1. Introduction: When “Identical” Isn’t Identical
In the rigorous environment of a plant biotechnology laboratory, micropropagation is employed to generate thousands of “clones” from a single donor plant. The scientific objective is genetic fidelity—ensuring that every offspring is “true-to-type.” However, a pervasive biological paradox exists: laboratory-produced clones often manifest novel traits not seen in the parent.

This phenomenon is somaclonal variation. It encompasses the genetic and epigenetic diversity that emerges during the in vitro culture process. While this variation poses a significant threat to commercial uniformity and germplasm preservation, it simultaneously offers a powerful mechanism for crop improvement. For the plant scientist, these “imperfect clones” are a reservoir of untapped traits, providing a unique path toward developing resistance and enhancing yields in sterile or polyploid crops.
This inherent instability is not a failure of the lab but a direct result of the specific laboratory conditions that trigger a profound reorganization of the plant genome.
2. The Laboratory Pressure Cooker: Environmental and Chemical Triggers
Tissue culture is fundamentally a high-stress environment. The process of wounding tissue, coupled with exposure to exogenous chemicals, acts as a mutagenic catalyst. This “pressure cooker” environment forces cells to abandon their specialized roles and enter a state of genomic vulnerability.
The Triggers of Instability
| Factor | Specific Example from Context | Biological Impact |
|---|---|---|
| Media Components | Synthetic auxins like 2,4-D and high levels of cytokinins like Benzyladenine (BA). | Induces polyploidy and endoreduplication; significantly increases DNA methylation rates. 2,4-D is linked to endoreduplication and DNA methylation changes; high BA levels can increase chromosome numbers. |
| Duration of Culture | Excessive subculture cycles or long-term maintenance. | Leads to the accumulation of variant karyotypes; the mutation rate often increases linearly with time in vitro. |
| Regeneration System | Indirect organogenesis (Regeneration via a Callus phase). | Forces dedifferentiation, causing a total loss of cellular control and a breakdown of the normal checks and balances of the cell cycle. |
| Explant Source | Meristematic tissues (pericycle, procambium) minimize variation. Highly differentiated tissues (roots, leaves) produce more variants. | Highly differentiated tissues require a callus phase for reprogramming, increasing mutation risk. |
The move from an organized meristem (like a shoot tip) to an unorganized callus is the most critical juncture for variation. During dedifferentiation, the cell cycle is no longer governed by the plant’s standard developmental architecture, making the genome susceptible to rapid, stochastic changes.

As these external pressures mount, they manifest internally through a state of heightened oxidative stress.
3. The Cellular Mechanism: From Stress to Mutation
The biochemical engine driving somaclonal variation is oxidative stress. Laboratory procedures—specifically wounding and chemical imbalances—trigger the production of Reactive Oxygen Species (ROS) such as superoxide and hydrogen peroxide. These ROS act as physical mutagens, attacking the DNA molecule directly.
Qualitative DNA Modifications Caused by ROS
- Chromosome Rearrangements: Large-scale breaking and reconnection of DNA strands.
- Base Deletions and Substitutions: Errors in the “letters” of the genetic code (A, T, C, G).
- Structural Irregularities: The formation of ring chromosomes, inversions, and acentric fragments.
- Ploidy Changes: Transitions from polyploid to aneuploid states.

It is vital for a biotechnologist to distinguish between these two classes of variation:
- Genetic Mutations: These involve permanent, heritable changes to the DNA sequence (e.g., deletions). They are stable across generations and form the basis of new cultivars.
- Epigenetic Changes: These involve modifications like DNA methylation or histone remodeling. They alter gene expression (silencing or activating genes) without changing the sequence. Crucially, these are sometimes reversible and may disappear once the stressor is removed.

These microscopic alterations eventually scale up, resulting in macroscopic, observable changes in the regenerated plant’s phenotype.


4. The Two Sides of the Coin: Beneficial vs. Harmful Outcomes
Somaclonal variation is a double-edged sword. In a commercial setting, it is often viewed as a contaminant; in a breeding program, it is a primary tool for innovation.
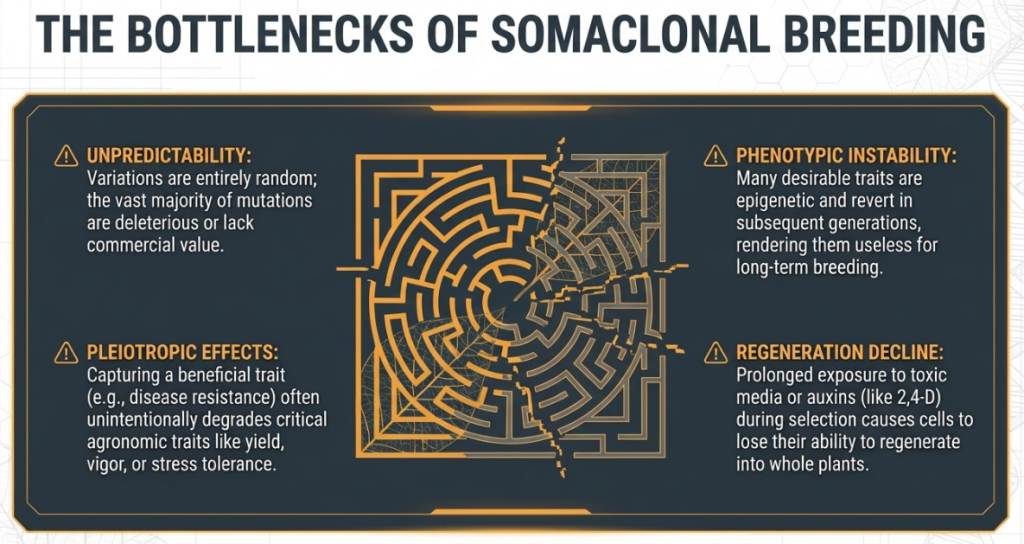
The Outcomes of Variation
Harmful / Unwanted
- Loss of Genetic Fidelity: Progeny fail to meet the performance standards of the elite donor parent.
- Dwarf Off-types: Frequently observed in ‘New Guinea Cavendish’ bananas, where the plant fails to reach commercial height.
- Deleterious Mutations: Mutations that impair plant vigor, fertility, or nutrient uptake.
Beneficial / Desirable
- Enhanced Agronomic Traits: Early flowering, heat tolerance (e.g., in potatoes), or increased essential oil content (e.g., in Patchouli).
- Improved Quality: High solid contents in tomatoes or non-browning varieties of potato.
- Durable Disease Resistance: Selection for resistance against devastating soil-borne pathogens like Fusarium.
The “So What?” for the Aspiring Scientist: While variation is a threat to laboratory uniformity, it serves as a critical tool for discovering traits that traditional breeding—especially in sterile triploids—cannot produce. We are currently transitioning from observing this stochastic (random) variation to using it as a map for targeted genome editing.
