1. Introduction
In the contemporary “Omics” era, Northern Blotting remains a foundational pillar for validating gene expression. While high-throughput deep sequencing and RT-PCR offer speed and sensitivity, they cannot replace the Northern Blot’s unique capability to provide direct visualization of transcript size and complexity. By displaying the abundance and molecular weight of multiple transcripts simultaneously, researchers can observe cellular control over structure and function during differentiation, morphogenesis, and diseased states. This guide provides the technical rigor necessary to leverage Northern Blotting for definitive transcript characterization, particularly when alternative splicing or small RNA processing is the primary research focus.

1.1 Prologue: Why We Track the Messenger
In the intricate library of the living cell, DNA serves as the permanent blueprint—the master archives stored safely in the nuclear vault. However, to actually build and operate an organism, the cell must generate active “work orders.” These messengers are known as mRNA (messenger RNA).
While DNA tells us what a cell can do, RNA tells us what it is actually doing at any given moment. We track these messengers for two primary reasons:
1. Tissue-Specific Expression: To discover which “department” of the body is active. For example, the ob (obese) gene is expressed specifically in white adipose tissue. Northern blotting allows us to see this specific activity, providing clues to the physiological function of the protein it encodes (such as leptin).
2. Gene Regulation: To understand how the cell responds to its environment—be it nutritional shifts (like fasting and refeeding), hormonal signals, or environmental changes like temperature.
Gene Expression: The process by which information from a gene is used to synthesize functional gene products. In Northern blotting, we measure this by detecting the presence, size, and abundance of the RNA “instructions” within a sample.
Before we can read the message, we must first extract it from the crowded city of the cell.
2. Foundations of Northern Blotting: From Concept to Gene Expression
2.1 Historical Development and Context
Developed in 1977 by James Alwine, David Kemp, and George Stark at Stanford University, the Northern Blot (a nomenclature that pays a playful nod to Edwin Southern’s DNA blot) initially used diazobenzyloxymethyl (DBM) paper. Modern protocols have largely transitioned to nylon membranes, which offer higher binding capacity and greater physical robustness.
While the Southern blot analyzes DNA, the Northern blot utilizes denaturing gel electrophoresis to separate RNA by size, followed by a transfer to a sturdy membrane—typically nylon—and subsequent detection via hybridization with a complementary probe. Unlike Southern blotting, the Northern technique must account for the inherent instability of RNA and its propensity for secondary structure, requiring specific denaturants to ensure separation is based solely on molecular weight.
2.2 Core Strategic Purpose
Direct Size Measurement: Northern analysis provides a direct measure of RNA length. By comparing the position of a band to a known ladder (such as the Millennium RNA marker), researchers can estimate the transcript’s coding capacity. This is vital for validating the expected size of the protein product it encodes.
Transcript Variant Detection: This technique is the premier tool for observing alternative splicing and transcript variants. It allows for the simultaneous visualization of multiple RNA species that share sequence identity, such as distinguishing between a mature microRNA (~20 nt) and its larger precursor form (~70 nt).
Relative Abundance: Enables direct comparison of message levels between samples on a single membrane.
Regulatory Analysis: Identifies tissue-specific expression and regulation in response to nutritional, hormonal, or environmental stimuli.
Utilization of Partial Homology Probes: Unlike the strict requirement for perfect primer-template matches in PCR, Northern blotting is exceptionally versatile. It allows researchers to use probes with only partial homology, such as using cDNA from a different species or genomic DNA fragments containing introns, to detect a target gene.

2.3 Comparative Analysis: Strategic Selection
| Feature | Northern Blotting | RT-PCR (Quantitative) | Nuclease Protection Assays |
|---|---|---|---|
| Primary Goal | Assessing transcript size and abundance (amount). | Detecting and quantifying specific RNA levels. | Quantifying message abundance and mapping ends. |
| Sensitivity Level | Standard; generally the lowest of the three. | Highest; can detect a few copies of a message. | High; more sensitive than standard Northern. |
| Throughput | Lower; labor-intensive for many samples. | High; capable of rapid, automated analysis. | Moderate. |
| Key Constraint | Requires significant amounts of intact RNA. | Cannot distinguish variants; prone to misleading data if RNA is degraded. | Does not provide data on full transcript size. |
Expert Observation: While RT-PCR is the choice for low-copy quantification, a Senior Scientist prioritizes Northern Blotting when transcript architecture is in question. Furthermore, while 32P-labeled probes remain the robust “gold standard” for many, the strategic move toward chemiluminescence is driven by safety and waste disposal considerations, though it requires a significant investment in dedicated instrumentation.
2.4 Physical Constraints and Resource Requirements
Northern analysis is a demanding discipline that requires meticulous attention to laboratory conditions and material resources.
• Sample Quantity and Enrichment: A standard lane requires 10–20 µg of total RNA. However, for low-abundance messages, Poly(A) RNA (mRNA) enrichment is recommended. Because mRNA comprises only 0.5–3% of total RNA, using 10 µg of purified Poly(A) RNA is equivalent to loading 300–350 µg of total RNA, offering a massive boost in signal resolution without saturating the membrane.
• Time Investment: Traditional protocols take 2+ days. Modern labs utilize vacuum blotting or alkaline transfer to reduce the transfer step to 1–2 hours, effectively trimming a full day off the workflow.
• Material Integrity (The “Horror Story” of Degradation): RNA is notoriously sensitive to ubiquitous RNases. To a methodologist, degradation is not just a minor issue; it is a dealbreaker. Consider this: a single cleavage in just 20% of 4 kb target molecules will decrease your detectable signal by 20%. Success requires DEPC-treated water, sterilized glassware, and constant glove changes.
3. The Standard Procedural Workflow: A Step-by-Step Analysis
Maintaining an RNase-free environment is paramount. RNA is highly susceptible to degradation by ubiquitous RNases. All reagents must be DEPC-treated or autoclaved, and the use of dedicated equipment and protective gloves is mandatory.
3.1 Safety and Rigor
• Chaotropic Agents: Guanidinium isothiocyanate, used in isolation, is a powerful protein denaturant; handle with care.
• Respiratory Hazards: Formaldehyde used in agarose gels must be handled in a fume hood.
• Neurotoxins: Acrylamide (used for small RNA dPAGE) is a cumulative neurotoxin; safety spectacles and gloves are non-negotiable.
3.2 The Standard Workflow (mRNA Focus)
3.2.1 Step 1: The Departure (RNA Isolation and Quality Control)
To begin our journey, we must liberate the RNA from the complex architecture of the cell. This requires chaotropic agents, such as guanidinium isothiocyanate, which act like a molecular demolition crew—disrupting cellular membranes and denaturing proteins.
However, the environment is perilous for RNA. Our primary “villains” are RNases—enzymes that are designed to shred RNA instantly. They are found everywhere, from tissues to your own fingertips. To protect our travelers, we must use specific inhibitors like DEPC, work in strictly nuclease-free conditions, and wear protective gloves at all times.
