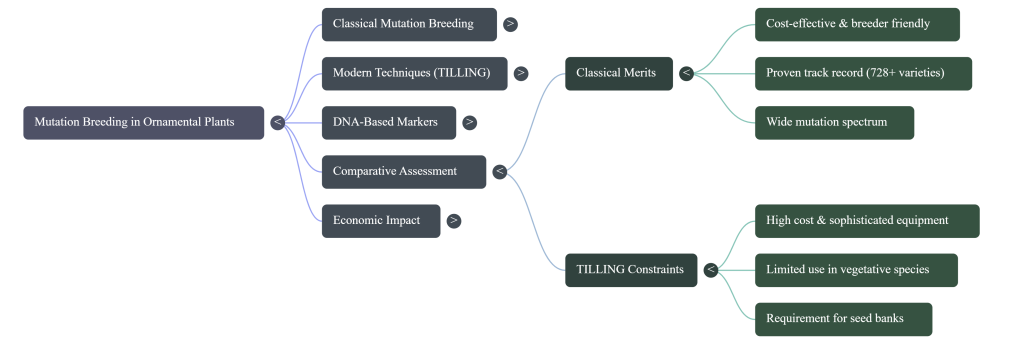
The strategic choice between “Forward Genetics” (Classical) and “Reverse Genetics” (TILLING) is often a matter of logistical maturity versus technical precision. While classical techniques wait for visible changes to manifest, TILLING starts with a known sequence and searches for a corresponding plant.
| Dimension | Classical Mutagenesis | TILLING |
|---|---|---|
| Genetic Approach | Forward Genetics (Phenotype to Gene) | Reverse Genetics (Gene to Phenotype) |
| Mutagens Used | Wide range (Gamma rays, X-rays, EMS, MMS) | Primarily EMS |
| Selection Method | Visual/Phenotypic eye-screening | High-throughput DNA screening |
| Mutation Spectrum | Wide (Color, shape, size, architecture) | Limited/Specific (Point mutations) |
| Targeting | Random and unfocused | Targets specific, known gene sequences |
| Technical Complexity | Simple, breeder-friendly, and standardized | Highly sophisticated; requires molecular expertise |
| Cost & Labor | Cost-effective; low equipment requirements | High; requires expensive reagents and equipment |
“Classical mutation technique is now well organized for almost all ornamental plants, but EMS-TILLING, EcoTILLING, NGS-TILLING promises added value. Classical mutation is a breeder-friendly technique, while TILLING is highly sophisticated.”
The “So What?” Layer
Despite TILLING’s precision, classical mutation remains the dominant force in ornamentals like Chrysanthemum. This is driven by the logistical challenges of polyploidy; in hexaploid species, designing genome-specific primers for multiallelic genes is exceptionally strenuous. Furthermore, molecular markers like RAPD often fail to differentiate flower color mutants because the bands specific to color resolution frequently fall outside the resolution capacity of tested primers. Consequently, the breeder’s ability to visually detect a “directive mutation” remains a more practical tool for immediate commercialization.
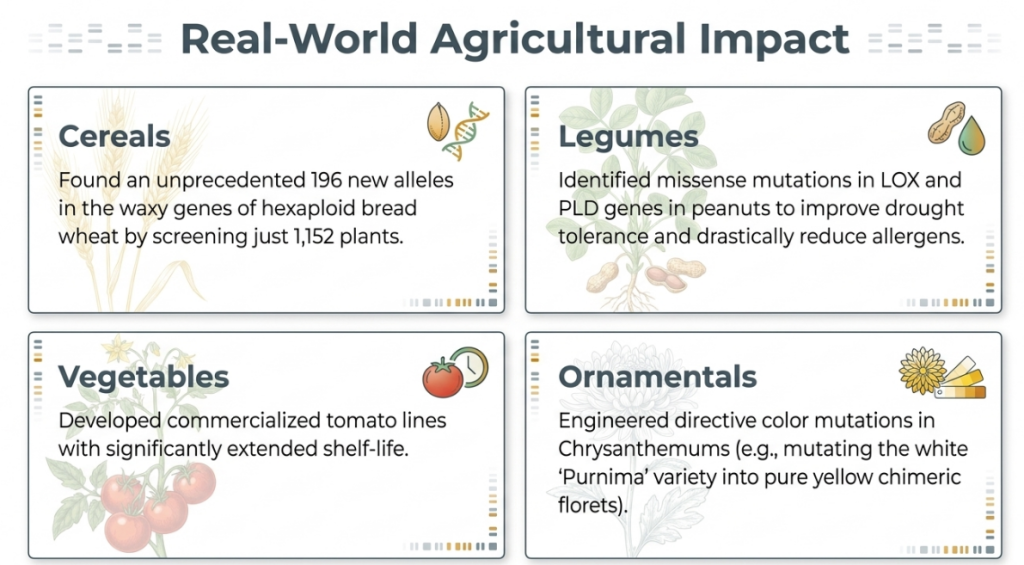
4. Strategic Applications and Impact on Global Sustainability
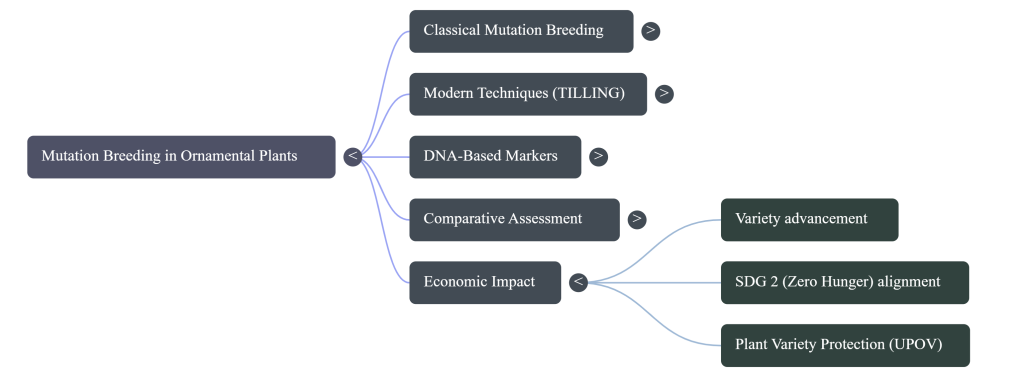
Mutation breeding is a primary driver for the United Nations Sustainable Development Goal 2 (Zero Hunger). By enhancing nutritional profiles and shelf-life, these technologies create a bridge between food security and commercial viability.
Improved Crops and Traits
- Agricultural Staples:
- Wheat: Varieties with upgraded total dietary fiber (TDF).
- Rice: Strains with significantly lower phytic acid.
- Peanuts: Development of varieties with lower allergy levels.
- Oats & Sorghum: Enhancements in protein digestibility.
- Tomato: Prolonged shelf-life via targeted metabolic mutations.
- Commercial Ornamentals:
- Chrysanthemum: The model plant for directive color mutations. Scientific logic dictates that parent pigment composition is a “consequential signal”: white varieties typically mutate to yellow, while red varieties yield either pure yellow or red/yellow mixtures.
The “So What?” Layer
A critical advantage of TILLING and classical mutation is the non-transgenic status of the resulting varieties. These crops bypass the rigorous and often prohibitive biosafety regulations that hinder GMO adoption. To identify these mutations at the biochemical level, scientists utilize Thin Layer Chromatography (TLC) and spectrophotometry—the simplest yet most effective methods to detect qualitative and quantitative pigment changes (using methanolic HCl extraction) long before full flowering.
