1. Introduction: The Strategic Imperative of Haploid Technology
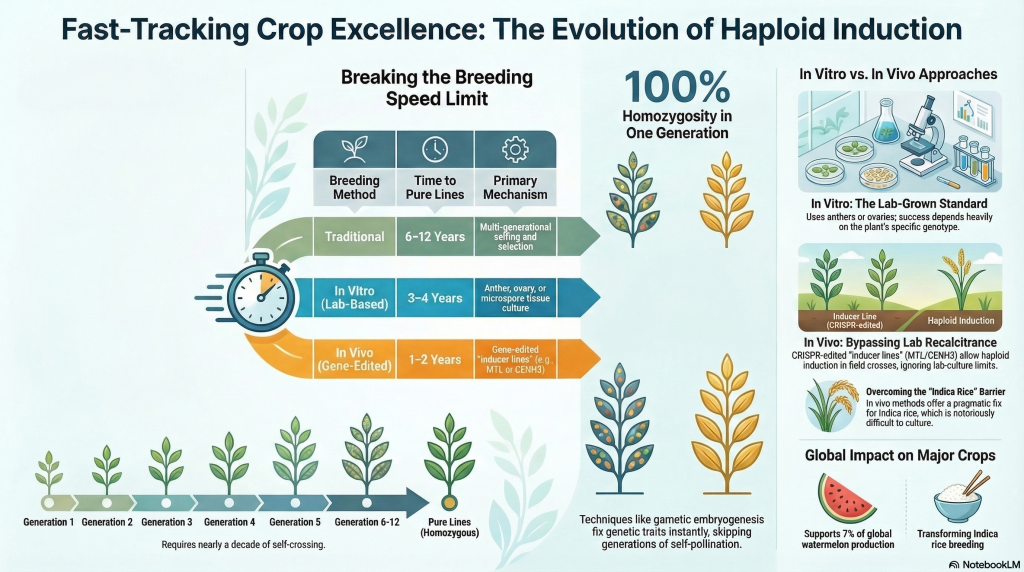
In the hyper-competitive landscape of global agricultural innovation, the speed of variety release is the ultimate differentiator. Traditional breeding programs are fundamentally tethered to a 6–12 year cycle to achieve the high levels of homozygosity required for stable performance. This timeline represents a significant opportunity cost. Strategically, the implementation of Haploid Induction technology allows for the production of 100% pure Homozygous Lines in just 1–2 years. By bypassing multiple generations of selfing, breeding programs can secure F1 parent line exclusivity and respond to shifting market demands—such as climate-resilient Oryza sativa or high-quality Citrullus lanatus L.—three to four times faster than competitors.
1.1 The Genomic Blueprint: Understanding Haploids vs. Diploids
To appreciate the DH shortcut, we must first master the biological states of the plants we manipulate. Most of our essential crops are diploids, carrying two complete sets of chromosomes (one from each parent). In the lab, however, we focus on haploids, which contain only a single set. For example, while a standard watermelon is a diploid with 2n = 22 chromosomes, its haploid counterpart (n) carries only 11.
| Criterion | Haploid (n) | Diploid (2n) |
|---|---|---|
| Chromosome Count | A single set (half the somatic number). | Two complete sets (the standard somatic state). |
| Physical Characteristics | Morphologically scaled-down; typically smaller with reduced vigor. | Standard size; robust growth and full fertility. |
| Role in Nature | Rare spontaneous occurrences (e.g., first observed in Datura stramonium in 1922). | The dominant state for growth and reproduction in most angiosperms. |
For a plant breeder, the “gold standard” is Homozygosity. This is a state where a plant carries identical alleles for a specific trait across both chromosome sets. Achieving homozygosity is the only way to “lock” a trait—ensuring that a plant’s superior characteristics, such as drought tolerance or disease resistance, do not segregate or “disappear” in future generations. While traditional methods take years to approximate this state, DH technology achieves 100% homozygosity immediately. This biological leap is essential because the traditional wait for genetic stability is the primary bottleneck in modern food security.