5. Case Study: Solving the Fusarium Wilt Crisis through Variation
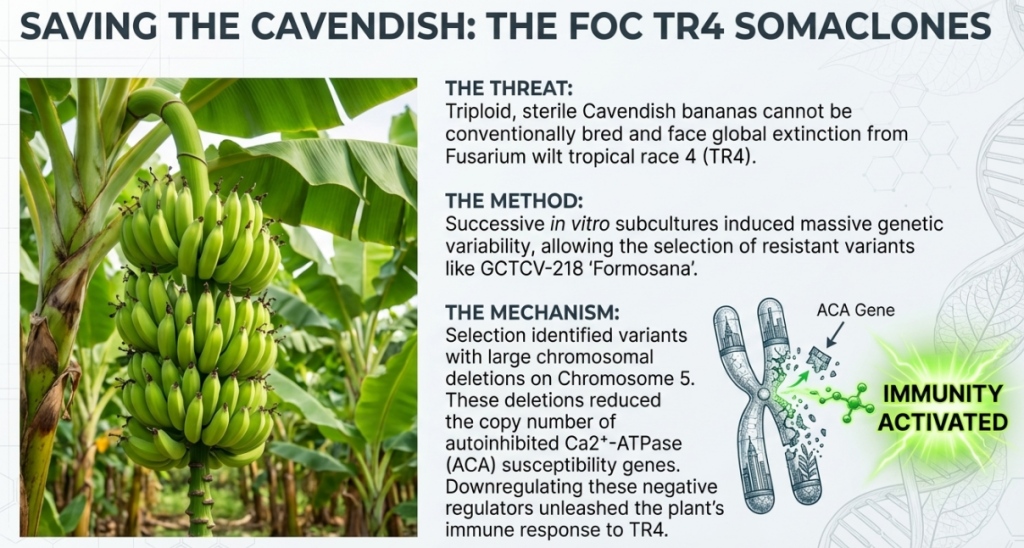
The global banana industry currently faces an existential threat from Fusarium oxysporum f. sp. cubense Tropical Race 4 (TR4), a virulent hemibiotrophic fungal pathogen responsible for the devastating Panama disease. This soil-borne fungus is exceptionally difficult to manage because it persists in the environment for decades via resilient chlamydospores, rendering traditional chemical and cultural controls ineffective. The crisis is exacerbated by the biological nature of the dominant Cavendish banana (AAA group); as a sterile triploid, it lacks the ability to be improved through conventional cross-breeding, creating a genetic bottleneck.

| Feature | Foc Race 1 (Gros Michel Era) | Foc Tropical Race 4 (Cavendish Era) |
|---|---|---|
| Primary Target | Gros Michel (AAA) | Cavendish (AAA) and diverse local varieties |
| Persistence | Decades in soil via chlamydospores | Decades in soil via chlamydospores |
| Chemical Control | Completely Ineffective | Completely Ineffective |
| Pathogen Virulence | High (Targeting Gros Michel) | Extreme (Broad host range, including Cavendish) |
| Management Complexity | High; required cultivar replacement | Absolute; requires non-conventional genomic strategies |
In this constrained landscape, somaclonal variation—the genetic and epigenetic diversity arising from plant cell or tissue culture—has emerged as a crucial resource for identifying stable disease resistance. While tissue culture is typically designed for clonal fidelity, the inherent instability of the in vitro environment can generate beneficial mutations. These variants offer a unique biotechnological pathway to bypass the limitations of sterility, providing a foundation for selecting and engineering “Giant Cavendish tissue-culture variants” (GCTCVs) that exhibit robust resistance to Fusarium wilt TR4. These tissue-culture-derived variants are not merely “off-types” but essential genomic resources for safeguarding global food security.
Using haplotype-resolved assemblies, scientists identified that resistance was caused by a Loss of Heterozygosity (LOH)—specifically, large chromosomal deletions that reduced the gene dosage of certain negative regulators of immunity.
5.1 Genomic Structural Variation: Chromosomal Deletions in GCTCV Lines
Identifying causal mutations in sterile triploids requires high-precision bioinformatic pipelines utilizing BWA for alignment and GATK HaplotypeCaller for variant discovery. Recent research has utilized Whole-Genome Sequencing (WGS) and Variant Allele Ratio (VAR) analysis to map structural variations. In triploid Cavendish genomes, heterozygous sites typically yield a VAR of 0.33 or 0.67. A shift to a VAR of 0.5 indicates an Alteration of Heterozygosity (AOH), while a shift to 0 or 1 represents a complete Loss of Heterozygosity (LOH).
Specific genomic findings have pinpointed recurrent deletions in resistant lines:
- The Chromosome 4 Deletion: In the TC5 variant, a specific ~30 kb deletion (chr04: 44,210,964–44,240,861 bp) was identified on one copy of chromosome 4, validated via split-read analysis.
- The Chromosome 5 Deletions: A much larger structural change—spanning approximately 6 Mb—was identified across FM (Formosana), TC7, GCTCV-217, and GCTCV-119. For the FM variant, the deletion boundaries were precisely mapped from 956,425 to 7,030,674 bp.
Key Differentiators and Technical Validation
- Haplotype-Specific Dosage: Coverage analysis of haplotype-resolved assemblies reveals that FM and TC7 lost the BXJ2-5/Ban05 haplotype, while GCTCV-217 and GCTCV-119 lost the BXJ1-5/Ze05 haplotype. This indicates that total gene dosage, rather than a specific allele, is the driver of resistance.
- Validation Tools: Deletions were confirmed through split-read analysis and collinearity analysis (using MCScanX), showing 74-78% collinear orthologs across deletion regions.
- Universal Mechanism: Similar deletions were observed in the TR4-resistant cultivar Nongke No. 1 (NK) compared to the susceptible Baxi (BX), proving this genomic mechanism is a universal feature of TR4 resistance across different breeding programs.
While large-scale structural deletions (~6 Mb) are typically lethal in diploids, the genomic flexibility of triploid Cavendish allows the plant to “shred” susceptibility genes while two functional chromosomal copies compensate for the loss.
The genomic evidence points to three critical insights regarding TR4 resistance:
- Alteration of Heterozygosity (AOH) and Dosage: In triploid Cavendish, heterozygous sites typically show a VAR of 0.33 or 0.67 (a 2:1 ratio). In resistant variants, these ratios shift to 0.5. This “AOH” is the mathematical proof of a single-copy loss, reducing the gene dosage from three copies to two.