Once isolated, we must perform a “Quality Control” check. Using electrophoresis, we look for two distinct, sharp stripes representing the 18S and 28S ribosomal RNA (rRNA). If these appear as a blurry smear, our messengers have been “mugged” by RNases, and the message is lost.
| RNA Type | Purity vs. Abundance | Purpose |
|---|---|---|
| Total RNA | High abundance; includes rRNA and tRNA. | Provides a general snapshot; 10 µg is a standard load. |
| Poly(A)+ mRNA | High purity; only 0.5–3% of total RNA. | Enriches the sample for protein-coding instructions to increase sensitivity. |
Now that we have our pure sample, our molecular travelers are ready for the great race.

3.2.2 Step 2: The Obstacle Course (Denaturing Electrophoresis)
RNA molecules are prone to tangling into “knots” called secondary structures. To sort them fairly by length, we must force them to stay in a straight line. We do this by running them through an “obstacle course” (the gel matrix) under strict denaturing conditions.
Note on Fragility: Unlike the Southern blot for DNA, we cannot use sodium hydroxide as a denaturant here, as it would completely degrade the fragile RNA. Instead, we use agents like formaldehyde or glyoxal to keep the molecules linear.
The Race Dynamics:
1. The Charge: RNA is negatively charged. When an electric current is applied, the molecules are repelled by the negative electrode and sprint toward the positive one.
2. The Matrix: The gel (agarose or polyacrylamide) acts as a molecular sieve. Large molecules get tangled in the pores and move slowly, while smaller ones zip through to finish first.
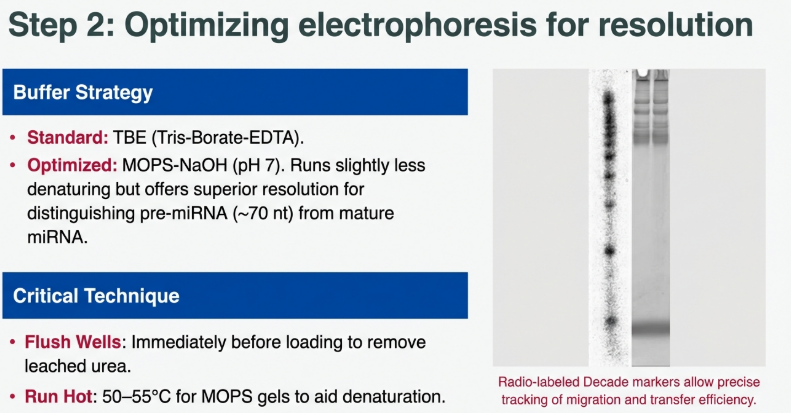
3. The Denaturant: While MOPS-NaOH provides the stable pH buffer environment (carefully chosen to avoid primary amines that might interfere with later steps), formaldehyde acts as the “molecular iron.” It smooths out the RNA “knots” so they run based on length alone.
The molecules are sorted, but they are trapped in a fragile gel “prison”; they need a more permanent home.

3.2.3 Step 3: The Leap of Faith (Transfer to the Membrane)
Gels are fragile and probes cannot easily penetrate them. To work with our sorted RNA, we must move it onto a durable nylon membrane. This “leap” is not a random jump; it is driven by physical forces that pull the RNA out of the gel and onto the membrane surface.
Transfer Methods:
• Capillary Transfer:
◦ The Mechanism: Uses “wicking” action, where a high-salt buffer is drawn upward through the gel by a stack of dry paper towels, carrying the RNA with it.
◦ Pros/Cons: Low-tech and requires no special equipment, but it is very slow (4–18 hours).
• Vacuum Transfer:
◦ The Mechanism: Uses a vacuum pump to actively pull the buffer and RNA through the gel onto the membrane.
◦ Pros/Cons: Fast (1–2 hours) and highly reproducible, but requires specialized vacuum equipment.
On the membrane, our molecules are in position, but they aren’t “home” until they are anchored.

3.2.4 Step 4: The Anchor (Immobilization/Cross-linking)
If we proceeded to the next step now, the RNA would simply wash away. We must “anchor” the molecules to the nylon fibers using a covalent bond.
| Method | Best For | Technical Mechanism |
|---|---|---|
| UV Cross-linking | Large mRNA (>70–100 nt) | Uses UV light to create covalent bonds between RNA bases (principally Uridine) and the membrane. |
| EDC Chemical Cross-linking | Small RNA (<40 nt) | Forms a covalent phosphoramidate bond between the 5′-terminal phosphate of the RNA and the amine groups on the membrane. |
The “So What?”: UV cross-linking is fast but can “over-fix” molecules, tangling the sequence and blocking the search party. The improved EDC method is vital for small RNAs (like miRNA or siRNA) because it tethers them by the “tail” (the 5′ end), leaving the entire sequence open and accessible. This can enhance detection sensitivity by up to 50-fold.
With the molecules locked in place, it’s time to send in the search party.

3.2.5 Step 5: The Secret Agent (Hybridization and Probes)
Now we must find our specific “needle” in a molecular haystack of millions. We send in a probe—a labeled “secret agent” (either DNA or RNA) that is perfectly complementary to our target sequence. This search is governed by Strict Base Pairing: Cytosine (C) always pairs with Guanine (G), and Adenine (A) pairs with Thymine (T) in DNA or Uracil (U) in RNA.
Successful Probe Factors:
• Specificity: To ensure the probe doesn’t stick to the wrong gene by chance, it must be a minimum of ~25 bases long. At 30 bases, the probability of a random match in a mammalian genome is 1 in 1 billion.
• Length: Longer probes bind more tightly and provide higher sensitivity, though they may take longer to hybridize.
• Labeling: Probes are tagged with radioactive isotopes (like 32P) or non-isotopic labels (like biotin or digoxigenin) so they can be visualized later.
Once the secret agents find their targets, we just need a way to make them glow in the dark.

3.2.6 Step 6: The Reveal (Washing and Detection)
After the search, the membrane is covered in “clumsy” probes that are either floating free or stuck weakly to the wrong targets. We perform a Stringency Wash—using heat and specific buffers—to rinse away everything except the probes that have found their perfect molecular match.
The Final Steps:
• Wash: Remove non-specifically bound probes to eliminate “background noise.”
• Develop: For non-isotopic probes, add enzymes (like alkaline phosphatase) that trigger a light-emitting chemical reaction.
• Image: Place the membrane against X-ray film, a phosphor screen, or a digital imager.
• Analysis: Use densitometry software to measure the bands.
The journey is complete; the blurry bands on the film tell us exactly how much of a gene was active.

Snapshot Summary
| Phase | Technical Action | Purpose (The “So What?”) |
|---|---|---|
| Separation | Denaturing Gel | Sorts molecules by size fairly; formaldehyde prevents folding while MOPS provides a safe buffer. |
| Transfer | Blotting | Moves RNA to a durable surface via physical forces (capillary/vacuum). |
| Fixation | Cross-linking | Covalently tethers RNA (UV for large; EDC for small) so it survives the washing process. |
| Detection | Hybridization | Uses a complementary probe to identify and quantify the specific target gene. |
4. Advanced Methodology: The Strategic Role of Northern Blotting in the Small RNA Era
In the modern molecular biology landscape, Northern Blotting remains a cornerstone technique for the study of gene expression. While high-throughput methods like deep sequencing and microarrays have accelerated the discovery of novel RNA species, and RT-PCR offers high sensitivity, Northern Blotting holds a unique strategic advantage: it is the only analytical method capable of simultaneously providing an accurate display of both the size and abundance of multiple RNA species. As noted in the literature (He, Page 2), this “size-dependent separation” is critical for validating transcript variants and verifying RNA processing—specifically distinguishing between mature microRNA (miRNA) of approximately 20 nucleotides (nt) and their precursors (approximately 70 nt) on a single blot.
