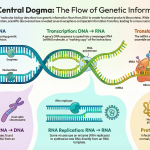
Introduction: From DNA Blueprint to RNA Message
Transcription is the foundational first step in gene expression, the process by which a cell reads its genetic instructions. Think of the cell’s nucleus as a central library containing a massive collection of master recipe books (the DNA). To bake a cake, you wouldn’t take the master book into the kitchen; instead, you’d carefully copy the specific recipe onto a notecard (the RNA). This copied message can then leave the library and be used in the kitchen (the cytoplasm) to assemble the final product. This guide explains the three main stages of this copying process in eukaryotes—Initiation, Elongation, and Termination—in a clear, step-by-step manner for new learners.

The master scribe responsible for reading the DNA blueprint and writing the RNA message is a powerful enzyme called RNA polymerase. Unlike prokaryotes which use a single polymerase for all jobs, eukaryotes employ a sophisticated division of labor with three specialized types. This separation is a significant evolutionary advantage, as it provides a mechanism to independently control the synthesis levels of different classes of RNA, which is crucial for processes like responding to growth-state changes. The distinct identities of these polymerases were famously revealed by experiments using α-amanitin, a deadly toxin from the “Death Cap” mushroom.

Our focus will be on RNA Polymerase II, as it is responsible for transcribing the vast majority of genes that code for proteins. The table below introduces all three key polymerases and their sensitivity to this revealing toxin.

| RNA Polymerase | Location in the Cell | Primary Job (RNA Transcribed) | α-Amanitin Sensitivity |
| Polymerase I | Nucleolus | Most ribosomal RNA (rRNA) | Insensitive |
| Polymerase II | Nucleus | All protein-coding pre-mRNAs | Extremely sensitive |
| Polymerase III | Nucleus | 5S rRNA, transfer RNAs (tRNAs), small nuclear RNAs | Moderately sensitive |
With our key players identified, let’s explore how the cell begins the crucial task of transcribing a gene.
1. Stage One: Initiation – Getting Transcription Started
Unlike its prokaryotic counterpart, eukaryotic RNA polymerase II cannot simply find a gene and start copying it. It requires a dedicated team of helper proteins called basal transcription factors to show it where to begin. This starting point is a specific region of DNA known as the promoter. For a new learner, two promoter sequences are especially important:
1. The TATA box: This is a sequence rich in Adenine (A) and Thymine (T) bases. Its primary benefit is that A-T bonds are held together by fewer hydrogen bonds than G-C bonds. This chemical instability creates a natural “unzip here” point in the otherwise robust DNA double helix, making it an efficient landmark for initiating transcription.
2. The CAAT box: Located further upstream (located further away from the gene’s start site in the 5′ direction) from the TATA box, this is another essential sequence that plays a key role in binding the necessary transcription factors.
The scale of the machinery involved in eukaryotic initiation is staggering. The polymerase itself is a 12-subunit complex, and it relies on a supporting cast of general initiation factors that comprise at least 32 different polypeptides. The initiation process unfolds in a precise, orderly sequence as these components assemble into a massive structure:
1. Recruiting the Team: The first responders are the basal transcription factors (conveniently named with “TFII” for “transcription factor for polymerase II”). These proteins recognize and bind to the promoter sequences, like the TATA box, marking the gene’s starting line.
2. Building the Complex: Like a master locksmith, each transcription factor binds in a precise sequence, creating a docking platform for the next, until the entire preinitiation complex is securely assembled. This colossal structure comprises nearly 60 proteins with a total mass of over 3 million daltons.
3. Calling the Scribe: Only when this preinitiation complex is fully formed does it successfully recruit the master scribe, RNA polymerase II, guiding it to the correct starting position on the gene.

Once this entire preinitiation complex is successfully assembled at the promoter, the polymerase is ready to begin synthesizing the RNA message.



2. Stage Two: Elongation – Building the RNA Strand
With the preinitiation complex securely in place, RNA polymerase II is released from the other transcription factors and begins its primary task. The core function of elongation is straightforward: the polymerase moves along the DNA template, reading the genetic code and synthesizing a complementary pre-mRNA molecule in the 5′ to 3′ direction.
However, the polymerase faces a major challenge that represents a fundamental design trade-off in eukaryotes: the need for massive data compaction versus the need for data access. Eukaryotic DNA is not a naked, easily accessible strand; it is tightly packaged around proteins called histones, forming structures known as nucleosomes. This organization, often compared to thread wrapped around a series of spools, keeps the vast amount of DNA organized but also physically blocks the path of the polymerase.