- Haplotype-Independent Resistance: The “smoking gun” for the dosage theory is haplotype identity. Deletions in FM and TC7 occurred on the BXJ2-5/Ban05 copy of Chromosome 5, whereas deletions in GCTCV-217 and GCTCV-119 occurred on the BXJ1-5/Ze05 copy. Because resistance was achieved regardless of which copy was lost, it proves that the total number of gene copies (dosage), not the specific allele, governs the immune response.
- Recurrent Selection & External Validation: This phenomenon is not isolated to TBRI variants. VAR analysis of independent data reveals that the resistant “Nongke No. 1” (NK) cultivar also harbors a similar deletion on Chromosome 5. This recurrent selection across different breeding programs confirms that reducing specific gene dosages is a primary evolutionary pathway to TR4 survival.
5.2 Analysis of Resistance Mechanisms in GCTCV Somaclonal Variants
The Taiwan Banana Research Institute (TBRI) screening program utilizes the stressful environment of in vitro culture—defined by high concentrations of plant growth regulators (PGRs) and repeated subculturing—to induce the genomic instability required for TR4 resistance. Through the analysis of haplotype-resolved assemblies (referencing the Li et al. and Huang et al. 2023/2024 genomes), we have determined that these variants utilize chromosomal deletions to achieve immune priming via dosage optimization.
- GCTCV-215-1-1A (Tai-Chiao No. 5 / TC5): Exhibits high commercial value and stable resistance. Genomic mapping (WGS) reveals a ~30 kb deletion on one copy of Chromosome 4. This structural variant results in the loss of one out of three chromosomal copies, specifically reducing the expression of the negative regulator MaACA7.
- GCTCV-218-1 (Formosana / FM): Carries a ~6 Mb deletion on one copy of Chromosome 5. Analysis confirms the loss of the BXJ2-5/Ban05 haplotype.
- GCTCV-105-1-1 (Tai-Chiao No. 7 / TC7): Shares the ~6 Mb Chromosome 5 deletion and the loss of the BXJ2-5/Ban05 haplotype with FM, indicating a recurrent genomic solution.
- GCTCV-119 & GCTCV-217: Contrast with FM by carrying larger deletions; GCTCV-119 exhibits nearly an entire chromosome 5 loss. Crucially, these variants lost the BXJ1-5/Ze05 haplotype.
The fact that different haplotypes were lost across these variants to achieve the same resistance phenotype is the “smoking gun” evidence that gene dosage, rather than a specific allelic identity, is the primary driver of resistance.
Mechanism of Resistance in GCTCVs
| Cultivars | Genomic Event | Target Gene & Homology | Immune Result |
|---|---|---|---|
| TC5 | ~30 kb deletion on Chromosome 4 | MaACA7 (ER-localized; homolog to AtACA1/2/7) | Reduced dosage of this calcium pump removes “brakes” on the immune system. |
| FM, TC7, GCTCV-217, GCTCV-119 | ~6 Mb deletion on Chromosome 5 | MaACA8 (Vacuole-localized; homolog to AtACA4/11) | Significant induction of SA marker genes: MaWRKY40 and MaDLO1. |
These variants succeed because the deletions target Susceptibility (S) genes. By reducing the dosage of these negative regulators of immunity, the plants exhibit a much stronger Salicylic Acid (SA) defense response upon fungal attack. Because the pathogen relies on these host components to suppress the plant’s defenses, the resistance gained through these deletions is expected to be more durable than traditional R-gene resistance.
5.3 The Molecular Basis of Resistance: S Gene Dosage and Calcium Pumps
The shift from R gene-mediated resistance to Susceptibility (S) gene modification represents a more durable strategy for crop protection. Reducing the dosage of negative regulators exploits the plant’s own “genomic flexibility” to achieve broad-spectrum protection. In the identified deletion regions of GCTCV lines, the critical candidates are two members of the autoinhibited Ca2+-ATPase (ACA) calcium pump family.
- MaACA7 (Chromosome 4): Homologous to ER-localized pumps (AtACA1/2/7). Its reduction in TC5 leads to a ~30% decrease in transcript levels.
- MaACA8 (Chromosome 5): Homologous to vacuole-localized pumps (AtACA4/11). This gene is consistently lost in the 6 Mb deletions of FM and TC7, resulting in a similar ~30% dosage-dependent reduction in expression.
“Our findings underscore the importance of structural variation and dosage of susceptibility genes in shaping robust disease resistance in triploid bananas. This work provides critical genomic insights for selecting or engineering TR4-resistant banana varieties.”
The downregulation of these calcium pumps in different compartments (ER and Vacuole) essentially removes the “brakes” on the plant’s immune system, maintaining a state of readiness that attenuates fungal colonization.
Understanding “Susceptibility Genes” (S Genes)
In molecular plant pathology, we distinguish between Resistance (R) genes and Susceptibility (S) genes. While R-genes act as specialized sensors for pathogen detection, S-genes are native plant genes that the pathogen either exploits or that the plant uses to maintain physiological homeostasis by “braking” the immune system. In the context of TR4, specific calcium pumps act as endogenous negative regulators. These are not genes hijacked by the fungus, but rather internal regulators whose natural presence prevents the plant from mounting a sufficiently aggressive defense. Consequently, “losing” an S-gene via mutation can unlock a latent immune potential that is often more robust than the gain of a single R-gene.
| Feature | R Genes (Race-Specific Defense) | S Genes (Negative Regulators) |
|---|---|---|
| Mechanism of Action | Specialized sensors that recognize specific pathogen effectors (the “lock and key” model). | Endogenous genes that suppress immune signaling or provide metabolic support to the pathogen. |
| Durability of Resistance | Low; pathogens rapidly evolve to evade specific recognition. | High; loss-of-function provides broad-spectrum, durable defense that is difficult to overcome. |
| Genetic Strategy | Transgenic “gain-of-function” (introducing new DNA). | Genome editing “loss-of-function” (silencing or deleting existing DNA). |
By analyzing resistant variants, researchers identified that the secret to TR4 survival lay in the deletion of these genetic “brakes,” allowing the plant’s defense system to remain primed for attack.
5.4 Implementation Framework: Identifying S-Genes in Other Clonal Crops
The genomic framework discovered in Musa serves as a blueprint for other polyploid, clonally propagated crops (e.g., olives, potatoes, strawberries) where conventional breeding is stymied.
- Induction of Variation: Leverage tissue culture stress, chemical mutagens, or gamma irradiation to generate a library of structural variations (SVs).
- Genomic Mapping via VAR: Utilize Variant Allele Ratio (VAR) and read coverage analysis to detect Loss of Heterozygosity (LOH). In triploid genomes, a shifted VAR of 0.5 or 0/1 is the primary diagnostic for one-copy loss.
- Candidate Screening: Prioritize orthologous S-genes, specifically calcium-binding proteins or pumps, referencing parallels such as TaHRC (wheat) and MLO (barley).