Industrial Security and Logistical Deterrents
A primary advantage for industrial security is TILLING’s “non-transgenic” status; derived varieties are generally exempt from the stringent biosafety regulations governing GMOs. However, the logistical deterrents are significant:
- Genomic Complexity: High-density background mutations are common, requiring extensive and “time-consuming back-crosses” to restore the parental background—a major barrier to rapid speed-to-market.
- Polyploidy Hurdles: Designing genome-specific primers for polyploid ornamentals is technically strenuous and prone to failure.
5. Future Frontiers: TILLING in the Age of CRISPR/Cas9
As we enter the era of targeted genome editing, the “close of the TILLING age” is a subject of intense debate. While CRISPR/Cas9 offers site-specific mutagenesis, TILLING remains a vital resource for species where genome sequencing is incomplete.
Limitations: TILLING vs. CRISPR/Cas9
- Background Mutations: TILLING often results in high-density “background mutations” that can negatively affect plant fitness, requiring years of back-crossing to eliminate.
- Polyploidy: As noted, hexaploid genomes make TILLING primer design difficult. However, CRISPR also struggles here, as a “transmitted gene often loses its transcriptional exercise” (gene silencing) in subsequent generations.
- Tissue Culture Bottleneck: CRISPR relies heavily on Agrobacterium-mediated transformation and tissue culture, which is not yet standardized for many high-value crops.
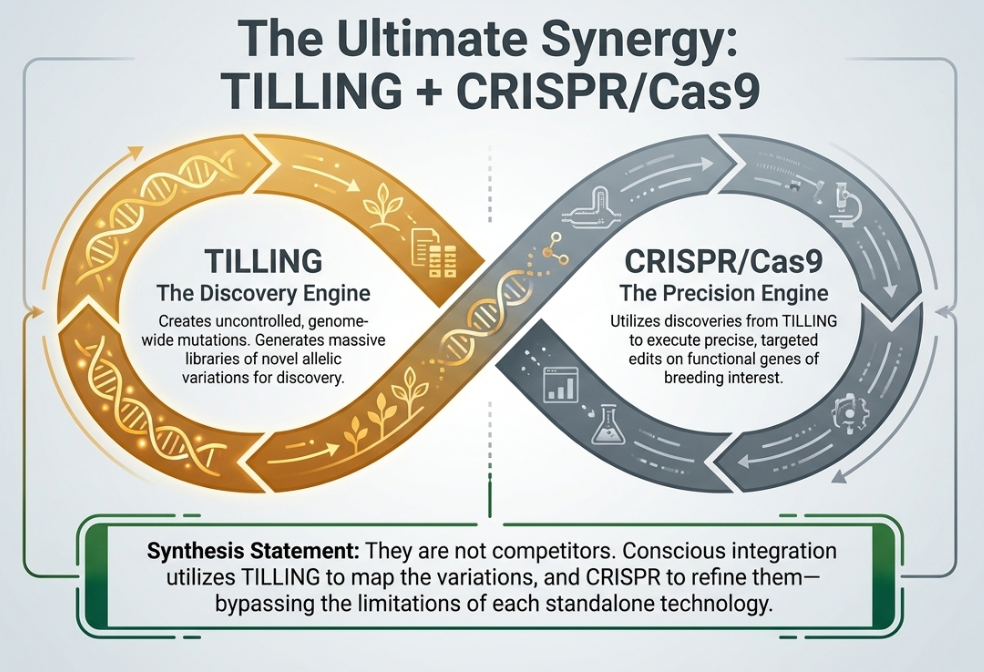
The “So What?” Layer
The modern scientist must adopt a “changed mindset,” selecting technology based on budget, technical maturity, and the biological constraints of the specific crop. While CRISPR represents the peak of precision, TILLING provides a stable, “non-transgenic” platform that is more accessible for species with high heterozygosity or complex ploidy levels where gene silencing remains a significant risk.
Comparative Assessment: Classical Mutagenesis vs. TILLING and CRISPR
Selecting the logistically appropriate breeding platform is essential for optimizing ROI and technical resources.
| Feature | Classical Mutation | TILLING | CRISPR/Cas9 |
|---|---|---|---|
| Mechanism | Forward Genetics (Phenotype first) | Reverse Genetics (Gene first) | Reverse Genetics (Target specific) |
| Targeting | Random / Directive | Gene-specific | Site-specific |
| Genomic Complexity | Ideal for polyploids | Difficult in polyploids | High complexity in hexaploids |
| Commercial Maturity | High (Proven Market ROI) | Preliminary | Infancy (in ornamentals) |
Strategic Recommendations for Industrial Breeding
To optimize R&D expenditure and ensure market novelty, the following directives are issued:
- Prioritize the Core Technology Package: Maintain “Classical Mutation + Chimera Management” as the primary engine for variety development. It remains the most cost-effective path to the M1v1 generation.
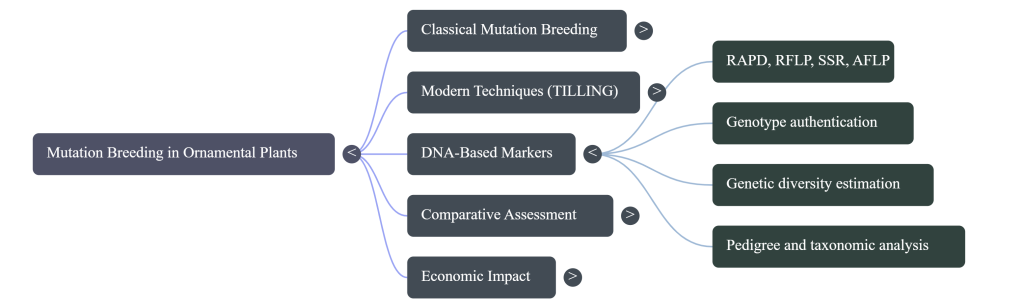
- Strategic Marker Implementation: Utilize DNA-based markers (RAPD, RFLP, SSR, VNTR, STS, and AFLP) specifically for variety authentication and intellectual property documentation rather than primary mutation detection.
- Mandatory Biochemical Screening: Integrate early-stage pigment screening using the Four-Pathway Logic (identifying increases/decreases in intensity, pigment blockage, new pigment synthesis, or combined synthesis/blockage). This reduces nursery costs by culling non-desirable variants early.
- Feasibility-First Transition: Conduct rigorous feasibility studies before shifting budgets to TILLING or CRISPR. These studies must account for high reagent expenses, skilled manpower costs, and the specific genomic complexities (polyploidy) of the target crop.

The TILLING Advantage: Why it Matters for Our Future
It is important to remember that while the mutation process is random, the “Targeting” in TILLING refers to our ability to screen for specific genes. This distinguishes it from classical methods while offering unique benefits:
- Non-Transgenic Status: Unlike GMOs, TILLING plants are produced through traditional chemical mutagenesis. This means they are exempt from the strict biosafety regulations imposed on transgenics, allowing improved crops to reach farmers much faster.
- Target Specificity: We can “aim” our screening at genes known to control vital traits, such as disease resistance or environmental adaptation.