However, as research has shifted toward regulatory small RNAs—including miRNA, short-interfering RNA (siRNA), and Piwi-interacting RNA (piRNA)—the inherent sensitivity limitations of traditional Northern Blotting have become a significant bottleneck. These molecules, typically under 40 nt, often escape robust detection. This paper identifies the immobilization step—the process of tethering RNA to the solid support—as the primary variable for optimization. By refining the chemistry of immobilization, researchers can overcome the “sensitivity gap” that has historically hindered small RNA analysis.
4.1 Molecular Mechanisms of RNA Immobilization: UV vs. EDC
The efficiency of hybridization is fundamentally dictated by the physical state of the RNA on the nylon membrane. How the RNA is tethered determines whether its sequence remains accessible to the complementary probe or becomes stochastically obscured by the immobilization chemistry.
4.1.1 The “Base-Consumption” Model of UV Cross-linking
Traditional protocols rely on UV irradiation (typically 1200 mJ) to fix RNA. The prevailing model suggests that UV produces reactive functional groups within the nucleotide bases—principally uridine residues—which then form covalent bonds with the free amine groups on the nylon surface. From an assay development perspective, this mechanism presents a fundamental conflict: it consumes the very bases required for hybridization. In small RNAs (<40 nt), where every nucleotide is vital for probe binding, this base-consumption inherently reduces accessibility. Furthermore, because the UV-triggered reaction is stochastic, “over-cross-linking” can occur, further hindering the probe’s ability to reach its target.
4.1.2 The “Terminal Tethering” Model of EDC Cross-linking
To address these shortcomings, a chemical cross-linking method utilizing 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide (EDC) facilitates the formation of a covalent phosphoramidate bond specifically between the 5’-terminal phosphate of the RNA and the primary amines of the nylon membrane.
Crucially, this reaction requires 1-methylimidazole (pH 8.0) to act as a catalyst and stabilizer for the reactive intermediate. This terminal tethering ensures the RNA is immobilized by its end, leaving the internal sequence entirely free for hybridization. For non-phosphorylated RNA, compatibility can be achieved via pre-treatment with T4 Polynucleotide Kinase (PNK) to add the necessary 5′-phosphate.
Mechanism Comparison: UV Irradiation vs. EDC Mediated Cross-linking
| Feature | UV Irradiation | EDC-Mediated Cross-linking |
|---|---|---|
| Bond Site | Internal bases (principally Uridine) | 5’-terminal phosphate |
| Reaction Time | Seconds to minutes (1200 mJ) | Approximately 2 hours |
| Probe Accessibility | Reduced (due to base consumption) | Maximum (end-tethered) |
| Sequence Integrity | Risk of over-cross-linking | High; sequence remains accessible |
4.2 Comparative Performance Analysis: Sensitivity and Transcript Size
The impact of transcript length on immobilization efficiency is a decisive factor in the resulting signal-to-noise ratio. Data from the Pall & Hamilton study reveals a dramatic performance divide based on transcript size.
4.2.1 The 50-Fold Sensitivity Factor and Compounding Gains
In trials detecting mmu-miR-292-3p in murine embryonic stem cells, EDC-mediated cross-linking demonstrated up to a 50-fold increase in sensitivity compared to UV (with a consistent 30-fold gain in specific ES cell tests). Strategic laboratory optimization can compound these results; for instance, the use of ULTRAhyb ultrasensitive hybridization buffer can increase signal intensity up to 100-fold, effectively reaching the sensitivity levels of nuclease protection assays.
4.2.2 Size-Dependency Thresholds
The benefits of EDC are concentrated in molecules under 40 nt and decline as transcript length increases:
• Small RNAs (<40 nt): Exhibit the highest sensitivity gains (up to 50-fold). Terminal tethering prevents the loss of crucial hybridization sites in these short sequences.
• Intermediate Species (~70 nt): For pre-miRNAs, the advantage of EDC over UV declines significantly.
• Large Transcripts (>100 nt): UV cross-linking remains the preferred method due to its speed and simplicity, as EDC offers no substantial enhancement for large molecules.
Sensitivity Comparison: The “So What?”
| Target Size | UV Cross-linking | EDC Cross-linking | Improvement |
|---|---|---|---|
| Small RNA (<40 nt) | Poor | High | Up to 50-fold |
| Pre-miRNA (~70 nt) | Moderate | Moderate | Negligible |
| Large mRNA (>100 nt) | High | Moderate | UV is superior |
4.3 Technical Implementation and Protocol Optimization
EDC cross-linking is technically straightforward but extremely sensitive to the chemical environment, necessitating a departure from traditional Northern Blot reagents.
4.3.1 Strategic Buffer Requirements: The Tris Conflict
EDC reacts with primary amines; therefore, the buffering system must be entirely free of Tris to prevent reagent neutralization. MOPS-NaOH (pH 7.0) must be used in the gel, the running buffer, and the transfer buffer. Because MOPS is slightly less denaturing than TBE, gels should be run at higher temperatures (50–55°C) to ensure accurate sizing of precursors.
4.3.2 Transfer and Membrane Optimization
For maximum signal, downward elution (passive, slightly alkaline) is preferred over traditional upward transfer as it is faster and results in “tighter” bands. Membrane choice is equally critical: neutral nylon (e.g., Hybond NX) provides the full 50-fold sensitivity gain, whereas positively charged membranes (Hybond N+) show only marginal enhancement with EDC compared to UV.
4.3.3 Parameters and Quality Control
• Incubation: Optimal cross-linking occurs at 60°C for 2 hours. Exceeding these parameters may induce over-cross-linking, resulting in artifactual bands or high background.
• Diagnostic Markers: [32P]-labeled Decade markers serve as a “yardstick.” Hand-held Geiger counter monitoring of the membrane during transfer is essential to ensure a >90% transfer efficiency before proceeding.
4.4 Strategic Selection Framework for Immobilization Strategies
Methodology selection should be a function of target transcript characteristics and laboratory throughput requirements.