- Functional Validation: Use transient VIGS systems to confirm immune priming through SA-pathway markers (WRKY and DLO homologs).
Broad-spectrum, durable resistance is best achieved by combining this S-gene “dosage optimization” with existing R-gene strategies.
5.5 Defense Signaling: The Role of Salicylic Acid (SA) in Somaclonal Immunity
The primary mediator of this enhanced immunity is Salicylic Acid (SA), central to plant defense against hemibiotrophic pathogens. The reduction in MaACA dosage directly impacts the SA pathway.
Experimental evidence from PNAS studies validates this mechanism:
- Induction of SA Markers: Resistant variants show significantly stronger induction of SA-responsive marker genes, specifically MaWRKY40 and MaDLO1, upon exposure to TR4.
- VIGS Validation: Using Virus-Induced Gene Silencing (VIGS) to knock down MaACA7 or MaACA8 in susceptible plants resulted in a spontaneous upregulation of SA defense genes. Silencing efficiency reached 77% for MaACA7 and 70% for MaACA8.
- Phenotypic Protection: Silenced plants exhibited markedly reduced rhizome discoloration, confirming that lower dosage of these calcium pumps is sufficient to enhance resistance.
The stronger SA response in variants like Formosana (FM) and TC7 provides a decisive advantage. By maintaining a sensitive SA-mediated defense trigger, these somaclonal variants effectively attenuate the progression of Fusarium, safeguarding the plant in infested soils.
6. Conclusion: Harnessing the Power of Plant Stress
Somaclonal variation is an inevitable byproduct of the stressful in vitro environment. Driven by media imbalances and the radical reprogramming of cells during dedifferentiation, it represents the plant genome’s attempt to reorganize under pressure.

Technical Comparison: Structural Deletions vs. Targeted Modification
While somaclonal deletions provided the initial proof-of-concept, the future of the industry lies in precision modifications that preserve agronomic quality and fruit yield.
| Feature | Large Deletion (Somaclonal) | Targeted Silencing/Editing (VIGS/CRISPR) |
|---|---|---|
| Precision | Low; spans megabases/hundreds of genes | High; targets specific S-gene sequences |
| Non-Target Impact | High; risk of “hitchhiker” trait loss | Minimal; preserves commercial characteristics |
| Agronomic Stability | Variable; requires extensive field testing | High; maintains “Pei-Chiao-like” quality |
| Regulatory Status | Currently lower complexity (natural mutation) | Higher; evolving GEC/GMO definitions |
| Technical Feasibility | Driven by screening volume and time | Requires established transformation protocols |
Validation of these targets was achieved via CMV-based Virus-Induced Gene Silencing (VIGS). Utilizing Agrobacterium-mediated delivery into the rhizome–pseudostem junction, researchers achieved 70-77% silencing efficiency for MaACA7 and MaACA8. This transient knockdown replicated the resistance phenotype of the somaclonal variants, confirming that these calcium pumps are the causal agents within the large deletions.
Silencing the Pumps
To validate the role of ACAs, researchers employed Virus-Induced Gene Silencing (VIGS). Using Agrobacterium to deliver viral constructs, they targeted the rhizome–pseudostem junction of susceptible “Pei-Chiao” (PC) bananas to manually “knock down” MaACA7 and MaACA8 expression.
The experimental results confirmed that reducing the dosage of these pumps transformed a susceptible plant into a resistant one:
- Before Silencing (Susceptible State):
- High expression of ACA calcium pumps acting as strong immune brakes.
- Muted Salicylic Acid (SA) response upon TR4 contact.
- Phenotype: Severe wilting and deep internal rhizome discoloration (necrosis) following TR4 exposure.
- After Silencing (Resistant State):
- Transcript levels reduced by ~30% (mimicking natural GCTCV deletions).
- Spontaneous activation of MaWRKY40 and MaDLO1 even in the absence of the pathogen.
- Phenotype: Attenuated disease symptoms and significantly reduced rhizome discoloration.
3 Essential Insights for a Learner
- Stress as an Engine for Diversity: The stressful conditions of tissue culture drive the creation of new traits that traditional breeding cannot reach, especially in sterile or complex polyploid genomes.
- S-Genes vs. R-Genes: The most critical paradigm shift in modern breeding is moving beyond finding new Resistance (R) genes to strategically deleting or silencing Susceptibility (S) genes (like the ACA calcium pumps) to provide durable, broad-spectrum resistance.
- From Random to Precise: By studying the mechanisms of somaclonal variation, we can move from waiting for “lucky” laboratory accidents to using precise tools like CRISPR to replicate these beneficial deletions without the burden of unwanted “off-type” mutations.

Strategic Takeaways:
- Transition to Dosage Optimization: Success in sterile triploids depends on fine-tuning the expression of susceptibility genes rather than the binary presence/absence of R-genes.
- Haplotype-Resolved Assemblies as a Prerequisite: Implementing this strategy requires high-quality, haplotype-resolved reference genomes to identify which specific chromosomal copies are being modulated.
- Immune Priming without Penalty: The goal is to achieve pre-activated defenses (priming) by reducing S-gene TPM counts just enough to stop Foc TR4 while maintaining elite fruit quality.
- S-Gene Primacy for Durability: Modulating calcium signaling negative regulators (like ACAs) offers more durable, broad-spectrum defense than race-specific interventions.