1.2 The 10-Year Shortcut: Turning a Decade into a Season
The “holy grail” for any plant breeder is achieving 100% homozygosity—creating a “pure” line where every gene is identical across chromosome pairs. In traditional breeding, this requires 6 to 12 years of repeated self-crossing.
Haploid induction technology changes the math. By generating haploids (plants with half the normal number of chromosomes) and then doubling those chromosomes to create doubled haploids (DH), breeders fix homozygosity in a single step. This biological pivot, where a reproductive cell is redirected to grow into a whole plant, is a process called gametic embryogenesis.
1.2.1 The “Time Barrier” in Traditional Plant Breeding
In conventional breeding, we rely on “self-cross and selection.” We take a hybrid and pollinate it with itself, selecting the best offspring and repeating the process for nearly a decade.
The Traditional Inbred Pure Line Process:
- Hybridization: Crossing two genetically diverse parents to create a heterozygous F1 generation.
- Repetitive Selfing: Manually pollinating the plant with its own pollen.
- Selection & Filtering: Identifying and keeping only the plants that display the desired traits.
- Generational Progression: Repeating this cycle for 7 to 10 generations.
The Reality of the “Time Barrier”:
- Watermelon: Requires 8–10 years of repetitive selfing to reach pure lines.
- Rice: Often takes 6+ years to stabilize a new cultivar.
- Brassicaceae (Cabbage/Mustard): Can require 6–12 years to fix traits.
Even after seven generations of labor-intensive “selfing,” a line only reaches approximately 99% homozygosity. In contrast, DH technology bypasses this repetitive labor entirely, producing a 100% homozygous “pure line” in the very first generation.
This technology allows for the immediate stabilization of desirable traits. As research in the field highlights:
“In a single phase, the entirely homozygous lines can be developed from heterozygous parents… this special property has consequences that can be exploited to the advantage of the plant breeders.”
This guide provides a high-level strategic analysis of in vitro systems (Androgenesis and Gynogenesis) and the emerging paradigm of in vivo induction. We focus on the procedural nuances required to produce Doubled Haploids (DH) across high-value crop models, addressing the biological bottlenecks that have historically characterized “recalcitrant” species.
2. Biological Foundations: Androgenesis and Gynogenesis Pathways
2.1 The Scientist’s Toolkit: In Vitro vs. In Vivo Induction
We utilize two primary pathways to trigger haploid development, depending on the species and its “recalcitrance”—a term we use for “stubborn” genotypes that resist standard protocols.
| Method Type | Biological Source | Key Examples |
|---|---|---|
| In Vitro | Anthers (male) or Ovaries (female) cultured in a sterile lab environment. | Watermelon, Indica Rice, Brassicaceae, Datura. |
| In Vivo | Specialized “Inducer Lines” used in controlled field crosses. | Maize, Indica Rice (via CRISPR/Cas9). |
The Androgenic Switch: Triggers and Stress Factors
To flip the switch from the gametophytic path (pollen) to the sporophytic path (embryo/callus), we apply targeted stress. This process is designed to increase the frequency of “P-grains” (embryogenic pollen grains)—specific populations of microspores that have the inherent potential to become embryos.
Stress Triggers vs. Biological Response
| Stress Trigger | Biological Benefit / Cellular Change |
|---|---|
| Cold Stress (10°C for 7–10 days) | Applied to intact or excised inflorescences. Causes mitochondrial condensation and cytoplasmic attenuation, disrupting normal pollen development. |
| Osmotic Stress / Starvation | Induces plasmolysis, which physiologically isolates cells within the anther, allowing them to develop independently into embryos. |
By applying these stresses, we aren’t just harming the tissue; we are deliberately redirecting the cell’s developmental map toward embryogenesis.