Strategy Decision Matrix
| Scenario | Recommendation | Rationale |
|---|---|---|
| A: High-Sensitivity Small RNA Analysis (<40 nt) | EDC Cross-linking | Maximizes probe accessibility; required for low-abundance miRNA/siRNA. |
| B: Standard Messenger RNA Analysis (>100 nt) | UV Cross-linking | Efficiency of UV is sufficient; speed and simplicity outweigh marginal gains. |
| C: Comparative Variant Analysis (pre- vs. mature miRNA) | EDC Priority | Ensures detection of the mature species; UV may fail to detect the 20 nt form entirely. |
| D: Non-phosphorylated Small RNA Analysis | PNK + EDC | Pre-treatment with T4 Polynucleotide Kinase adds the 5′-phosphate for tethering. |
4.5 Bridging the Sensitivity Gap: Modern Enhancements
To approach the sensitivity of more modern assays, we have developed several tactical enhancements:
1. EDC Chemical Cross-linking: For small RNAs (<40 nt), traditional UV cross-linking is suboptimal because it creates random bonds with nucleotide bases (principally Uridine), which can interfere with probe hybridization. By using EDC-mediated cross-linking, we create a covalent phosphoramidate bond specifically at the 5′ terminal phosphate. This tethers the RNA by its end, leaving the entire sequence accessible for the probe and increasing sensitivity by up to 50-fold.
2. Advanced Hybridization Buffers: Utilizing specialized buffers like ULTRAhyb can increase sensitivity 100-fold by pushing hybridization to completion. This allows for the detection of as few as 10,000 to 100,000 molecules.
3. Antisense RNA Probes (Riboprobes): High-specific-activity RNA probes provide a 10-fold increase in sensitivity over DNA probes. Their higher affinity for the target allows for more stringent washing conditions, which effectively lowers background noise and yields a cleaner signal.
5. Research and Clinical Applications: Analyzing Gene Expression Patterns
Case Study: The ob (Obese) Gene and Leptin
Northern Blotting was instrumental in identifying the ob gene product, Leptin.
• Tissue Specificity: Analysis showed ob is expressed exclusively in white adipose tissue (WAT), serving as a signal for fat depot size.
• Nutritional Regulation: Figure 3B of the primary research shows ob mRNA levels are drastically reduced during fasting (24 h) and rapidly restored within 6 h of refeeding, providing critical insights into metabolic signaling.
Case Study: Metallothionein-1
Studies in rats revealed that Metallothionein-1 is induced by cold (4°C) and Zinc (10 mg/kg) in brown adipose tissue (BAT) and the liver. Zinc acts as a major inducer, showing a much greater effect on the liver than 24 h of cold exposure. Interestingly, this gene is not expressed in white adipose tissue under either condition, demonstrating tissue-specific regulation.
Clinical Relevance
• Huntington’s Disease: Assessing neurodegenerative gene expression patterns.
• Breast Cancer: Analyzing oncogene and tumor suppressor transcription rates in various biopsy tissues.
6. Technical Optimization and Troubleshooting: Hard-Won Laboratory Wisdom
Top 5 Strategies for Increased Sensitivity
1. ULTRAhyb Buffer: Can increase hybridization sensitivity up to 100-fold.
2. High Specific Activity Probes: Aim for ≥108 cpm/µg. Asymmetric PCR-generated DNA probes are 3–5 fold more sensitive than random-primed DNA.
3. Antisense RNA Probes: RNA “riboprobes” provide a 10-fold increase in sensitivity over DNA probes.
4. Poly(A) Selection: Enriching for mRNA reduces background noise.
5. EDC Cross-linking: Mandatory for all small RNA species (<40 nt).
Troubleshooting Matrix: The EDC-dPAGE Workflow
| Problem | Possible Cause | Solution |
|---|---|---|
| No signal on blot | EDC decomposition | Use fresh EDC powder; store strictly at -20°C. |
| Low signal/turing | Over-transfer | Marker RNA has passed through to the 3MM paper; reduce transfer time/voltage. |
| Poor retention | Incorrect pH | 1-methylimidazole must be exactly pH 8.0. Check pH meter calibration. |
| Incomplete cross-linking | Buffer Interference | Ensure no Tris was used in the gel or transfer solutions. |
| Artifactual bands | Over-cross-linking | Reduce EDC incubation time or temperature (stay ≤ 60°C). |
Expert Pro-Tip: The Radioactive “Yardstick” Utilize 32P-labeled Decade markers as a procedural monitor. Use a hand-held Geiger counter to track the RNA’s progress. Specifically, monitor the membrane during transfer; if counts are detected on the underlying 3MM filter paper or the positive electrode, over-transfer has occurred, necessitating a reduction in transfer time for future runs.
7. Conclusion: The Future of RNA Blotting
Northern Blotting remains a definitive resource for high-precision gene expression studies. Its versatility in handling diverse probe types—from radiolabeled riboprobes to chemiluminescent anti-sense oligonucleotides—ensures its longevity. By strategically bifurcating the workflow based on target size—utilizing dPAGE and EDC cross-linking for small regulatory species and Agarose/UV for larger transcripts—researchers can achieve unmatched clarity in visualizing the transcriptome. These extensive notes provide the necessary framework for optimizing this classic technique to meet the rigorous demands of modern molecular biology.
7.1 Decision Framework: “When to Use Which?”
Use this reference to align your methodology with your experimental objectives.
| If your experimental goal is to… | Recommended Method |
|---|---|
| Determine the exact size of an mRNA transcript or validate protein size. | Northern Blotting |
| Distinguish between a mature miRNA and its precursor (pre-miRNA). | Northern Blotting |
| Use a probe from a different species to detect a gene of interest. | Northern Blotting |
| Detect a few copies of a message in a very limited sample. | RT-PCR |
| Compare relative abundance across 96 tissue samples quickly. | RT-PCR |
| Analyze samples with questionable RNA integrity. | Prioritize Integrity Checks (Northern will fail). |
Image Summary




References
Pollard, J.W., Perry, C.R., Thurston, C.F. (1988). Northern Blotting. In: Walker, J.M. (eds) New Nucleic Acid Techniques. Methods in Molecular Biology, vol 4. Humana Press. https://doi.org/10.1385/0-89603-127-6:13
Northern blotting: transfer of denatured RNA to membranes. Nat Methods 2, 997–998 (2005). https://doi.org/10.1038/nmeth1205-997
Pall, G., Hamilton, A. Improved northern blot method for enhanced detection of small RNA. Nat Protoc 3, 1077–1084 (2008). https://doi.org/10.1038/nprot.2008.67
He SL, Green R. Northern blotting. Methods Enzymol. 2013;530:75-87. doi: 10.1016/B978-0-12-420037-1.00003-8. PMID: 24034315; PMCID: PMC4287216.
Lovatt, D., & Eberwine, J. (2013). Northern blotting. https://doi.org/10.1016/B978-0-12-374984-0.01065-2
https://en.wikipedia.org/wiki/Northern_blot
https://www.genome.gov/genetics-glossary/Northern-Blot
https://www.nature.com/scitable/definition/northern-blot-287/
https://www.thermofisher.com/np/en/home/references/ambion-tech-support/northern-analysis/general-articles/the-basics-northern-analysis.html
https://www.goldbio.com/blogs/articles/an-overview-of-northern-blot?srsltid=AfmBOoqLaWMq5s3uqAg2tWbBJmJi2V6c6uSi8JChU8UZl09-yLGnSW8V
Questions/Answers
1. How does northern blotting compare to modern techniques for analyzing gene expression?
While modern high-throughput technologies like RNA-Seq and microarray analysis have largely superseded northern blotting for large-scale gene expression profiling, northern blotting remains a standard and essential method due to several unique capabilities.