Image Summary
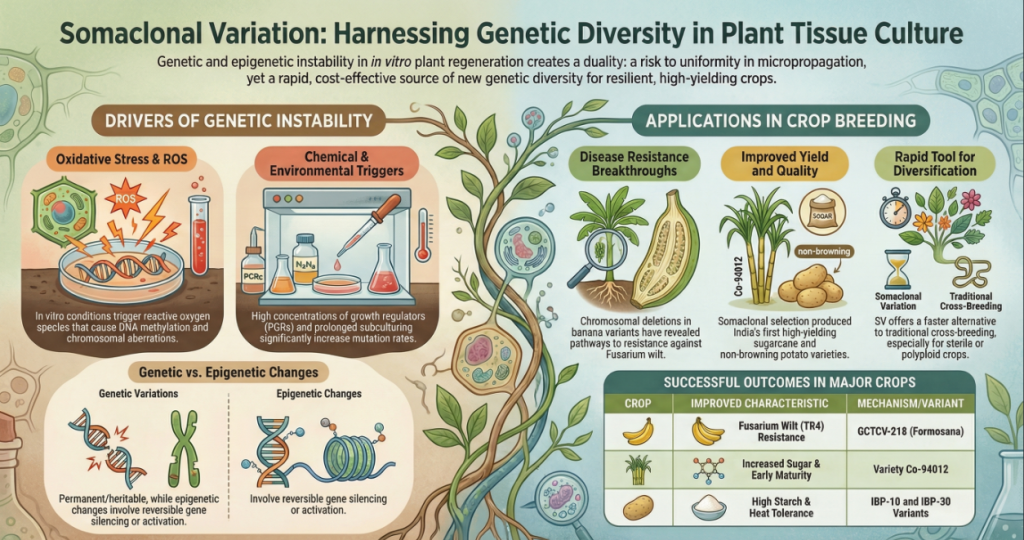


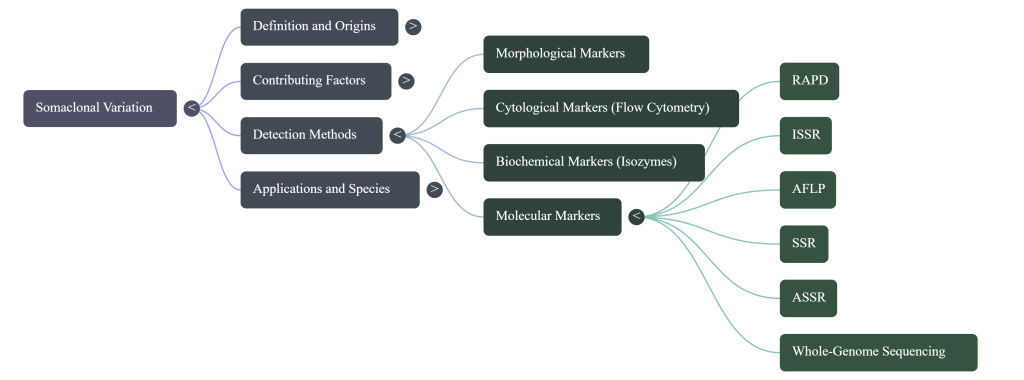

Questions/Answers
1. What causes somaclonal variation during the plant tissue culture process?
Somaclonal variation—defined as genetic and epigenetic variation occurring among plants regenerated from in vitro cell or tissue culture—is caused by a complex interaction of factors related to the starting material, the culture environment, and the molecular mechanisms triggered during the micropropagation process.
The primary causes of somaclonal variation include:
1. Explant Source and Genotype
- Tissue Type: Highly differentiated tissues such as roots, leaves, and stems are more susceptible to genetic alterations during culture than meristematic tissues like shoot tips or axillary buds. Meristematic cells are generally more genetically stable and less prone to epigenetic changes.
- Pre-existing Variation: Somaclonal variation can stem from somatic mutations or chimeras already present in the donor plant’s tissues.
- Genotype: Different genomes respond differently to the stresses of tissue culture, with some cultivars exhibiting higher inherent instability than others.
- Ploidy Level: Polyploid species often show higher rates of somaclonal variation (specifically chromosomal instability) because their multiple chromosome sets can buffer the effects of gross genetic alterations.
2. Mode of Regeneration
The pathway by which a plant is regenerated significantly impacts its genetic stability:
- Indirect Organogenesis: Processes that involve a callus phase (disorganized growth) carry a high risk of genetic instability due to uncontrolled cell divisions and dedifferentiation.
- Direct Organogenesis: Bypassing the callus stage and forming structures directly from the explant minimizes instability and helps maintain genetic fidelity.
3. Media Components and Chemical Factors
- Plant Growth Regulators (PGRs): High concentrations or unbalanced ratios of auxins and cytokinins are powerful agents of variation.
- 2,4-D (Dichlorophenoxyacetic acid): This synthetic auxin is frequently associated with polyploidy, endoreduplication, and DNA methylation changes.
- Cytokinins: High levels of BAP (Benzylaminopurine) or BAP have been linked to increased chromosomal counts and mutations in specific cultivars.
- Imbalances: Nutrient or chemical imbalances in the artificial media can cause cellular stress, disrupting normal cell cycle controls.
4. Culture Duration and Subculturing
- Number of Subcultures: The risk of genetic variation increases with each successive subculture. Mutations and chromosomal aberrations tend to accumulate sequentially over multiple cycles.
- Culture Age: Prolonged duration in culture, particularly for cell suspensions and callus cultures, significantly heightens the frequency of somaclonal variants.
5. Environmental Stress
- Oxidative Stress: The artificial in vitro environment—including wounding during explant preparation, exposure to sterilants, lighting conditions, and high humidity—induces oxidative stress.
- Reactive Oxygen Species (ROS): This stress leads to the overproduction of free radicals (ROS), which can damage DNA, cause chromosome strand breaks, and alter DNA methylation patterns.
6. Molecular and Cellular Mechanisms
- Genetic Changes: These include changes in chromosome number (aneuploidy or polyploidy), structural rearrangements (deletions, inversions, translocations, duplications), and point mutations.
- Transposable Element Activation: “Genomic shock” from the culture process can activate dormant transposable elements (transposons), which move within the genome and disrupt gene function or linearity.
- Epigenetic Modifications: Changes in DNA methylation (hypermethylation or hypomethylation) and histone modifications can profoundly alter gene expression without changing the underlying DNA sequence. These epigenetic markers can sometimes be passed to progeny, though they are often reversible.
2. Can somaclonal variants be used to improve other vegetable crops?
Yes, somaclonal variation is a powerful and widely used tool for the genetic improvement of various vegetable crops. It offers a rapid alternative to traditional breeding, especially for crops that are difficult to breed due to high heterozygosity, polyploidy, or vegetative propagation.
The following vegetable crops have been successfully improved through the selection of somaclonal variants:
Potato (Solanum tuberosum)
Potato is frequently cited as one of the most important vegetable crops improved via this method.
- Disease Resistance: Variants have been developed with stable resistance to Late blight (Phytophthora infestans) and Early blight (Alternaria solani).
- Abiotic Stress Tolerance: Researchers have identified potato somaclones with improved tolerance to salinity, drought, heat, and cadmium.
- Nutritional and Industrial Quality: The variant “Ros 119” was selected for its outstanding starch accumulation, showing significant increases in tuber weight and starch content. Other variants offer non-browning characteristics or higher antioxidant capacity.
Tomato (Solanum lycopersicum)
Tomato crops have been improved for several key agronomic traits:
- Yield: Selection has led to high-yielding varieties with an increased number of fruits per plant.
- Disease Resistance: Somaclonal variants with improved resistance to Fusarium wilt have been identified and used as parents for further varietal production.
- Fruit Quality: Variants have shown improved fruit size, shape, and solid content.
Carrot (Daucus carota)
- Stress Resistance: Somaclonal variation has produced carrot lines with enhanced drought resistance.
- Disease Resistance: Variants have been selected for resistance to leaf spot (Alternaria dauci) and cavity spot (Pythium violae).
Other Vegetable and Root Crops
- Eggplant (Solanum melongena): Somaclones have been developed with significantly improved salt stress tolerance, exhibiting higher dry weight and fruit weight under saline conditions.