2.1.1 The In Vitro Pathway: Reprogramming the Cell
This involves “tricking” reproductive cells into growing into whole plants without fertilization.
- Androgenesis: Developing haploids from male gametes (anthers/microspores).
Timing the “Booting Stage”: The Window of Opportunity
The secret to successful androgenesis lies in catching the microspore at its most plastic state. This occurs during the booting stage, when the panicle is still enclosed in the leaf sheath. At this moment, the microspores are near the first mitotic division—specifically at the mid- to late uni-nucleate or early bi-nucleate stages. This is the biological “trigger point” where the cell can be diverted from becoming a pollen grain to becoming an embryo.
Because microscopic staging is labor-intensive, we use reliable morphological markers to identify this window:
- Internode Distance: The gap between the flag leaf and the penultimate leaf.
- Panicle Length: The size of the developing inflorescence within the “boot.”
- Anther-to-Spikelet Ratio: The physical size of the anther relative to the protective spikelet.
These markers are merely the prelude; once identified, the microspore must undergo a cellular “shock” to initiate its reprogramming.
- Gynogenesis: Developing haploids from female gametes (ovaries). We use this when a genotype is recalcitrant to androgenic methods.
The fundamental strategic value—the “So What?”—of gametic embryogenesis is the deliberate reprogramming of cellular fate. In vivo, gametic cells are genetically programmed for fertilization and subsequent development. Through targeted laboratory interventions, we force a switch from the gametophytic to the sporophytic development pathway. This effectively tricks a single-set (n) microspore or egg cell into behaving like a zygote, enabling the fixation of the entire genome in a single generation.
While androgenesis remains the primary workhorse due to high starting material volume, the comprehensive breeder’s toolkit must include maternal pathways:
| Feature | Androgenesis (Anther/Microspore Culture) | Gynogenesis (Ovary/Ovule Culture) | Parthenogenesis (Irradiated Pollen) |
|---|---|---|---|
| Definition | Development from male gametophytes. | Development from female gametophytes. | Embryo development triggered by sterile pollen. |
| Primary Target Tissues | Anthers or isolated microspores. | Unpollinated ovaries or ovules. | In situ unfertilized egg cells. |
| Efficiency Levels | Theoretically high; thousands of microspores per anther. | Generally the least preferred due to low coherence and limited cells. | High specificity; bypasses some in vitro tissue culture hurdles. |
| Common Applications | Standard for cereals and Brassicaceae; problematic in indica rice. | Vital fallback for genotypes non-responsive to androgenesis (e.g., cucumber). | Emergent tool for specific cucurbit species (Watermelon). |
2.1.2 The In Vivo Pathway: Breaking the Impasse
For years, Indica rice remained a “main contender” for zero hunger but was notoriously recalcitrant to lab culture. The breakthrough came via Haploid Inducer Lines. By using CRISPR/Cas9 to knock out specific genes—specifically MTL (Matrilineal) and CENH3 (Centromere Histone)—we create plants that, when crossed with an elite variety, trigger the elimination of their own chromosomes in the resulting seed. This leaves only the elite parent’s DNA, creating a haploid naturally in the field and effectively solving the Indica rice “impasse.”
Methodological Divergence: In Vitro vs. In Vivo Systems
The transition from in vitro to in vivo methodology is dictated by the biological limitations of indica gametic embryogenesis. Traditional methods require reprogramming a microspore to an embryo in an artificial environment, a process that is highly genotype-dependent and prone to failure across most recalcitrant species, including watermelon and indica rice.