- Broad Application & Food Security: TILLING has been successfully utilized to address global health and nutrition goals (aligning with the UN’s Zero Hunger/SDG 2), including:
- Wheat: Upgrading total dietary fiber.
- Tomato: Maximizing shelf-life.
- Rice: Reducing phytic acid.
- Oats & Sorghum: Elevating digestibility.
- Peanuts: Lowering allergy levels.

6. Conclusion
TILLING provides a powerful reverse-genetic platform that has modernized the detection and utilization of plant mutations. However, its successful application requires balancing technical sophistication with the practical, “breeder-friendly” goals of traditional mutagenesis. While we are at the dawn of the genome-editing era, the integration of multidisciplinary approaches—combining classical cytology, biochemical TLC analysis, and molecular precision—remains the most reliable path for ensuring global food security and the continued evolution of the floral trade.
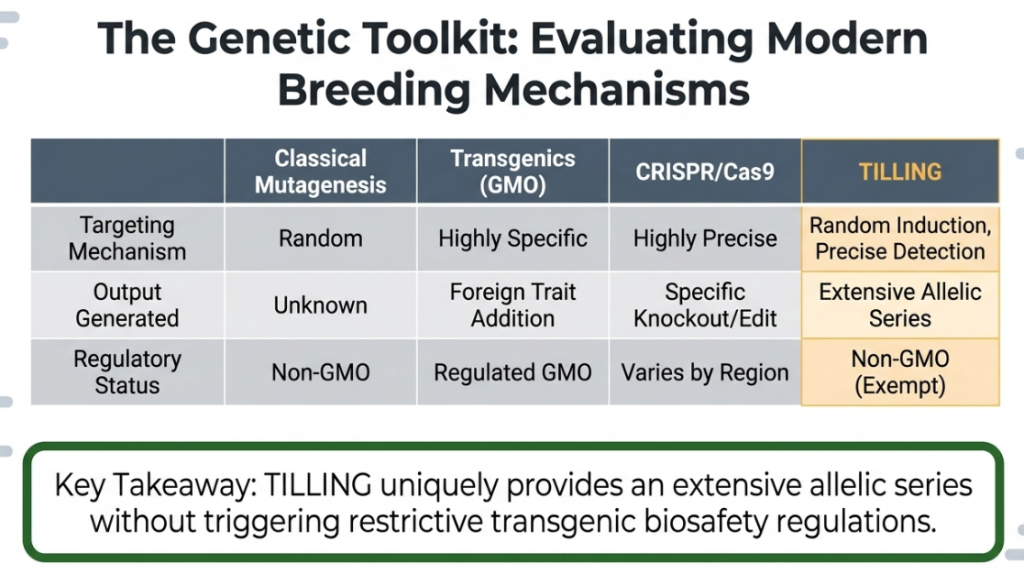

Image Summary
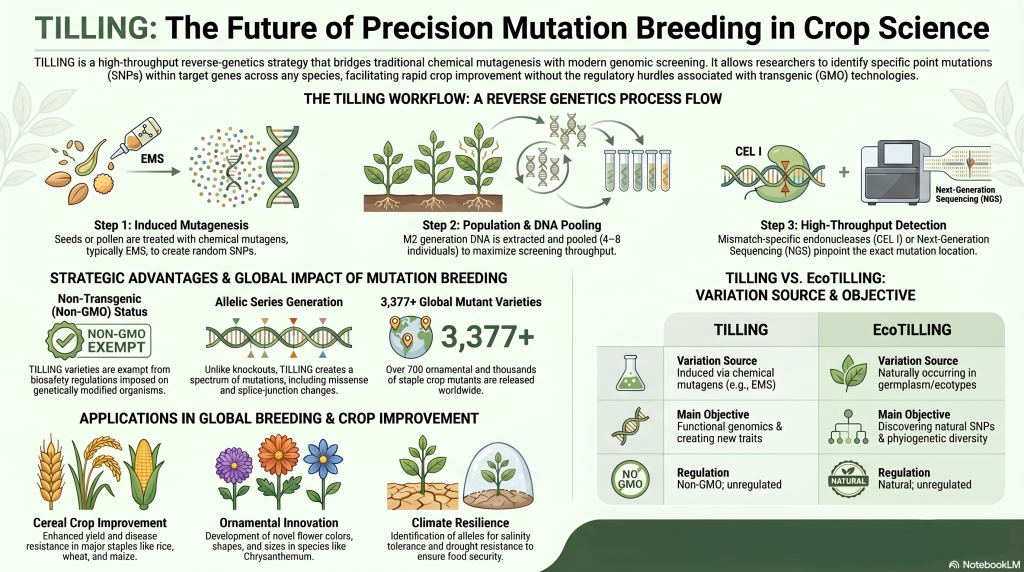
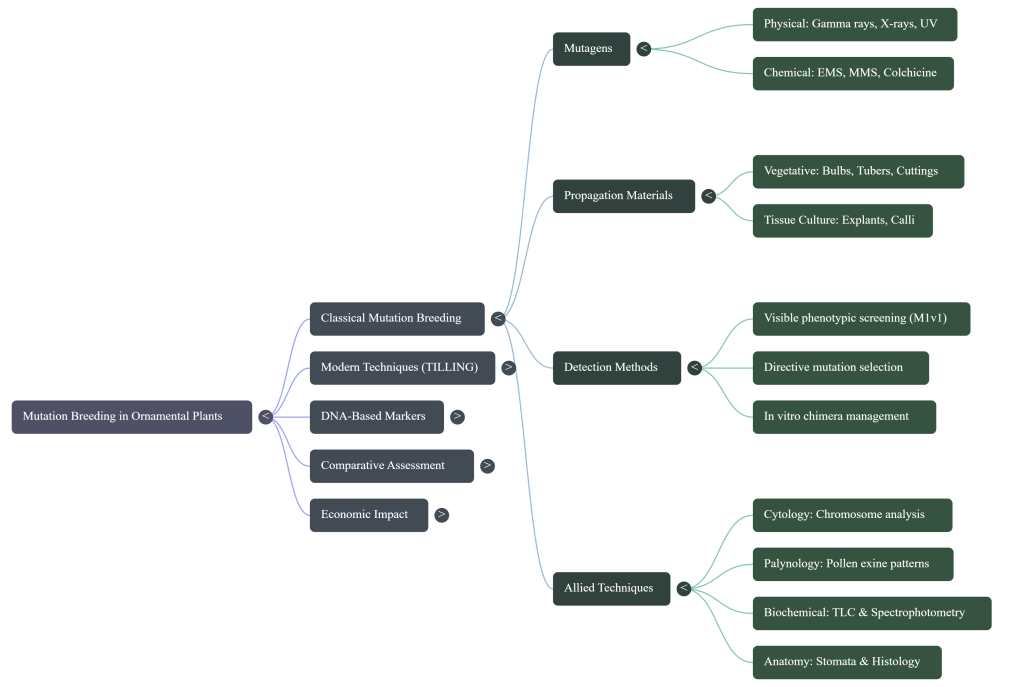
Questions/Answers
1. Explain the step-by-step process of creating a TILLING library.
The creation of a TILLING library (also referred to as a TILLING platform) is a multi-year process that combines traditional mutagenesis with modern high-throughput DNA screening techniques. This process establishes a permanent, renewable resource for identifying genetic variations in genes of interest.