Sensitivity and Throughput
In comparison to modern techniques, conventional Northern blot analysis is generally less sensitive and offers lower throughput. Real-time PCR (RT-PCR) and RNase protection assays provide significantly higher sensitivity; RT-PCR, in particular, is capable of measuring expression levels as low as a few copies of a specific mRNA in a tissue sample. Furthermore, while high-throughput methods like deep sequencing (RNA-Seq) and microarrays can quantify thousands of mRNA species simultaneously, northern blotting typically analyzes only one or a very small number of target genes.
Size Determination and Accuracy
The primary advantage of northern blotting over newer techniques is its unmatched ability to accurately display RNA size. While modern bioinformatic tools and sequencing can quantify abundances, they do not provide direct information regarding the physical size of the transcript. Northern blotting allows researchers to:
• Determine the exact size of a specific transcript, which provides an estimate for its protein-coding capacity.
• Identify alternatively spliced transcripts and splice variants.
• Simultaneously display the sizes and amounts of multiple RNAs that share sequence identity, such as the mature and precursor forms of microRNAs (miRNAs).
• Detect deletions or errors in transcript processing.
Specificity and Reliability
Northern blotting is favored for its high specificity, which is critical for reducing false-positive results, even though it has lower sensitivity than RT-PCR. The technique is exceptionally versatile, allowing the use of probes with only partial homology, such as cDNA from a different species or genomic DNA fragments that might contain introns. Additionally, northern blot membranes are durable; they can be stored and reprobed for years, whereas most modern electronic or sequencing-based data are fixed once the procedure is complete.
Sample Integrity Assessment
A key procedural benefit of northern blotting is the ability to visually assess RNA quality and quantity on the gel prior to the blotting process. This is vital because even slightly degraded RNA samples can severely compromise data quality; for instance, a single cleavage in just 20% of target molecules can decrease the returned signal by 20%. Modern high-throughput techniques often rely on automated quality checks rather than this direct visual confirmation of sample integrity.
2. Why must RNA samples be denatured before undergoing size separation by electrophoresis?
RNA samples must be denatured before and during electrophoresis to ensure that the separation of molecules is based strictly on their molecular weight rather than their physical shape.
Prevention of Secondary Structures
Unlike double-stranded DNA, single-stranded RNA has a strong tendency to form complex intramolecular base pairs, resulting in secondary and tertiary structures. Denaturation ensures that these RNA strands are unfolded and that there is no bonding between or within strands. If these structures are not disrupted, the RNA molecules will not move through the gel matrix as linear chains, which is essential for accurate identification.
Impact on Migration Rate
The formation of secondary structures can significantly change the migration rate of RNA in several ways:
• Hydrodynamic Volume: Folded RNA occupies a different physical volume (hydrodynamic volume) than linear RNA, which alters how it navigates the pores of the gel matrix.
• Charge Distribution: Structural folding can change the distribution of the negative charge provided by the phosphate backbone, affecting how the molecule responds to the electric field.
• Anomalous Sizing: Without complete denaturation, certain RNA molecules—such as longer pre-microRNAs—may run faster than expected, leading to incorrect size estimates.
Methods of Denaturation
To maintain RNA in an unfolded state, researchers use a combination of chemical and thermal treatments:
• Chemical Denaturants: Formaldehyde, formamide, or glyoxal/DMSO are commonly added to the gel or the loading buffer to prevent the re-formation of RNA secondary structures during the run.
• Heat Treatment: Samples are typically heat-denatured at 65°C to 95°C before loading.
• Snap Cooling: Following heat treatment, samples are often snap-cooled on ice to inhibit the renaturation of RNA structures before they can be loaded into the gel.
By ensuring the RNA remains linear, electrophoresis can precisely separate thousands of different transcripts based on their physical length, providing a direct measure of their size and coding capacity.
3. How does EDC cross-linking improve small RNA detection sensitivity?
1-ethyl-3-(3-dimethylaminopropyl) carbodiimide (EDC) cross-linking improves detection sensitivity for small RNA molecules (typically <40 nt) by up to 50-fold compared to conventional UV cross-linking. This enhancement is achieved through a more efficient and less intrusive immobilization mechanism that preserves the RNA sequence for hybridization.
Mechanism of Improvement
• Terminal Tethering: EDC facilitates the formation of a covalent phosphoramidate bond between the 5′-terminal phosphate of the small RNA and the primary amine groups on the surface of a neutral nylon membrane. This resulting linkage tethers the RNA by only one end.
• Full Sequence Accessibility: Because the RNA is immobilized only at its terminus, the entire remaining sequence remains fully available and accessible for hybridization with complementary probes.
• Prevention of “Consumption” of Bases: Conventional UV cross-linking is thought to react with the nucleotide bases (principally uridine) to form bonds with the membrane. This process “consumes” these bases, making them unavailable for base-pairing and potentially hindering probe accessibility through over-cross-linking.
Impact on Sensitivity and Size Specificity
• Critical for Short Sequences: For very short sequences like microRNAs (~20 nt) or siRNAs, the loss of even a few bases to UV cross-linking significantly reduces the stability and signal of probe hybridization. EDC preserves these bases, leading to a much stronger signal.
• Size-Dependent Benefits: The sensitivity gains provided by EDC are most significant for RNA molecules under 40 nucleotides. The advantage declines as the transcript size increases; for larger molecules, such as miRNA precursors (~70 nt), EDC offers no significant enhancement over standard UV methods.
• Reduced Exposure Times: Due to the massive increase in signal intensity, researchers can use shorter exposure periods to detect the target RNA after hybridization compared to traditional methods.
4. Why is northern blotting still used for small RNAs?
Northern blotting remains a popular and valuable analytical method for small RNAs because it is unmatched in its ability to accurately display the physical size of RNA molecules. While modern high-throughput methods like deep sequencing and microarrays can quantify abundance, they do not provide direct information regarding the physical size and size complexity of the transcripts.
The continued use of northern blotting for small RNAs is driven by the following factors:
Simultaneous Detection of Related Species
Northern blotting can simultaneously display the sizes and amounts of multiple small RNAs that share significant sequence identity. For example, a single blot can detect both the mature form (~20 nt) and the precursor form (~70 nt) of a microRNA (miRNA). This specific pattern provides strong evidence that a sequence is indeed a miRNA, making the technique essential for the validation and functional characterization of novel small RNAs.
High Specificity and Verification
The technique is favored for its high specificity, which is critical for reducing false-positive results that can occur with other expression analysis methods. As new RNA species are discovered through high-throughput sequencing, northern blotting is frequently used as the standard for characterization and verification of their presence in tissues and cells.
Enhanced Sensitivity Through Technological Improvements
While conventional northern blots were once considered less sensitive, the development of EDC-mediated chemical cross-linking has significantly improved detection for small RNA molecules under 40 nucleotides. This method improves detection sensitivity by up to 50-fold compared to traditional UV cross-linking. By tethering the RNA to the membrane at only one end, the full sequence remains accessible for hybridization, which results in much stronger signals and shorter exposure times.