- Cucumber (Cucumis sativus): Variants have been selected for determinate growth habits, increased main shoot length, and a higher number of lateral shoots.
- Garlic (Allium sativum): Selection has yielded variants with consistently higher bulb yields and resistance to pathogenic fungi such as Sclerotium cepivorum.
- Pea (Pisum sativum): Somaclonal variants have demonstrated improved resistance to root rot caused by Fusarium solani.
- Celery (Apium graveolens): Research has produced varieties resistant to Fusarium yellows and multiple-resistant somaclones exhibiting both disease and insect resistance.
- Sweet Potato (Ipomoea batatas): Improvements include higher salinity tolerance and more desirable, stable skin color.
Strategic Advantages for Vegetable Breeding
Somaclonal variation is particularly valuable because it allows for the development of new traits within the best available elite cultivars without needing extensive back-crossing. It can also expand the narrow genetic base of self-pollinated and vegetatively propagated vegetables. Furthermore, unlike genetically modified (GM) crops, somaclonal variants generally face fewer social, ethical, and regulatory obstacles, making them a more accessible tool for crop improvement.
3. How does oxidative stress trigger DNA damage in tissue culture?
Oxidative stress triggers DNA damage during tissue culture through a process initiated by an oxidative burst, which leads to the overproduction of Reactive Oxygen Species (ROS). These ROS, which include free radicals such as superoxide anions, hydrogen peroxide, hydroxyl, peroxyl, and alkoxyl radicals, inflict various forms of molecular and structural damage on plant DNA.
The mechanisms by which oxidative stress causes DNA damage include:
1. Direct Structural Damage
The overproduction of ROS leads to significant physical disruptions in the genome. Specifically, oxidative stress is responsible for:
- Chromosome strand breaks: ROS can directly cause breakages in DNA strands.
- Chromosomal rearrangements: These breaks can lead to structural aberrations, including deletions, duplications, inversions, and translocations.
- Aneuploidy and Polyploidy: Stress can disrupt normal cell cycle controls, leading to numerical chromosomal changes such as the loss or gain of entire chromosomes.
2. Genetic Mutations
Oxidative stress induces permanent changes at the nucleotide level, which can alter the function of genes. These include:
- Point mutations: ROS cause single-nucleotide substitutions within the DNA sequence.
- Deletions and Substitutions: Small-scale deletions or substitutions of DNA bases frequently occur as a result of ROS activity.
3. Epigenetic Modifications
Beyond direct sequence changes, oxidative stress triggers epigenetic alterations that modify how genes are expressed:
- DNA Methylation Changes: ROS are known to cause both hyper-methylation and hypo-methylation of DNA. These changes in methylation patterns can activate previously silent transposable elements or silence essential genes.
- Histone Modifications: Stress can alter chemical marks on histone proteins, further impacting chromatin structure and transcriptional activity.
Triggers of Oxidative Stress in Culture
The artificial in vitro environment provides numerous stressors that induce this oxidative damage, including:
- Explant preparation: Wounding during excision and exposure to harsh sterilizing agents.
- Media components: Imbalances in nutrients, pH, or high concentrations of plant growth regulators (particularly synthetic auxins like 2,4-D and cytokinins).
- Environmental factors: Exposure to high light intensity, high humidity, and temperature fluctuations.
The cumulative effect of these ROS-mediated molecular disruptions is a primary driver of the genetic and epigenetic instability known as somaclonal variation.
4. Can somaclonal variation improve abiotic stress tolerance in vegetables?
Somaclonal variation has proven to be a valuable tool for enhancing abiotic stress tolerance in various vegetable crops, offering a faster alternative to traditional breeding methods. By applying specific in vitro selection pressures (such as adding salt or chemicals to the culture medium), researchers can identify and recover rare variants that possess superior resilience to environmental challenges.
Key examples of vegetable and root crops improved for abiotic stress tolerance through this process include:
Potato (Solanum tuberosum)
Potato is one of the most frequently improved vegetable crops via somaclonal variation.
- Salinity and Drought: Researchers have successfully used in vitro screening to produce potato lines with improved tolerance to both salinity and drought.
- Heat and Heavy Metals: Somaclonal variants have been selected for heat tolerance and resistance to toxic levels of cadmium.
Eggplant / Brinjal (Solanum melongena)
Somaclonal variants of eggplant have shown significant improvements in handling saline environments.
- Salt Tolerance: Callus lines selected on medium containing up to 120 mM NaCl yielded salt-tolerant plants.
- Performance: These salt-tolerant somaclones exhibited higher dry weight, better water retention, and increased fruit weight compared to their original parent plants.
Carrot (Daucus carota)
Somaclonal variation has been utilized to address water scarcity in carrot production.
- Drought Resistance: Several studies have successfully isolated and characterized carrot somaclonal lines with enhanced drought resistance.
Sweet Potato (Ipomoea batatas)
- Salinity Tolerance: Researchers have identified sweet potato somaclonal variants with a high degree of tolerance to salinity.
Tomato (Solanum lycopersicum)
- Osmotic and Abiotic Stress: While often used for disease resistance, biotechnological approaches in tomato breeding—including somaclonal variation—are actively used to alleviate abiotic stresses. Studies have identified specific signatures of osmotic stress-responsive genes in somaclonal variant cells.
General Benefits for Abiotic Improvement
- Rapid Diversity: It provides a source of genetic variability relatively quickly without the need for sophisticated transgenic technologies.
- New Pathways: The process can generate novel pathways of resistance that may not be accessible through conventional cross-breeding.
- Efficiency: Screening for desirable traits like stress tolerance in vitro requires less space and time than large-scale field trials for seedlings of perennial or long-cycle crops.
5. Can gene editing replicate the resistance found in somaclonal variants?
Recent research suggests that gene editing can indeed be used to replicate the resistance mechanisms discovered in somaclonal variants, providing a more precise and predictable alternative to the random nature of the tissue culture process. While somaclonal variation is a valuable source of novel traits, its random genetic alterations often result in undesirable side effects; gene editing allows for the induction of specific beneficial mutations while preserving the agronomic integrity of the parent cultivar.
Key ways gene editing can replicate or enhance somaclonal resistance include:
1. Targeted Replication of Identified Mutations
Once the molecular basis for resistance in a somaclonal variant is identified through genomic analysis, gene editing tools like CRISPR/Cas9 can induce the same specific change in susceptible cultivars.
- Banana (TR4 Resistance): Researchers found that certain somaclonal variants resistant to Fusarium wilt (TR4) possessed large chromosomal deletions that reduced the dosage of susceptibility (S) genes, specifically the ACA gene family. This discovery provides a “genomic framework” for using targeted gene editing to precisely modify ACA gene dosage in other susceptible banana varieties.