The step-by-step process is as follows:
1. Mutagenesis and Dose Optimization
The process begins with the selection of plant material (typically seeds) and a mutagen. Ethyl methane sulfonate (EMS) is the most prominent chemical mutagen used because it reliably induces a high density of random point mutations, primarily G/C to A/T transitions.
- LD50 Standardization: Before treating a large population, a pilot experiment is conducted to determine the LD50 (Lethal Dose 50%), which is the concentration of mutagen that allows 50% of the treated material to survive. This ensures a high frequency of mutations without excessive lethality.
2. Developing the M1 and M2 Generations
- M1 Generation: The treated seeds (M0) are sown to grow the M1 generation. Because the initial mutagenic treatment affects cells differently, M1 plants are chimeric, meaning different parts of the same plant may carry different mutations.
- M2 Generation: To establish stable, heritable mutations and eliminate chimerism, M1 plants are self-pollinated to produce M2 seeds. One or a few seeds from each M1 plant are typically grown into M2 plants to maximize the number of independent mutations represented in the population while avoiding the repeated selection of the same mutation.
3. Sampling and DNA Extraction
Tissue (usually leaf material) is collected from thousands of individual M2 plants for genomic DNA extraction. Proper documentation is essential at this stage to link each DNA sample to its corresponding physical plant and seeds.
4. Archiving and Platform Establishment (The Library)
A mutated population officially becomes a TILLING library once the DNA and seeds are archived for long-term use:
- DNA Library: Extracted DNA samples are normalized to a standard concentration and stored in plates (often 96-well formats).
- Seed Bank: Corresponding M3 seeds harvested from each M2 plant are cataloged and stored in a seed bank. This physical resource allows researchers to retrieve and grow the specific mutant plants identified during screening. A typical library contains between 3,000 and 5,000 unique M2 families.
5. DNA Sample Pooling
To increase screening throughput and reduce costs, individual DNA samples are pooled together. Common strategies include 1D, 2D, or 3D pooling:
- In an 8-fold pool, DNA from eight individual plants is mixed into a single well, allowing one PCR reaction to screen eight genomes simultaneously.
- Complex multidimensional pooling allows for the precise intersection of signals, helping to identify the exact mutant carrier within a large population more quickly.
6. Mutation Identification (Screening)
Once the library is established, it can be “TILLed” for any gene of interest:
- Target Amplification: Gene-specific primers (often labeled with fluorescent dyes) are used to amplify the target region via PCR.
- Heteroduplex Formation: PCR products are heat-denatured and slowly reannealed. If a mutation is present in a pool, the mutant DNA will pair with wild-type DNA to form a heteroduplex with a mismatched base pair.
- Cleavage and Detection: A mismatch-specific endonuclease (such as CEL I from celery) is added to cleave the DNA at the mismatched site. The fragments are then separated and visualized using methods like denaturing polyacrylamide gel electrophoresis (PAGE) or High-Resolution Melting (HRM).
7. Deconvolution and Confirmation
When a mutation is detected in a pool, the individual samples that made up that pool are screened separately to identify the specific mutant plant. Finally, the target gene segment in the identified individual is sequenced to confirm the exact location and type of nucleotide change (e.g., missense, nonsense, or silent).
2. How does EcoTILLING detect natural variation in different species?
EcoTILLING (Ecotype TILLING) is a high-throughput reverse genetic technique used to discover natural polymorphisms within populations, including single nucleotide polymorphisms (SNPs), small insertions/deletions (indels), and variations in satellite sequences. Unlike standard TILLING, which detects mutations induced by chemical or physical agents, EcoTILLING surveys existing genetic variation in natural germplasm, such as wild relatives, landraces, or diverse cultivars.
Detection Mechanism
The standard EcoTILLING process follows several technical steps to pinpoint variations:
- DNA Preparation: Genomic DNA is extracted from various individuals or accessions within a natural population.
- Target Amplification: Gene-specific primers, often labeled with fluorescent dyes, are used to amplify a target region of interest (typically up to 1.5 kb).
- Heteroduplex Formation: The unknown sample DNA is mixed with a known reference sequence (usually in a 1:1 ratio). The mixture is heated to denature the DNA strands and then cooled slowly to allow them to re-anneal. If the sample DNA differs from the reference, they form “heteroduplexes” containing mismatched base pairs at the polymorphic sites.
- Enzymatic Cleavage: A mismatch-specific endonuclease, most commonly CEL I from celery, is added to the mixture. This enzyme specifically cleaves one strand of the DNA at the site of the mismatch while leaving perfectly matched “homoduplexes” intact.
- Visualization and Mapping: The cleaved fragments are separated and visualized using high-resolution systems like LI-COR gel analyzers or denaturing polyacrylamide gel electrophoresis (PAGE). The size of the resulting bands indicates the precise location of the polymorphism relative to the labeled ends of the PCR product.
- Haplotype Identification: Individuals with identical band patterns represent the same haplotype. This makes the process highly cost-effective, as researchers only need to sequence one representative individual for each unique haplotype to determine the exact nucleotide change.
Modifications for Complex Species
EcoTILLING has been adapted to handle specific genomic complexities:
- Self-EcoTILLING: Developed for polyploid species or multigene families where multiple highly similar copies of a gene exist. Instead of mixing with an external reference, one internal gene locus can serve as a reference for another to identify mismatches between them.
- DEco-TILLING: A variation that uses complete digestion and unlabeled primers to reduce ascertainment bias and cost.
- EcoTbyS (EcoTILLING by Sequencing): Combines EcoTILLING with next-generation sequencing (NGS) to discover natural variations by sequencing multiplexed PCR amplicons from pooled samples.
Applications Across Species
EcoTILLING has been successfully applied to identify natural variations associated with important traits in many species:
- Rice: Used to find alleles related to salinity tolerance (OsCPK17), drought resistance, and starch synthesis.
- Wheat: Employed to characterize natural variation in genes for vernalization (VRN-A1), kernel hardness (Pina, Pinb), and starch synthesis (TaSSIV).
- Trees: In Populus trichocarpa, it has helped evaluate heterozygosity levels and linkage disequilibrium in wild populations.
- Vegetatively Propagated Crops: Used to assess polymorphisms in the Musa (banana) gene pool and in potato.
- Legumes: Mined for natural allelic variants in transcription factors associated with seed weight in chickpea and diversity in mung bean.
What are the advantages of EMS-TILLING over classical mutation?
EMS-TILLING (Targeting Induced Local Lesions IN Genomes) represents a significant technical advancement over classical mutation breeding by combining traditional random mutagenesis with high-throughput molecular screening to identify specific genetic variations.
The primary advantages of EMS-TILLING over classical mutation include:
- Transition from Forward to Reverse Genetics: Classical mutation breeding is a forward genetics approach where breeders must first observe a visible phenotype and then work to identify the underlying gene. In contrast, TILLING is a reverse genetics strategy that starts with a known gene sequence and identifies mutations within that specific gene regardless of whether they produce an immediate visible change.
- DNA-Level Detection Prior to Phenotyping: TILLING identifies mutations at the DNA level before phenotypic evaluation. This allows researchers to discover mutations in genes that might not have easily detectable morphological effects or those that are “masked” by genomic redundancy.
- High-Throughput and Efficiency: While classical mutation is labor-intensive and slow, TILLING utilizes DNA pooling strategies (where samples from 8 or more plants are screened simultaneously) and automated analyzers (like LI-COR or NGS) to process thousands of individuals rapidly and cost-effectively.