Procedural Advantages
• Quality Control: Researchers can visually assess the quality and integrity of the RNA on the gel (using EtBr staining) before the blotting process, ensuring the samples are not degraded.
• Versatility: The process is highly versatile, allowing for the use of probes with only partial homology, such as cDNA from different species or genomic fragments containing introns.
• Durability: Unlike digital sequencing data, northern blot membranes are physically durable and can be stored and reprobed for years.
• Refined Separation: For small RNAs, the use of denaturing polyacrylamide gel electrophoresis (PAGE) with urea provides much higher resolution for fragmented or microRNA-sized molecules than standard agarose gels.
5. Why is EDC cross-linking less effective for long RNA?
EDC cross-linking is less effective—or rather, provides no significant advantage—for long RNA compared to conventional methods because the specific problems it solves are only critical for very short sequences.
According to the sources, the effectiveness of EDC cross-linking declines as transcript size increases due to the following factors:
Negligible Impact of “Base Consumption”
Conventional UV cross-linking facilitates immobilization by producing reactive functional groups within the nucleotide bases (principally uridine), which then form covalent bonds with the membrane. This process “consumes” those bases, making them unavailable for hybridization with a probe.
• For short RNA: In a 20-nt microRNA, the loss of even a few bases to UV cross-linking significantly destabilizes hybridization and reduces the signal.
• For long RNA: For transcripts longer than 70–100 nucleotides, the loss of a few bases has a negligible impact on the overall stability and signal intensity because the vast majority of the sequence remains available for the probe.
Lack of Sensitivity Enhancement
While EDC can improve detection sensitivity for small RNAs (<40 nt) by up to 50-fold, researchers found that no such enhancement occurs for larger molecules. Data indicate that the sensitivity gain provided by EDC declines as the size of the RNA increases; for instance, it offers no significant benefit for miRNA precursors that are approximately 70 nucleotides long.
Simplicity vs. Benefit Trade-off
Conventional immobilization methods, such as UV irradiation (254 nm) or baking at 80°C, are rapid, inexpensive, and well-established for transcripts in the mRNA size range. Because EDC cross-linking is a more involved chemical process—taking 15 minutes to 2 hours compared to the near-instantaneous nature of UV—it is not the preferred method when it does not provide a clear sensitivity advantage. Consequently, UV cross-linking remains the standard choice for RNA greater than 70 nucleotides due to its greater simplicity.
6. What is the size limit for EDC cross-linking?
The effectiveness of 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide (EDC) cross-linking is highly dependent on the length of the RNA molecule, with its primary benefits restricted to very small sequences.
Functional Size Limits
• Optimal Range (<40 nt): The enhancement in detection sensitivity provided by EDC—which can be up to 50-fold—is most significant for RNA molecules under 40 nucleotides in length. This includes regulatory species such as microRNA (miRNA), short-interfering RNA (siRNA), and Piwi-interacting RNA.
• Threshold for Declining Benefit (~70 nt): No significant detection enhancement is observed for larger molecules, such as miRNA precursors, which are approximately 70 nucleotides long. As the size of the RNA transcript increases, the specific advantages of EDC cross-linking decline.
• Transition to Standard Methods (>70 nt): Because conventional UV cross-linking is more technically straightforward and works well for transcripts in the mRNA size range, it remains the method of choice for any RNA greater than approximately 70 nucleotides.
Binding Capacity Limits
Beyond the nucleotide length of the transcript, there is also a physical limit to how much RNA can be immobilized. While EDC facilitates a terminal covalent bond, it is likely subject to a similar upper binding capacity limit as UV cross-linking, which is dictated by the availability of free amine groups on the surface of the nylon membrane. This limit may be reached when using highly enriched RNA fractions rather than total RNA preparations.
7. How does RNA structure affect migration rate during gel electrophoresis?
RNA structure significantly affects migration rate during gel electrophoresis by interfering with the relationship between a molecule’s physical length and its movement through the gel matrix. In its natural state, single-stranded RNA has a strong tendency to form complex intramolecular base pairs, resulting in secondary and tertiary structures.
The specific ways these structures impact migration include:
• Alteration of Hydrodynamic Volume: Folded RNA molecules occupy a different physical volume (hydrodynamic volume) than linear chains of the same length. This change in shape affects how easily the RNA fragments can navigate the sieve-like pores of the gel matrix.
• Changes in Charge Distribution: While RNA has a net negative charge proportional to its length, structural folding can change the distribution of this charge, affecting how the molecule responds to the electric field driving it toward the positive anode.
• Anomalous Migration Speeds: Without proper denaturation, RNA molecules may not migrate according to their actual size. For example, researchers have found that longer precursor microRNAs (~70 nt) may run faster than expected (appearing as ~50 nt) when electrophoresis buffers are not sufficiently denaturing.
• Reduced Resolution: Failure to maintain RNA in a linear, unfolded state can result in less sharp bands and a “smear” on the gel, making it difficult to precisely separate and identify transcripts of different lengths.
To ensure that migration is strictly a function of molecular weight, researchers use denaturing agents such as formaldehyde, glyoxal, or urea to disrupt these structures. Additionally, samples are often heat-denatured (typically at 65°C to 95°C) and run in preheated buffers to aid in maintaining a linear conformation for accurate sizing.
8. What is the size limit for EDC cross-linking?
The effectiveness of 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide (EDC) cross-linking is primarily determined by the length of the RNA transcript, with its unique advantages restricted to a specific size range.
Functional Size Limits
• Optimal Range (<40 nt): The significant enhancement in detection sensitivity provided by EDC—which can be up to 50-fold—is most effective for small RNA molecules under 40 nucleotides in length. This includes species such as microRNA (miRNA), short-interfering RNA (siRNA), and Piwi-interacting RNA.
• Declining Benefit (40–70 nt): The gains in sensitivity offered by the EDC method decline as the size of the RNA of interest increases.
• Practical Threshold (~70 nt): Research indicates that no substantial enhancement is detected for larger molecules, such as miRNA precursors that are approximately 70 nt long. Because chemical cross-linking is more technically involved than conventional methods, UV cross-linking remains the standard choice for any RNA greater than approximately 70 nt.
Membrane Binding Capacity Limits
In addition to transcript length, there is a physical limit to the short RNA-binding capacity of the neutral nylon membranes used in this process. It is presumed that EDC cross-linking has a similar upper binding limit to UV cross-linking, which is dictated by the density of free amine groups available on the surface of the membrane. This saturation limit is most likely to be reached when researchers use highly enriched RNA fractions instead of standard total RNA preparations.
9. What type of nylon membrane works best for EDC?
For EDC-mediated chemical cross-linking, the most effective option is a neutral nylon membrane, specifically nylon 6,6 neutral membranes.
The sources provide the following details regarding membrane selection for this technique:
• Recommended Model: Researchers routinely use and recommend Hybond NX (Amersham/Pharmacia). While other neutral nylon 6,6 membranes should theoretically work just as well, this specific model is the standard for the improved small RNA detection protocol.
• Mechanism: EDC facilitates a covalent phosphoramidate bond between the 5′-terminal phosphate of the small RNA and the primary amine groups located on the surface of the neutral nylon membrane.