To solve this problem, the cell uses a protein complex called FACT (which stands for “facilitates chromatin transcription”). The FACT complex acts as a mobile chromatin manager, performing a two-step process to clear the way for the polymerase.
• Clearing the path: As RNA polymerase II moves along the DNA, the FACT complex travels just ahead of it, pulling the histone spools out of the way. This allows the polymerase to access and read the otherwise blocked DNA sequence.
• Restoring the structure: After the polymerase has passed by and transcribed the section of DNA, the FACT complex puts the histones back where they belong. This step reassembles the nucleosome, ensuring the DNA is neatly repackaged and the chromatin structure is maintained.
This elegant process allows transcription to proceed efficiently along the entire length of the gene. Once the polymerase has transcribed the complete genetic message, the cell must employ a unique strategy to signal the end of the process.
3. Stage Three: Termination – Ending the Process
The method for ending transcription in eukaryotes varies between the different polymerases, and the process for RNA polymerase II is particularly unique. Instead of stopping at a precise termination signal located at the very end of the gene, the polymerase continues to transcribe, adding an extra tail of 1,000 to 2,000 nucleotides to the pre-mRNA.
This seemingly inefficient “read-through” is not a quirk, but rather a key feature of an integrated system of mRNA maturation. This long tail is not part of the final genetic message. During the subsequent step of mRNA processing, this extra sequence is removed by cleavage at a specific site. This cut serves a dual purpose: it defines the correct end of the mRNA molecule, where a protective poly-A tail will be added, and it simultaneously signals the termination of transcription, causing the polymerase to detach from the DNA.
This mechanism stands in contrast to the other polymerases, which rely on more direct signals:
• RNA Polymerase I recognizes a specific 18-nucleotide sequence that recruits a termination protein.
• RNA Polymerase III terminates after transcribing a sequence that forms an mRNA hairpin structure, similar to a mechanism seen in prokaryotes.
4. Conclusion: A Tightly Regulated Process
Eukaryotic transcription is a sophisticated and highly controlled process, unfolding in three distinct stages: the assembly of a massive preinitiation complex at the promoter (Initiation), the synthesis of a pre-mRNA strand through organized chromatin (Elongation), and the cleavage-signaled release of the transcript (Termination). It is markedly more complex than transcription in prokaryotes, involving a suite of specialized enzymes and proteins. This intricate, multi-layered control is not complexity for its own sake; it is the molecular engine that allows a single genome to produce the vast diversity of cells—from neurons to muscle to skin—that build a complex organism. The presence of multiple polymerases, armies of transcription factors, and chromatin-modifying complexes like FACT provides the multiple layers of regulation necessary to ensure that the cell transcribes precisely the pre-mRNAs it needs, when it needs them, to carry out the essential functions of life.
Image Summary




References
R.D. Kornberg, The molecular basis of eukaryotic transcription, Proc. Natl. Acad. Sci. U.S.A. 104 (32) 12955-12961, https://doi.org/10.1073/pnas.0704138104 (2007).
Roeder, R.G. 50+ years of eukaryotic transcription: an expanding universe of factors and mechanisms. Nat Struct Mol Biol 26, 783–791 (2019). https://doi.org/10.1038/s41594-019-0287-x
Krishnamurthy, S., & Hampsey, M. (2009). Eukaryotic transcription initiation. Current Biology, 19(4), R153-R156. 10.1016/j.cub.2008.11.052
Sekine SI. Toward structural understanding of eukaryotic transcription elongation. Proc Jpn Acad Ser B Phys Biol Sci. 2025;101(7):414-430. doi: 10.2183/pjab.101.024. PMID: 40721381; PMCID: PMC12462365.
Juanjuan Xie, Domenico Libri, Odil Porrua; Mechanisms of eukaryotic transcription termination at a glance. J Cell Sci 1 January 2023; 136 (1): jcs259873. doi: https://doi.org/10.1242/jcs.259873
https://en.wikipedia.org/wiki/Eukaryotic_transcription
https://bio.libretexts.org/Bookshelves/Introductory_and_General_Biology/General_Biology_(Boundless)/15%3A_Genes_and_Proteins/15.06%3A_Eukaryotic_Transcription_-_Initiation_of_Transcription_in_Eukaryotes