- Creation of an Allelic Series: TILLING generates an allelic series for a single gene, providing a spectrum of mutation types including missense, nonsense, and splice-site variations. This provides a much more detailed “toolkit” for functional genomics than classical methods, allowing for the study of subtle protein changes rather than just complete knockouts.
- Suitability for Polyploid Species: In polyploid crops like wheat, redundant gene copies often prevent mutations from manifesting as visible phenotypes in classical forward screens. TILLING overcomes this by using homoeologue-specific primers to detect mutations in each individual genome copy, making it a highly effective tool for complex crops.
- Saturation Mutagenesis with Smaller Populations: Because chemical mutagens like EMS induce a high density of point mutations, a relatively small TILLING population (typically 3,000 to 5,000 individuals) can achieve mutational saturation, meaning every gene in the genome is likely to be mutated at least once within that library.
- Permanent and Renewable Resource: Once established, a TILLING library (comprising archived DNA and a corresponding seed bank) serves as a permanent resource that can be “TILLed” repeatedly for any number of different genes over many years.
- Non-Transgenic Status with High Precision: While it offers precision comparable to some gene-editing techniques, TILLING is considered a non-transgenic (non-GMO) approach. Varieties developed through TILLING generally do not require the same extensive regulatory approval as transgenic crops, making it more attractive for commercial agricultural applications.
3. How does EcoTILLING differ from standard TILLING?
EcoTILLING (Ecotype TILLING) is a high-throughput modification of the standard TILLING (Targeting Induced Local Lesions IN Genomes) method. While they share a similar technical foundation for mutation detection, they differ fundamentally in the source of genetic variation they analyze, the populations used, and their primary applications.
1. Source of Genetic Variation
- Standard TILLING: Detects induced mutations created by treating plant material (typically seeds or pollen) with chemical mutagens like ethyl methane sulfonate (EMS) or physical mutagens like fast neutrons.
- EcoTILLING: Detects natural polymorphisms that already exist within a species, including single nucleotide polymorphisms (SNPs), small insertions/deletions (indels), and microsatellite variations.
2. Population Development
- Standard TILLING: Requires a multi-year process to create and maintain a mutagenized population (e.g., M1 and M2 generations) before screening can begin.
- EcoTILLING: Screens natural germplasm, such as wild relatives, landraces, diverse cultivars, or ecotypes, eliminating the need for an initial mutagenesis step.
3. Technical Procedures and Pooling
- DNA Pooling:
- Standard TILLING typically uses multidimensional pooling (e.g., 4-fold to 8-fold) of DNA from different mutagenized plants to increase screening throughput and reduce costs.
- EcoTILLING often does not use sample pooling because multiple natural polymorphisms can exist within a single DNA segment, which would complicate detection in a mixed pool.
- Heteroduplex Formation:
- In Standard TILLING, heteroduplexes are formed between wild-type and mutant DNA within a pool after PCR amplification.
- In EcoTILLING (especially for inbred, homozygous species), the unknown sample DNA is mixed with a known reference sequence (typically in a 1:1 ratio) to ensure that if a natural polymorphism exists, it will form a detectable mismatch.
4. Primary Objectives and Applications
- Standard TILLING: Focuses on functional genomics (determining gene function) and creating novel alleles that may not exist in nature to improve traits like shelf life, starch quality, or disease resistance.
- EcoTILLING: Primarily used for allele mining to discover useful natural variations, evaluating levels of heterozygosity, characterizing phylogenetic diversity, and performing association mapping and linkage disequilibrium analysis.
Summary of Differences
| Feature | Standard TILLING | EcoTILLING |
|---|---|---|
| Variation Type | Induced (Chemical/Physical) | Natural (SNPs/Indels) |
| Population | Mutagenized (M1, M2, M3) | Natural (Ecotypes/Landraces) |
| Pooling | DNA from 4–8 individuals | 1:1 mixture with reference DNA |
| Primary Use | Gene function discovery | Biodiversity/Natural allele mining |
| Regulatory Status | Non-transgenic (Exempt) | Natural variants (Exempt) |
4. What challenges exist for TILLING in ornamental plants?
The application of TILLING in ornamental plants faces several unique technical, genomic, and economic challenges that have resulted in the technology still being in its infancy for this sector compared to major field crops.
The primary challenges identified in the sources include:
1. Vegetative Propagation and Chimerism
Most ornamental plants are vegetatively propagated (via bulbs, tubers, corms, or cuttings) rather than by seed.
- Difficulty in Library Creation: Establishing a TILLING population for these species is a slow and difficult process compared to seed-propagated crops.
- Chimera Management: Mutations in vegetatively propagated ornamentals often appear as chimeras (where only a part of a branch or petal is mutated). Isolating these into solid, stable mutants requires specialized tissue culture techniques that are not yet standardized for all major ornamental species.
- Inheritability: Stabilization of mutations through successive rounds of tissue culture is necessary to ensure they are heritable, which adds significant time and cost.
2. Genomic Complexity and Polyploidy
Many commercially important ornamentals have complex polyploid genomes.
- Primer Specificity: Designing genome-specific primers is strenuous and often impossible when multiple highly similar homeologous gene copies exist.
- Genetic Redundancy: In polyploids, redundant gene copies can mask the effects of a single induced mutation, making it difficult to identify recessive variants without extensive and time-consuming backcrossing.
3. Technical and Economic Barriers
The sophistication of TILLING presents a significant hurdle for traditional floriculture breeders.
- High Cost and Sophistication: The requirement for expensive automated equipment (like LI-COR analyzers or NGS platforms), specialized reagents, and highly skilled manpower restricts the economic feasibility for many ornamental programs.
- Efficiency and Mutation Density: Low mutation rates in certain species can drastically increase the cost and labor required for screening. Conversely, high doses of mutagens can lead to acute injury or high lethality, making it difficult to strike a balance between a high mutation density and plant survival.
4. Commercial Trait Desirability
In ornamentals, the “desirability” of a mutation is strictly tied to specific economic characteristics like flower color, shape, or vase life.
- Random Nature of Mutations: TILLING induces random point mutations across the genome, many of which may not affect the specific commercial traits desired by the floriculture trade.
- Background Mutations: Because mutagenesis is random, background mutations occur throughout the genome. These can negatively affect the plant’s phenotype and must be eliminated through multiple generations of backcrossing, which is particularly difficult in species with long generation times.
5. Lack of Standardized Protocols
While classical mutation techniques are well-organized for ornamentals, TILLING protocols are still in a preliminary phase for most species. There is a critical need to standardize effective procedures for inducing mutations in different propagules and for the long-term preservation of DNA and seed/tissue banks for these diverse plants.
If you would like a detailed comparison between TILLING and classical mutation techniques specifically for ornamental variety development, I can create a tailored report for you. Would you like me to do that?
5. What are the differences between TILLING and CRISPR/Cas9?
The primary difference between TILLING and CRISPR/Cas9 lies in the precision and nature of the mutations they induce. While CRISPR/Cas9 is a targeted genome-editing tool that creates specific changes at a predetermined location, TILLING is a reverse-genetics approach that identifies mutations within a population that has been subjected to random, uncontrolled mutagenesis.
The following are the key differences between the two technologies:
1. Precision vs. Randomness
- CRISPR/Cas9 is highly targeted and precise, using a Cas9 nuclease and a single guide RNA (sgRNA) to create double-strand breaks at a specific location in the genome.