• Comparison to Positively Charged Membranes: While positively charged nylon membranes (such as Hybond N+) are often the most effective for standard Northern blotting because they have a high affinity for negatively charged nucleic acids, they are less effective for EDC cross-linking. Testing shows that using EDC with a positively charged membrane provides only a slight enhancement in detection sensitivity compared to conventional UV cross-linking.
In contrast, using EDC with a neutral nylon membrane can improve the detection sensitivity of small RNA molecules (under 40 nucleotides) by up to 50-fold compared to traditional UV methods.
10. How do formaldehyde and urea effectively disrupt RNA secondary structure?
Formaldehyde and urea act as chemical denaturing agents during gel electrophoresis to ensure that single-stranded RNA remains in an unfolded, linear state. Single-stranded RNA has a natural tendency to form intramolecular base pairs, creating complex secondary and tertiary structures that can alter its migration rate by changing its hydrodynamic volume and charge distribution.
Formaldehyde is traditionally added to agarose gels to limit the formation of these secondary structures. It is also frequently included in RNA loading buffers, often in combination with formamide, to prepare the sample for separation. Formaldehyde is used in these procedures because alkaline denaturants, such as the sodium hydroxide used in Southern blotting, would degrade the RNA.
Urea is primarily used as a denaturant in polyacrylamide gel electrophoresis (dPAG), which is the preferred method for analyzing smaller RNA species like microRNAs or fragmented RNA. In these protocols, high concentrations of urea—typically 7 M—are incorporated directly into the gel matrix. This ensures the RNA stays denatured while moving through the electric field.
By disrupting structural bonding, these chemicals ensure that RNA molecules migrate through the gel matrix strictly according to their molecular weight and physical length. This precise separation is essential for determining the accurate size of transcripts and their protein-coding capacity.
11. What advantages do mRNA enrichment methods provide for sensitivity?
mRNA enrichment methods provide significant advantages for detection sensitivity in northern blotting by increasing the concentration of target molecules and overcoming the physical limitations of the electrophoresis process.
According to the sources, the specific advantages include:
Increased Concentration of Target Molecules
Because messenger RNA (mRNA) comprises only about 0.5% to 3% of total RNA extracted from a tissue, the vast majority of a total RNA sample consists of ribosomal RNA (rRNA) and transfer RNA (tRNA). By isolating mRNA through poly-A (+) selection (using oligo-dT columns or beads), researchers can load a much higher effective amount of the target gene’s RNA into a single gel lane. For example:
• 10 µg of purified mRNA is equivalent to loading approximately 300–350 µg of total RNA.
• This purification process can increase the detection limit by approximately 30-fold.
Overcoming Physical Constraints
Sensitivity in standard northern blots is often limited by the physical capacity of the gel and the membrane. There is an upper limit to the total quantity of RNA that can be loaded onto a gel without losing resolution or saturating the transfer membrane. mRNA enrichment allows researchers to bypass these limits by removing the high-abundance rRNA and non-coding RNA that would otherwise take up the limited space on the gel and membrane.
Reduced Interference
Removing non-mRNA components increases the sensitivity of downstream applications by eliminating most of the non-coding RNA and rRNA, which can otherwise interfere with the analysis of the specific mRNA of interest. This results in a higher signal-to-noise ratio, making it easier to detect genes with low expression levels that might be below the detection limit in a total RNA preparation.
12. How does poly-A selection isolate mRNA from total RNA?
Poly-A selection isolates messenger RNA (mRNA) from total RNA by exploiting a specific structural feature common to most eukaryotic mRNA: the poly-A tail.
The Binding Mechanism
The procedure utilizes affinity purification techniques, specifically oligo-dT cellulose chromatography columns or beads primed with oligo-dT. “Oligo-dT” refers to short synthetic chains of deoxythymidine, which have a natural affinity for the long string of adenine nucleotides that make up the poly-A tail. When total RNA is passed through the column or incubated with the beads, the oligo-dT sequences selectively bind to the poly-A tails of the mRNA molecules through base-pairing.
Selective Removal of Non-mRNA
Because other major types of RNA—such as ribosomal RNA (rRNA), transfer RNA (tRNA), and various non-coding RNAs—typically lack a poly-A tail, they do not bind to the oligo-dT. These non-mRNA components remain in the solution or pass through the column, leading to their selective removal from the sample. Once the high-abundance rRNA and tRNA have been washed away, the isolated mRNA can be released from the oligo-dT substrate for use in downstream applications.
Impact on Sensitivity
This isolation is a critical enrichment step because mRNA comprises only about 0.5% to 3% of the total RNA extracted from a tissue or cell sample. By removing the interfering non-mRNA components, which represent the vast majority of the sample, researchers can increase the detection sensitivity of techniques like northern blotting by approximately 30-fold. This allows for the analysis of specific gene transcripts that might otherwise be below the detection limit in a total RNA preparation.
13. Does heat-denaturing RNA carry a high risk of transcript degradation?
Heat-denaturing RNA is a standard procedural step, but it does carry an inherent risk of transcript degradation, particularly if high temperatures are maintained for extended periods without protective chemical agents.
Degradation Risks and Mitigation
• Threshold for Damage: During the hybridization phase of northern blotting, maintaining temperatures at or above 65°C is considered undesirable because RNA degradation is significantly increased under these conditions.
• Use of Chemical Denaturants: To mitigate this risk, researchers frequently use formamide in transfer and hybridization buffers. Formamide lowers the required annealing temperature for probe-RNA interactions, allowing the process to occur at 37–42°C and eliminating the need for higher temperatures that could compromise RNA integrity.
• Brief High-Heat Exposure: In the initial preparation for electrophoresis, RNA samples are commonly heat-denatured at 95°C, but this exposure is strictly limited to 1–5 minutes and is immediately followed by snap cooling on ice. This brief pulse is used to unfold secondary structures while minimizing the time the RNA is vulnerable to thermal breakdown.
Impact on Sensitivity and Sample Loss
• Membrane Loss: High temperatures are also problematic during the “stripping” process, where an initial probe is removed so a membrane can be reused. Using high heat to strip probes accelerates the loss of RNA from the membrane, which can limit the useful lifespan of the blot.
• RNase Activity: A critical factor in degradation risk is the presence of RNases, which are ubiquitous and highly stable enzymes. Because even slight degradation can severely compromise data quality—for instance, a single cleavage in 20% of target molecules reduces the signal by 20%—meticulous care and the use of RNase-free reagents are required alongside temperature control.
In summary, while brief heat exposure is necessary for denaturation, prolonged high temperatures are avoided in favor of chemical denaturants to prevent significant transcript degradation.
14. What physical constraints limit RNA loading on gels?
Physical constraints limiting RNA loading on gels involve the capacity of the equipment, the resolution of the separation, and the binding limits of the transfer materials.
Well Volume and Sample Concentration
The first physical constraint is the capacity of the indented wells created by the gel comb during preparation. The volume of the RNA sample, combined with its loading dye, must not exceed the physical space available in the well. If a sample is too diluted to fit within these dimensions, researchers must use salt precipitation to concentrate the RNA into a smaller volume before loading.