- S-Gene Disruption: Broad-spectrum resistance is often linked to the loss of function in susceptibility genes. For example, mlo-based resistance to powdery mildew, originally found as a mutation in barley, has been successfully replicated in other crops through gene editing.
2. Precision vs. Random Variation
The primary advantage of gene editing over somaclonal variation is its precision.
- Avoiding Negative Pleiotropy: Somaclonal variation is random and frequently generates “inferior lines” or “off-types” alongside resistant variants. In bananas, for instance, early TR4-resistant variants often had poor yield or fruit quality. Gene editing can replicate the resistance mutation while “minimizing unintended effects on other loci” to preserve commercial traits.
- Clonal Fidelity: CRISPR/Cas9 supports the maintenance of clonal fidelity by offering controlled mutations, whereas somaclonal variation is spontaneous, uncontrolled, and unpredictable.
3. Strategic Limitations and Challenges
Despite the potential of gene editing, several factors mean that the selection of somaclonal variants remains a relevant breeding tool:
- Genetic Knowledge Requirement: Gene editing requires prior knowledge of the specific gene responsible for a trait. Somaclonal variation is often more useful for novel traits where the underlying genetic basis is not yet identified or isolated.
- Technological Barriers: The application of genome editing across many cultivars (such as different banana varieties) is still constrained by variable regeneration and transformation efficiencies.
- Regulatory and Social Factors: In some contexts, somaclonal variants are more accessible because they generally face fewer social, ethical, and regulatory obstacles than genetically modified (GM) or gene-edited crops.
In summary, while somaclonal variation serves as an “exploratory” tool to discover new resistance pathways, gene editing is the “precision” tool that can replicate those discoveries to produce stable, high-quality, resistant cultivars.
6. Explain the role of ACA genes in plant immunity.
ACA genes (autoinhibited Ca2+-ATPase) encode calcium pumps that act as negative regulators of plant immunity, specifically suppressing the salicylic acid (SA)-mediated defense response. Because their presence and normal function actively inhibit immune pathways, they are classified as susceptibility (S) genes.
The role of ACA genes in plant immunity includes the following key aspects:
1. Negative Regulation of Defense Signaling
- Suppression of Salicylic Acid (SA): ACA genes serve to keep the plant’s immune system in check by suppressing the expression of SA-responsive genes.
- Upregulation upon Downregulation: When ACA genes are downregulated or deleted, marker genes for SA-mediated defense—such as MaWRKY40 and MaDLO1—are significantly induced, even in the absence of a pathogen. This “primes” the plant’s immune system for a faster and stronger response during an actual infection.
2. Calcium Signaling and Localization
- Calcium Efflux Pumps: These genes encode pumps that move calcium ions (Ca2+) across membranes to regulate cellular signaling.
- Cellular Compartmentalization: Different members of the ACA family are localized to specific organelles. For example, MaACA7 (and its Arabidopsis homologs AtACA1, 2, and 7) is localized in the endoplasmic reticulum (ER), while MaACA8 (homologous to AtACA4 and 11) is localized in the vacuole. Disruption of these pumps in either location has been shown to activate defense-related programmed cell death pathways.
3. Impact on Pathogen Resistance
- Fungal Resistance (Banana): In Cavendish banana somaclonal variants, large chromosomal deletions that reduce the copy number (dosage) of MaACA7 or MaACA8 lead to stable resistance against Fusarium wilt (TR4). Reducing the expression of these genes attenuates disease symptoms by allowing the plant to maintain a more robust SA-mediated defense.
- Broad-Spectrum Immunity: Beyond fungi, the knockdown or knockout of ACA genes in model plants like Arabidopsis, rice, and Nicotiana benthamiana has been shown to enhance resistance to a wide range of bacterial and viral pathogens.
4. Breeding and Engineering Potential
Because ACA genes function as susceptibility genes, they are primary targets for gene editing (e.g., CRISPR/Cas9) to enhance disease resistance. By precisely modifying the dosage or expression of these negative regulators, researchers can replicate the robust resistance found in somaclonal variants while preserving the commercial and agronomic integrity of the crop.
References
Adly, W. M. R. M., Niedbała, G., EL-Denary, M. E., Mohamed, M. A., Piekutowska, M., Wojciechowski, T., Abd El-Salam, E. S. T., & Fouad, A. S. (2023). Somaclonal Variation for Genetic Improvement of Starch Accumulation in Potato (Solanum tuberosum) Tubers. Plants, 12(2). https://doi.org/10.3390/plants12020232
Anil, V. S., Bennur, S., & Lobo, S. (2018). Somaclonal variations for crop improvement: Selection for disease resistant variants in vitro. In Plant Science Today (Vol. 5, Number 2, pp. 44–54). Horizon e-Publishing Group. https://doi.org/10.14719/pst.2018.5.2.382
Bairu, M. W., Aremu, A. O., & van Staden, J. (2011a). Somaclonal variation in plants: Causes and detection methods. Plant Growth Regulation, 63(2), 147–173. https://doi.org/10.1007/s10725-010-9554-x
Bairu, M. W., Aremu, A. O., & van Staden, J. (2011b). Somaclonal variation in plants: Causes and detection methods. Plant Growth Regulation, 63(2), 147–173. https://doi.org/10.1007/s10725-010-9554-x
Bulbarela-Marini, J. E., Gómez-Merino, F. C., Galindo-Tovar, M. E., Pastelín-Solano, M. C., Murguía-González, J., Núñez-Pastrana, R., & Castañeda-Castro, O. (2023). Ratio of Somaclonal Variation and the Phytohormonal Content of Citrus × latifolia in Three In Vitro Culture Systems. Journal of Plant Growth Regulation, 42(6), 3356–3364. https://doi.org/10.1007/s00344-022-10796-x