- TILLING relies on chemical (e.g., EMS) or physical mutagenesis, which induces random mutations throughout the entire genome. Researchers must kemudian screen thousands of individuals to find a mutation in their specific gene of interest.
2. Transgenic Status and Regulation
- CRISPR/Cas9 typically requires genetic transformation (often via Agrobacterium) to deliver the editing machinery into plant cells, which can trigger strict GMO (Genetically Modified Organism) biosafety regulations in many regions.
- TILLING is considered a non-transgenic (non-GMO) technique. Because it is essentially an extension of traditional mutation breeding, varieties developed through TILLING are generally exempt from the regulatory approvals required for transgenic crops.
3. Technical Requirements and Applicability
- CRISPR/Cas9 is limited by the need for efficient plant tissue culture and transformation protocols, which have not yet been regularized for all major crop species.
- TILLING is applicable to virtually any organism that can be mutagenized, regardless of its genome size, ploidy level, or whether transformation protocols exist for that species.
4. Mutation Spectrum
- CRISPR/Cas9 is most commonly used to create knockouts by inducing small insertions or deletions (indels) that disrupt a gene’s function.
- TILLING identifies an allelic series of mutations, including missense, nonsense, and silent mutations. This allows researchers to study subtle changes in protein function rather than just a complete loss of function.
5. Side Effects and Off-Targets
- CRISPR/Cas9 can produce off-target effects, where unintended mutations occur at genomic locations with sequences similar to the intended target.
- TILLING does not have “off-targets” in the same sense, but it results in a high density of background mutations throughout the genome. These background mutations must be eliminated through multiple rounds of time-consuming backcrossing with the parent line.
Summary of Differences
| Feature | TILLING | CRISPR/Cas9 |
|---|---|---|
| Precision | Random (Genome-wide) | Precise (Site-specific) |
| Method | Chemical/Physical Mutagenesis | sgRNA-guided Cas9 Nuclease |
| Transgenic Status | Non-Transgenic | Often Transgenic (GMO) |
| Mutation Type | Allelic Series (Point mutations) | Targeted Indels (Often Knockouts) |
| Applicability | Any species | Limited by transformation success |
6. Why is Chrysanthemum considered a model plant for TILLING research?
Chrysanthemum is considered a model plant for TILLING research primarily because of its extensive history in mutation breeding and the highly standardized protocols available for its genetic manipulation. It serves as a benchmark for evaluating technological advancements, such as the transition from classical mutagenesis to high-throughput DNA screening techniques like TILLING.
The specific reasons Chrysanthemum is regarded as a model in this field include:
- Extensive Mutagenesis History: It is one of the major ornamental crops where the most comprehensive work has been performed regarding improvement through induced mutagenesis. Globally, it holds the record for the highest number of mutant varieties developed (approximately 293), which is significantly higher than any other ornamental species.
- Standardized Experimental Protocols: Ideal experimental methods have been endorsed and standardized for the species, covering everything from the selection of mutagens to the use of various propagules like seeds, cuttings, suckers, and explants.
- Ease of Phenotypic Detection: Morphological changes in both vegetative and floral characters are easily detectable by the naked eye, which provides a significant advantage for identifying the results of genetic variations.
- Advanced Tissue Culture Techniques: Chrysanthemum has perfectly standardized tissue culture protocols, which are essential for managing chimerism—a common issue in mutation breeding where only a portion of the plant is mutated. These techniques allow for the isolation of chimeric tissues through direct regeneration to establish stable, solid mutant lines.
- Predictable Allelic Spectrum: Research on Chrysanthemum has demonstrated that the spectrum of induced mutations is broad and, in some cases, predictable. For example, the flower color of a parent cultivar can serve as a “directive signal” for predicting new flower color mutations, allowing breeders to target specific market demands.
- Proven Functional Amelioration: The species has already been used to successfully develop distinctive functional mutations, such as salt (NaCl) tolerance, through in vitro mutagenesis techniques.
Because of these established foundations, Chrysanthemum provides a reliable platform for researchers to compare the efficiency of newer reverse-genetic tools like TILLING and CRISPR/Cas9 against traditional forward-genetic breeding outcomes.
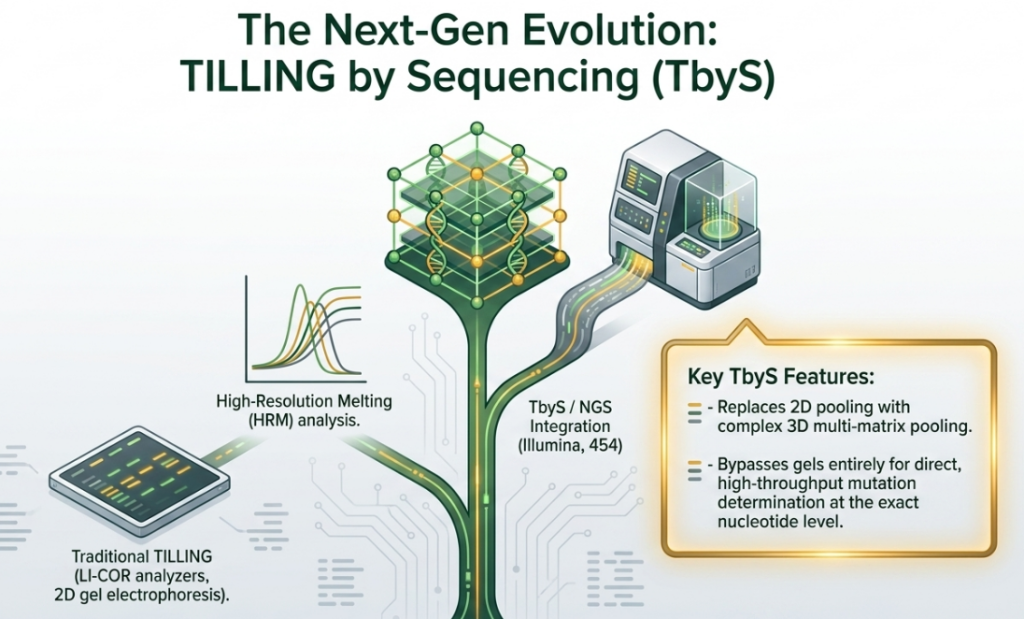
7. How does NGS-TILLING improve the mutation detection process?
NGS-TILLING (also referred to as TILLING by Sequencing or TbyS) significantly improves the mutation detection process by replacing traditional, labor-intensive enzymatic cleavage and gel-based visualization with automated, high-throughput sequencing technologies.
The key improvements provided by NGS-TILLING include:
1. Direct and Precise Mutation Discovery
Unlike traditional TILLING, which relies on the visual inspection of gel bands to estimate a mutation’s location, NGS-TILLING provides the exact nucleotide sequence of each mutation. This allows researchers to immediately determine the mutation’s impact on the protein sequence (e.g., missense, nonsense, or silent) without needing separate confirmation steps for every candidate.
2. Enhanced Sensitivity and Reliability
NGS-based methods are more sensitive than gel-based systems like LI-COR, identifying many mutants that typically go unnoticed. The process is driven by objective statistical methods for SNP calling rather than subjective visual interpretation, which reduces the risk of identifying false positives due to highly redundant sequencing.