| Feature | In Vitro (Anther/Microspore Culture) | In Vivo (Inducer-Mediated) Induction |
|---|---|---|
| Genotype Dependency | High; limited to specific responsive lines | Low; universal trigger for recalcitrant lines |
| Tissue Culture Requirement | Total; requires specialized lab facilities | Minimal; primarily field-based pollination |
| Risk of Albinism | High; major barrier in indica rice | None; bypasses plastid genome issues |
| Technical Complexity | Operational: Requires constant lab maintenance | Front-Loaded: Initial engineering only |
| Scalability for Indica | Poor; hindered by albinism and recalcitrance | High; democratized for tropical breeding stations |
The biological “why” behind indica’s failure in vitro is the timing of plastid development. Indica varieties exhibit a faster transition from proplastids to amyloplasts compared to japonica. This rapid differentiation “locks” the cells into a state where they cannot revert to green chloroplasts during regeneration, leading to high rates of albinism and non-photosynthetic progeny. In vivo systems bypass this plastid bottleneck entirely by utilizing natural fertilization followed by selective chromosome elimination. The success of these systems relies on precisely engineered genetic triggers within the inducer’s genome.
2.2 The Transformation: How a Haploid Becomes “Doubled.”
The magic of the DH process is the transition from a single-set (n) sterile plant to a double-set (2n) fertile, homozygous plant. In the lab, we deploy anti-microtubular agents—also known as chromosome-doubling agents—to arrest the cell cycle. Chemicals like Colchicine (the traditional gold standard) or Oryzalin (a powerful alternative highlighted in recent rice studies) interfere with spindle fiber formation. This prevents the cell from splitting during division, effectively doubling the DNA within a single nucleus.
The Two Pathways to Doubling:
- Spontaneous Doubling: This occurs naturally through endomitosis (chromosome duplication without nuclear division). In rice, we see this natural “gift” in 30–40% of cases, where the plant repairs its own ploidy without our intervention.
- Induced Doubling: This is our active laboratory intervention. By applying doubling agents, we force the genetic material to replicate, creating two identical sets of chromosomes from the original single set.
The “So What?” of this transformation is profound: we reach 100% homozygosity in one generation, compressing a decade of traditional breeding into roughly 12 to 24 months. To trigger this doubling, however, we must first master the art of inducing the haploid state itself.
2.3 Strategic Roadmap: Transitioning to Inducer-Led Breeding Workflows
Moving from laboratory CRISPR engineering to a field-ready indica pipeline requires a phased implementation:
- Step 0: Pre-requisite Engineering: Develop the initial CRISPR knockouts in tissue-culture-responsive “model” genotypes (e.g., specific japonica or responsive indica lines) to serve as universal donors.
- Inducer Line Engineering: Validate the HIR and ensure the stability of the OsMTL or CENH3 knockout line.
- Intraspecific Crossing: Use the engineered line as the pollen parent to fertilize elite, recalcitrant indica cultivars.
- Haploid Identification: Screen seeds to isolate maternal haploids (carrying only the elite parent’s genome).
- Chromosome Doubling: Restore fertility and 100% homozygosity through spontaneous doubling or agents like colchicine.
The use of CRISPR/Cas9 is the legal and technical cornerstone of this roadmap. The resulting DH progeny are null-segregants; they contain no traces of the CRISPR transgene nor any part of the modified inducer genome. Because the final seeds are genetically identical to the elite parent—minus the heterozygosity—they should be exempt from GMO classifications in many jurisdictions, significantly lowering the regulatory barrier for new variety releases.
2.4 Overcoming the Final Hurdles: Albinism and Ploidy
The final stage of the journey often presents the most difficult technical barriers, particularly for indica researchers:
- Albinism: This is the regeneration of white shoots that lack chlorophyll. In rice, this is caused by deletions in the plastid genome and a premature cellular transition where proplastids differentiate into amyloplasts (starch storage) rather than chloroplasts. This makes the plants non-functional as they cannot photosynthesize.
- Ploidy Validation: Because regenerants can be haploid, diploid, or mixoploid, we must verify their ploidy.
2.5 The Laboratory Journey: A Step-by-Step Process
The transition from a donor plant to a fertile, homozygous green plantlet follows a rigorous protocol:
- Donor Plant Cultivation: Growth under high solar radiation (>18 MJ m⁻²) and specific day/night temperatures (34°C/24°C) is crucial.