Das, S., Datta, S., Kundu, M., Bhattacharjee, P., Wangchu, L., Ningombam, L., Singh, S., Nandita, K., Tani, M., Das, S., & Shil, S. (2025). From induction to innovation: investigating somaclonal variation induced by tissue culture and its role in advancing fruit crop improvement. In Plant Cell, Tissue and Organ Culture (Vol. 161, Number 1). Springer Science and Business Media B.V. https://doi.org/10.1007/s11240-025-03044-z
Dewi Ibrahim, M. S., Hartati, R. S., Rubiyo, R., Reflinur, R., Purwito, A., & Sudarsono, S. (2024). Exploring indirect somatic embryogenesis and somaclonal variation for propagation of three Coffea arabica L. cultivars. Chilean Journal of Agricultural Research, 84(1), 15–27. https://doi.org/10.4067/S0718-58392024000100015
Duta-Cornescu, G., Constantin, N., Pojoga, D. M., Nicuta, D., & Simon-Gruita, A. (2023). Somaclonal Variation—Advantage or Disadvantage in Micropropagation of the Medicinal Plants. In International Journal of Molecular Sciences (Vol. 24, Number 1). MDPI. https://doi.org/10.3390/ijms24010838
Dyduch-Siemińska, M., & Gawroński, J. (2025). In Vitro Regeneration of Stevia rebaudiana Bertoni Using Somaclonal Variation as a Tool for Genetic Diversification. Genes, 16(10). https://doi.org/10.3390/genes16101203
Ferreira, M. dos S., Rebouças, T. A., Rocha, A. de J., Oliveira, W. D. dos S., Santos, A. C. L. S. dos, Jesus, J. P. F. L. de, Ramos, A. P. de S., Ferreira, C. F., Santos-Serejo, J. A. dos, Haddad, F., & Amorim, E. P. (2024). Selection and Characterization of Somaclonal Variants of Prata Banana (AAB) Resistant to Fusarium Wilt. Agronomy, 14(8). https://doi.org/10.3390/agronomy14081740
Ferreira, M. dos S., Rocha, A. de J., Nascimento, F. dos S., Oliveira, W. D. dos S., Soares, J. M. da S., Rebouças, T. A., Morais Lino, L. S., Haddad, F., Ferreira, C. F., Santos-Serejo, J. A. dos, Fernández, J. S., & Amorim, E. P. (2023). The Role of Somaclonal Variation in Plant Genetic Improvement: A Systematic Review. In Agronomy (Vol. 13, Number 3). MDPI. https://doi.org/10.3390/agronomy13030730
Gu, X., Feng, W., Zhou, Z., Li, W., Zhang, H., Wei, L., Cao, Y., & Hu, S. (2026). Cellular and molecular characterizations of a thick secondary cell wall somaclonal variant in Dendrocalamus farinosus. Industrial Crops and Products, 241. https://doi.org/10.1016/j.indcrop.2026.122777
Hou, B. H., Tsai, Y. H., Chiang, M. H., Tsao, S. M., Huang, S. H., Chao, C. P., & Chen, H. M. (2022). Cultivar-specific markers, mutations, and chimerisim of Cavendish banana somaclonal variants resistant to Fusarium oxysporum f. sp. cubense tropical race 4. BMC Genomics, 23(1). https://doi.org/10.1186/s12864-022-08692-5
Kaeppler, S. M., Kaeppler, H. F., & Rhee, Y. (2000). Epigenetic aspects of somaclonal variation in plants. In Plant Molecular Biology (Vol. 43).
Karp, A. (1995). Somaclonal variation as a tool for crop improvement. In Euphytica (Vol. 85).
Krishna, H., Alizadeh, M., Singh, D., Singh, U., Chauhan, N., Eftekhari, M., & Sadh, R. K. (2016). Somaclonal variations and their applications in horticultural crops improvement. In 3 Biotech (Vol. 6, Number 1, pp. 1–18). Springer Verlag. https://doi.org/10.1007/s13205-016-0389-7
Kumar Maurya, P., & Bahadur, V. (2022). Somoclones and Somaclonal Variants: A Review. Environment and Ecology, 40(4C), 2532–2539.
Leva, A. R., Petruccelli, R., & Rinaldi, L. M. R. (2012a). Somaclonal Variation in Tissue Culture: A Case Study with Olive. In Recent Advances in Plant in vitro Culture. InTech. https://doi.org/10.5772/50367
Leva, A. R., Petruccelli, R., & Rinaldi, L. M. R. (2012b). Somaclonal Variation in Tissue Culture: A Case Study with Olive. In Recent Advances in Plant in vitro Culture. InTech. https://doi.org/10.5772/50367
López-Hernández, F., & Cortés, A. J. (2022). Whole Transcriptome Sequencing Unveils the Genomic Determinants of Putative Somaclonal Variation in Mint (Mentha L.). International Journal of Molecular Sciences, 23(10). https://doi.org/10.3390/ijms23105291
Majumder, S., Igamberdiev, A. U., & Debnath, S. C. (2025). Somaclonal Variation and Clonal Fidelity in Commercial Micropropagation: Challenges and Perspectives. In Agronomy (Vol. 15, Number 6). Multidisciplinary Digital Publishing Institute (MDPI). https://doi.org/10.3390/agronomy15061489
Manchanda, P., Sharma, D., Kaur, G., Kaur, H., & Vanshika. (2025). Exploring the Significance of Somaclonal Variations in Horticultural Crops. In Molecular Biotechnology (Vol. 67, Number 6, pp. 2185–2203). Springer. https://doi.org/10.1007/s12033-024-01214-6
Martínez-Velasco, I., & Arzate-Fernández, A. M. (2022). ANALYSIS OF THE SOMACLONAL VARIATION IN TWO IN VITRO REGENERATED AGAVE SPECIES † [ANÁLISIS DE LA VARIACIÓN SOMACLONAL EN DOS ESPECIES DE AGAVE REGENERADAS IN VITRO]. In Tropical and Subtropical Agroecosystems (Vol. 25).
Mujib, A., Banerjee, S., & Ghosh, P. (2013). Tissue Culture Induced Variability in some Horticultural Important Ornamentals. Biotechnology, 12(3).
Sianipar, N. F., Assidqi, K., So, I. G., Maulidha, A. R., & Asikin, Y. (2024). SOMACLONAL VARIATIONS INDUCED BY BENZYLAMINOPURINE TO ENHANCE THE FRUIT MORPHOLOGY OF HORN BANANA. Sabrao Journal of Breeding and Genetics, 56(5), 2045–2055. https://doi.org/10.54910/sabrao2024.56.5.27
Singh, P. T., Hou, B. H., Tsai, Y. H., Tzean, Y., Chao, C. P., Zheng, P. X., Lin, Y. C., Shen, W. C., Yeh, H. H., & Chen, H. M. (2025). Chromosomal deletions in banana somaclonal variants reveal negative regulators of immunity underlying Fusarium wilt resistance. Proceedings of the National Academy of Sciences of the United States of America, 122(48). https://doi.org/10.1073/pnas.2511842122
Thipwong, J., Kongton, K., & Samala, S. (2022). Micropropagation and somaclonal variation of Doritis pulcherrima (Lindl.). Plant Biotechnology Reports, 16(4), 401–408. https://doi.org/10.1007/s11816-022-00766-w
Tripathy, S. K., Panda, A., Nayak, P. K., Dash, S., Lenka, D., Mishra, D. R., Kar, R. K., Senapati, N., & Dash, G. B. (2016). Somaclonal variation for genetic improvement in grasspea (Lathyrus sativus L.). Legume Research, 39(3), 329–335. https://doi.org/10.18805/lr.v0iOF.6853