3. Scalability and High Throughput
NGS-TILLING allows for the simultaneous screening of thousands of samples and multiple target genes in a single sequencing run. While traditional methods are limited by the number of lanes on a gel, NGS can handle massive data volumes, making it feasible to screen very large populations efficiently.
4. Cost-Effectiveness and Resource Efficiency
The transition to NGS improves economic feasibility in several ways:
- Elimination of Labeled Primers: TbyS does not depend on expensive fluorescently labeled primers required for LI-COR systems.
- No Mismatch-Specific Enzymes: It bypasses the need for endonucleases like CEL I, which can be expensive or inconsistent.
- Falling Sequencing Costs: The dramatic decrease in DNA sequencing costs over the last decade has made NGS more affordable than many older genotyping methods.
5. Advanced Pooling Strategies
NGS-TILLING utilizes complex multidimensional pooling (e.g., 2D or 3D). A 3D pooling strategy allows for the precise molecular recognition of a mutation and its specific carrier plant through the intersection of signals across different dimensions, often eliminating the need for additional “deconvolution” (re-screening individuals from a pool).
6. Applicability to Complex Genomes
NGS-TILLING is particularly beneficial for polyploid species (like wheat) and multigene families. Techniques like exome capture allow researchers to sequence only the coding regions (exons) of the genome, ignoring the vast, non-functional sequences (introns) that characterize large plant genomes. Specialized bioinformatics tools, such as MAPS (mutations and polymorphisms surveyor), further help distinguish between induced mutations and natural variations among highly similar homeologous genes in polyploids.
8. What are the benefits of Self-EcoTILLING in polyploid species?
Self-EcoTILLING is a specialized modification of the EcoTILLING protocol designed to identify single nucleotide polymorphisms (SNPs) specifically within polyploid species and multigene families. It addresses the technical difficulties that arise when multiple, highly similar copies of a gene exist within the same genome.
The primary benefits of Self-EcoTILLING in polyploid species include:
- Handling Genomic Complexity: In polyploid plants, many genes belong to multigene families that share partial to high sequence homology, often leading to nonspecific PCR amplification. Self-EcoTILLING overcomes this by allowing the identification of mismatches between these internal gene copies (homologues).
- Internal Reference System: Unlike standard EcoTILLING, which requires mixing an unknown sample with an external reference sequence, Self-EcoTILLING uses one internal gene locus to serve as the reference for another. PCR products from different homologues are allowed to form heteroduplexes with each other, which are then cleaved at mismatch sites by endonucleases like CEL I.
- Significant Cost and Time Savings: Because it eliminates the experimental steps required to prepare and mix external reference and query DNA, the technique is considerably more cost-effective and rapid than standard TILLING or EcoTILLING.
- High Precision in Multigene Families: The technique was successfully demonstrated in the allotetraploid species Monochoria vaginalis, where it pinpointed specific SNPs in the acetolactate synthase (ALS) multigene family that conferred herbicide resistance.
- Detection of Transgenes and Edits: Beyond natural variation, Self-EcoTILLING has the potential to be used for identifying transgene integration sites or screening for specific mutations induced by CRISPR/Cas9 gene editing.
While highly efficient for identifying mutations, the sources note that Self-EcoTILLING does not automatically provide information on the functional consequences of the identified variations; it is often combined with other techniques like RNA-Sequencing or proteomics to understand the impact on gene expression or protein function.
9. Can TILLING be applied to vegetatively propagated ornamental plants?
Yes, TILLING can be applied to vegetatively propagated ornamental plants, but the technology for this group is currently in its infancy and preliminary phases compared to seed-propagated crops. While classical mutation techniques are well-organized for almost all ornamental plants, advanced techniques like EMS-TILLING and EcoTILLING are seen as promising “added value” tools that require further standardization for vegetative species.
The application of TILLING to these plants involves specific modifications and faces unique challenges:
1. VeggieTILLING and Explant Selection
For plants that do not produce seeds or are traditionally cloned, a specialized approach often called VeggieTILLING is used. Instead of mutagenizing seeds, researchers treat meristematic tissues, such as:
- Vegetative parts: Cuttings, bulbs, tubers, corms, rhizomes, or suckers.
- In vitro cultures: Explants like shoot apices, leaves, petals, or calli used in tissue culture.
2. Managing Chimerism
A major hurdle in vegetatively propagated ornamentals is that mutations usually appear as chimeras (where only a portion of the tissue is mutated).
- In seed-propagated plants, chimerism is eliminated through self-pollination to the M2 generation.
- In vegetative plants, chimeras must be managed through successive rounds of tissue culture and direct regeneration to isolate and establish “solid,” non-chimeric mutant lines.
3. Technical and Practical Challenges
- Difficult Library Creation: Establishing a stable TILLING population (platform) for these species is a slow and difficult process.
- Mutation Density and Heritability: Ensuring that mutations are mitotically stable and heritably maintained through successive vegetative generations is more complex than in sexual reproduction.
- Lack of Protocols: Effective TILLING protocols for many major ornamental species are still being identified, particularly regarding the best type of propagule and mutagen dose to use without causing acute injury to the plant.
4. EcoTILLING as a Practical Alternative
Because creating induced mutant populations in vegetative species with long generation times is challenging, EcoTILLING is often suggested as a more immediate solution. This method avoids mutagenesis entirely by screening for natural polymorphisms within existing germplasm, such as wild relatives or diverse cultivars. This approach has already been successfully applied to other clonally propagated crops like banana and potato to identify useful natural variations.
If you are interested in a specific ornamental plant, like Chrysanthemum, which is often used as a model for these techniques due to its well-standardized tissue culture protocols, I can provide more details on that specific case. Would you like me to do that?
10. What are the limitations of TILLING in polyploid species?
While TILLING is considered a powerful tool for complex crops, polyploid species (those with multiple sets of chromosomes, such as hexaploid wheat or oats) present several unique technical and biological limitations that complicate the process.
The primary limitations of TILLING in polyploid species include:
1. Difficulty in Designing Specific Primers
Designing genome-specific or homoeologue-specific primers is strenuous and sometimes impossible because homologous genes in polyploids often share a very high degree of sequence homology. If primers are not specific enough, they may amplify multiple loci simultaneously, which alters the composition of DNA pools and can interfere with the formation of detectable heteroduplexes.
2. Phenotypic Masking by Genetic Redundancy
A major biological hurdle is genetic redundancy, where the effects of a single induced mutation are “enshrouded” or masked by the presence of functional wild-type copies (homoeologues) in the other sub-genomes. Because most induced mutations are recessive, they often fail to manifest as a visible phenotype in polyploids unless all copies of the gene are mutated.
3. Requirement for Mutation Stacking
To overcome phenotypic masking, researchers must perform “pyramiding” or stacking of mutations. This involves identifying individual mutants for each homoeologue (e.g., the A, B, and D genomes in wheat) and then crossing them to produce double or triple homozygous mutants. This process is labor-intensive and adds significant time to breeding programs.
4. Reduced Sensitivity in DNA Pooling
The increased number of genome equivalents per individual in polyploids means that more DNA targets are amplified during PCR. This can lead to:
- Reduced Detection Throughput: Researchers may need to use smaller pooling sizes (fewer individuals per pool) to maintain the sensitivity required to detect a single mutated allele amidst many wild-type copies.