Loss of Resolution
Gels have an upper limit to the total quantity of RNA they can accurately separate; exceeding this amount leads to a significant loss of resolution. Overloading a lane can cause RNA fragments to migrate poorly, resulting in smeared bands rather than sharp, distinct signals. This smearing makes it impossible to precisely determine the size of the transcript or distinguish between different splice variants.
Membrane Binding Capacity
The detection limit cannot be increased indefinitely by simply loading more RNA because the transfer membrane has a fixed binding capacity.
• Saturation: Nylon membranes rely on free amine groups or specific reactive sites to immobilize RNA through UV or chemical cross-linking. If too much RNA is loaded, the membrane becomes saturated, and the excess target molecules will not be captured, leading to inaccurate quantification.
• Small RNA Limits: This capacity limit is particularly relevant when using highly enriched RNA fractions, where the concentration of short sequences might approach the membrane’s physical saturation point.
Transfer Efficiency and Over-transfer
The physical mechanics of moving RNA from a gel to a membrane also impose limits. Large quantities of total RNA can interfere with the efficiency of the transfer process. Furthermore, attempting to force large amounts of RNA onto the membrane can result in “over-transfer,” where target molecules pass through the membrane entirely and are deposited on the underlying filter paper or electrode.
Gel Fragility and Matrix Integrity
The physical stability of the gel matrix itself acts as a constraint. Agarose and polyacrylamide gels are inherently fragile, making them difficult to handle during the complex sandwich assembly required for transfer. Additionally, the flimsiness of the gel increases as the concentration of the matrix (such as acrylamide) decreases, which can limit the types of RNA that can be analyzed effectively.
15. How do snap cooling and chemical denaturants protect RNA integrity?
Snap cooling and chemical denaturants protect RNA integrity by maintaining transcripts in a linear, unfolded state and preventing damage from both excessive heat and enzymatic activity.
Snap Cooling and Structural Control
• Inhibiting Renaturation: During electrophoresis preparation, RNA samples are typically pulse-heated (often at 95°C for 1–5 minutes) and then immediately snap-cooled on ice. This process inhibits the renaturation of RNA structures, ensuring the molecules remain unfolded and linear.
• Facilitating Sizing: By preventing the re-formation of intramolecular base pairs, snap cooling ensures that the RNA occupies a consistent hydrodynamic volume and charge distribution, which is essential for accurate size determination during electrophoresis.
• Ease of Handling: Practically, keeping samples on ice after denaturation also makes sample loading easier for the researcher.
Chemical Denaturants and Thermal Protection
• Formamide and Temperature Regulation: Chemical denaturants like deionized formamide are added to loading and hybridization buffers to maintain RNA integrity. Formamide works by lowering the annealing temperature of the probe-RNA interaction.
• Preventing Thermal Degradation: By allowing hybridization and transfer to occur at lower temperatures (37–42°C) rather than 65°C or higher, formamide eliminates the need for high heat that would otherwise significantly increase RNA degradation.
• Maintaining Linearity: Formaldehyde and urea are used in gel matrices to prevent the formation of secondary structures during the run. This ensures that migration is strictly a function of molecular weight rather than physical shape.
Protection Against Enzymatic Damage
• Inactivating RNases: Chaotropic agents used during the extraction process, such as guanidinium isothiocyanate, protect integrity by disrupting cells while simultaneously denaturing and inactivating RNases.
• Broad-Spectrum Inhibition: Because RNases are ubiquitous and highly stable, using chemical inhibitors and denaturants is vital; even slight degradation—such as a single cleavage in 20% of target molecules—can result in a 20% loss of signal.
• Standard Precautions: To further protect integrity, researchers use RNase-free reagents and equipment, such as DEPC-treated water or specialized decontamination solutions, to prevent environmental contamination.
16. What causes RNA to pass through membranes during transfer?
The primary cause of RNA passing through a membrane during the transfer process is excessive duration, a phenomenon often referred to as “over-transfer”.
Causes of Over-transfer and Sample Loss
• Blotting Time: If the transfer process—whether capillary, vacuum, or electrophoretic—is allowed to proceed for too long, the RNA molecules may not stop at the membrane surface but instead pass entirely through it. In these instances, the RNA is deposited on the underlying 3MM filter paper or the positive electrode of the transfer unit.
• Binding Capacity Saturation: Nylon membranes have a finite binding capacity determined by the density of free amine groups on their surface. If a researcher loads highly enriched RNA fractions that exceed this physical limit, the excess RNA molecules will not be captured by the membrane and may pass through into the support materials.
• Inadequate Immobilization: While not strictly a cause of passing “through” during the initial move, a failure to properly cross-link the RNA (via UV light, heat, or EDC) immediately after transfer will cause the RNA to be lost from the membrane during subsequent hybridization and washing steps.
• Thermal Loss during Stripping: If a membrane is being reused, the high temperatures typically used to strip an old probe can accelerate the physical loss of the immobilized RNA from the membrane surface.
Monitoring and Prevention
Researchers can monitor whether RNA is passing through the membrane by using radioactive size markers. By monitoring the transfer unit with a hand-held Geiger counter, a scientist can ensure the majority of the counts are retained on the membrane. If significant radioactivity is detected on the 3MM paper below the nylon, it indicates that over-transfer has occurred, which typically leads to a severely reduced signal from the target sample.
To prevent this, transfer times must be optimized for the specific equipment being used, and the membrane must be positioned correctly between the gel and the positive electrode. Additionally, using positively charged nylon membranes is generally more effective for retention because they have a high affinity for negatively charged nucleic acid molecules.
17. How does RNA overloading impact band resolution on gels?
RNA overloading significantly compromises data quality by causing a loss of resolution during the separation process. Because gel electrophoresis and membrane transfer have physical constraints, there is a fixed upper limit to the quantity of total RNA that can be loaded into a single lane.
The impact of exceeding this limit includes:
• Band Smearing: Instead of appearing as sharp, distinct signals, overloaded RNA often results in a “smear” on the gel. This makes it difficult or impossible to accurately determine the physical size of a transcript or distinguish between multiple transcripts of similar size, such as alternative splice variants.
• Reduced Quantitative Accuracy: Simply increasing the amount of RNA loaded does not indefinitely increase the detection limit; once the gel’s separation capacity is exceeded, the resulting loss of resolution and potential saturation of the transfer membrane prevent reliable quantification.
• Retrospective Indicators: In total RNA preparations, researchers often assess the clarity and sharpness of high-abundance bands (such as tRNA or ribosomal subunits) to determine if the run was successful; if these bands appear as a heavily stained smear, it indicates that the loading was too high or the sample integrity was poor.
To avoid these resolution issues while still detecting low-abundance genes, researchers often use mRNA enrichment (poly-A selection). This allows them to load a higher effective concentration of target molecules—equivalent to roughly 300–350 µg of total RNA—into a single lane while staying within the physical loading limits required to maintain sharp band resolution. Additionally, technical factors such as urea leaching from the gel matrix into the wells before loading can also negatively impact resolution independently of the RNA quantity.