- Experimental Complexity: If two targets from different sub-genomes are amplified with different efficiencies, the resulting pool composition can be further skewed, making mutation identification unreliable.
5. Increased Requirement for Backcrossing
Polyploid mutants often carry a high density of background mutations due to their higher tolerance for mutagenic agents. Eliminating these undesirable background changes requires a large number of time-consuming backcrosses to the parent line—potentially four or more cycles—to restore the desired performance of the crop variety.
6. Bioinformatic Challenges
Analyzing results in polyploids requires advanced bioinformatics tools to distinguish between induced mutations and natural divergence among the highly similar homoeologous sequences. Tools such as MAPS (mutations and polymorphisms surveyor) have been developed specifically to handle these complexities.
11. What does a TILLING project for Chrysanthemum cost?
The sources do not provide a specific total dollar amount for a TILLING project in Chrysanthemum, as the technology for this species is still in its preliminary phases and standardized protocols are still being identified. However, the sources detail the significant cost drivers and some specific fees associated with established TILLING services for other species that provide a baseline for understanding the financial commitment required.
The cost of a TILLING project for Chrysanthemum is influenced by the following factors:
1. Library Establishment (High Initial Investment)
Creating a TILLING “platform” or library is a multi-year, labor-intensive process.
- Timeframe: Establishing a population and preparing DNA samples from thousands of plants typically takes the better part of two years.
- Population Size: A typical library requires between 3,000 and 5,000 unique families to achieve mutational saturation.
- Vegetative Challenges: Because Chrysanthemum is vegetatively propagated, the process is slower and more difficult than for seed-propagated crops. Managing chimerism (where only part of the tissue is mutated) requires successive rounds of tissue culture to establish stable mutant lines, which adds significant time and expense.
2. Operational and Reagent Costs
The technical steps of TILLING involve several ongoing expenses:
- Equipment: High-throughput screening requires expensive automated analyzers, such as LI-COR gel systems or Next-Generation Sequencing (NGS) platforms.
- Reagents: Standard TILLING depends on fluorescently labeled primers and specialized mismatch-specific endonucleases (like CEL I), which can be costly. Some researchers reduce costs by using crude celery juice extract instead of purified enzymes.
- Manpower: These projects require highly skilled manpower to manage the library and interpret complex molecular data.
3. Benchmarking Against Established Services
For comparison, established public TILLING facilities for other species charge fees to help offset their costs:
- User Fees: The Arabidopsis TILLING Project (ATP) charged a fee of $500 for an initial screen or for screening the remainder of its collection.
- Subsidies: The sources note that such fees often only partially offset expenses, with the remainder being covered by large government or institutional grants.
4. Comparison to Classical Breeding
The sources characterize TILLING as being beyond the reach of many classical mutation breeders due to its sophistication and the required investment in equipment and reagents. In contrast, classical mutation breeding (using gamma rays and visual selection) is described as a money-saving and budget-friendly alternative that has already successfully produced hundreds of Chrysanthemum varieties.
In summary, while a specific price for a Chrysanthemum TILLING project is not cited, it is described as a sophisticated and expensive undertaking compared to traditional breeding, requiring specialized facilities, several years of labor, and high-end molecular reagents.
References
Datta, S. K. (2026). Impact of classically induced mutations on ornamental plants and comparative assessment of advanced techniques (TILLING) for induction and detection of mutations and mutants. In Nucleus (India). Springer. https://doi.org/10.1007/s13237-026-00665-3
De-Kai, W., Zong-Xiu, S., & Yue-Zhi, T. (2006). Application of TILLING in Plant Improvement. 33(11).
Derek L Stemple. (2004). TILLING — a high-throughput harvest for functional genomics. Nature Reviews Genetics, 5.
Gilchrist, E. J., & Haughn, G. W. (2005). TILLING without a plough: A new method with applications for reverse genetics. In Current Opinion in Plant Biology (Vol. 8, Number 2, pp. 211–215). Elsevier Ltd. https://doi.org/10.1016/j.pbi.2005.01.004
Henikoff, S., Till, B. J., & Comai, L. (2004). TILLING. Traditional mutagenesis meets functional genomics. In Plant Physiology (Vol. 135, Number 2, pp. 630–636). American Society of Plant Biologists. https://doi.org/10.1104/pp.104.041061
Irshad, A., Guo, H., Zhang, S., & Liu, L. (2020). TILLING in cereal crops for allele expansion and mutation detection by using modern sequencing technologies. In Agronomy (Vol. 10, Number 3). MDPI AG. https://doi.org/10.3390/agronomy10030405
Kurowska, M., Daszkowska-Golec, A., Gruszka, D., Marzec, M., Szurman, M., Szarejko, I., & Maluszynski, M. (2011). TILLING – a shortcut in functional genomics. In Journal of Applied Genetics (Vol. 52, Number 4, pp. 371–390). https://doi.org/10.1007/s13353-011-0061-1
Manzanares, C., Yates, S., Ruckle, M., Nay, M., & Studer, B. (2016). TILLING in forage grasses for gene discovery and breeding improvement. In New Biotechnology (Vol. 33, Number 5, pp. 594–603). Elsevier B.V. https://doi.org/10.1016/j.nbt.2016.02.009
Nouman Khalid, M., Amjad, I., Vamuyah Nyain, M., Saleem, M. S., Asif, M., Ammar, A., & Rasheed, Z. (2021). A Review: TILLING Technique Strategy for Cereal Crop Development. In International Journal of Applied Chemical and Biological Sciences (Vol. 2, Number 5). www.visnav.in/ijacbs/
Palan, B., Bhattacharya, A., & Char, B. (2021). TILLING in the era of precise genome editing. In Indian Journal of Biotechnology (Vol. 20).
R, A. A., & Vighneswaran, V. (2025). Targeting Induced Local Lesions in Genome (TILLING): An Approach towards Genetic Mutations in Rice. Journal of Experimental Agriculture International, 47(10), 169–181. https://doi.org/10.9734/jeai/2025/v47i103802
Simsek, O., & Kacar, Y. A. (2010). Discovery of mutations with TILLING and ECOTILLING in plant genomes. Scientific Research and Essays, 5(24), 3799–3802. http://www.academicjournals.org/SRE
Singh, D., Chaudhary, P., Taunk, J., Singh, C. K., Chinnusamy, V., Sevanthi, A. M., Singh, V. J., & Pal, M. (2024). Targeting Induced Local Lesions in Genomes (TILLING): advances and opportunities for fast tracking crop breeding. In Critical Reviews in Biotechnology (Vol. 44, Number 5, pp. 817–836). Taylor and Francis Ltd. https://doi.org/10.1080/07388551.2023.2231630
Slade, A. J., & Knauf, V. C. (2005). TILLING moves beyond functional genomics into crop improvement. In Transgenic Research (Vol. 14, Number 2, pp. 109–115). https://doi.org/10.1007/s11248-005-2770-x
Wen, N., Osorio, C. E., Brew-Appiah, R. A. T., Mejías, J. H., Alam, T., Kashyap, S., Reinbothe, S., Reinbothe, C., Moehs, C. P., von Wettstein, D., & Rustgi, S. (2022). Targeting Induced Local Lesions in the Wheat DEMETER and DRE2 Genes, Responsible for Transcriptional Derepression of Wheat Gluten Proteins in the Developing Endosperm. Frontiers in Nutrition, 9. https://doi.org/10.3389/fnut.2022.847635