- Student Insight: Field-grown plants are generally superior to greenhouse plants because these natural temperature fluctuations and light intensities significantly boost embryogenic potential.
- Panicle Harvest at Booting: Maturity check using morphological markers.
- Student Insight: Since maturity varies along the inflorescence, ensure your samples represent the entire panicle to maximize your P-grain yield.
- Surface Disinfection & Cold Pre-treatment: The primary trigger to initiate the androgenic switch.
- Student Insight: This “shock” is the most critical step; without it, the microspores will simply commit to their natural path of becoming sterile pollen.
- Anther Excision and Inoculation: Placing anthers on induction medium (e.g., N6 or MS).
- Student Insight: The precise balance of auxins and cytokinins here determines whether you get a mass of undifferentiated callus or organized embryogenic structures.
- Callus Formation: Initial growth from the reprogrammed microspores.
- Student Insight: Careful observation is needed to ensure the callus originates from the gametic microspores rather than the somatic tissues of the anther wall.
- Regeneration & Chromosome Doubling: Moving callus to a regeneration medium to produce green shoots.
- Student Insight: While spontaneous doubling is common in rice (30–40%), we often use anti-microtubular agents like colchicine to ensure the resulting plants are fertile diploids rather than sterile haploids.
3. Critical Success Factors in In Vitro Culture
Achieving commercial-scale haploid induction requires managing a multidimensional matrix of environmental and hereditary variables. Success is not a “recipe” but a strategic alignment of the donor plant’s physiological state with laboratory conditions.
3.1 Genotype Dependency
Strategic analysis of current protocols confirms that genetics account for over 90% of androgenic variability. The severity of this dependency is best illustrated in rice: while the japonica variety ‘Taipei 309’ can achieve a 41.9% callus induction rate, the indica variety ‘Suweon 290’ often yields a 0% response under identical conditions. For a Strategist, this necessitates an early assessment of “Tissue Culture Competence” before committing resources to a specific germplasm.
3.2 Microspore Maturity
The “uni-nucleate” to “early bi-nucleate” stage represents the critical window of developmental plasticity. Because cytological staining is too slow for high-throughput labs, breeders must utilize morphological markers: internode distance in rice or precise bud size (± 0.25 mm) in Brassicaceae are the standard indicators of readiness for sporophytic reprogramming.
3.3 Stress Pre-treatments
Stress is the mandatory trigger for the sporophytic switch. It disrupts the normal gametophytic cycle and induces cytoplasmic attenuation.
- Cold Stress: Treating panicles at 10°C for 7–10 days is the standard for rice.
- Heat Shock: In watermelon, incubation at 32–35°C for 2–20 days is often required to initiate the developmental shift.
3.4 Media and Growth Regulators
The selection of nutrient media (MS or N6) is secondary to the strategic management of hormone ratios. The “So What?” of utilizing Auxins (2,4-D/NAA) versus Cytokinins (BAP/Kinetin/TDZ) is the redirection of metabolic flux. In recalcitrant genotypes, this balance lies between stalled callus growth and the successful induction of embryogenic structures.
4. Overcoming Recalcitrance: Case Studies in High-Value Crops
The “Recalcitrance Impasse” is the primary barrier to ROI in biotechnology-led breeding. Overcoming it requires crop-specific procedural modifications.
4.1 Indica Rice: The Albinism Barrier
Indica rice frequently fails due to poor callus induction and high rates of albinism, often linked to deletions in the plastid genome. From an innovation standpoint, when in vitro regeneration fails to produce green shoots, the shift toward in vivo induction (Section 5) is mandatory to bypass tissue culture altogether.
4.1.1 The “Stubborn” Seed: Why Your Favorite Rice is Hard to Hack
While the technology is transformative, not all plants cooperate. Scientists refer to this resistance as recalcitrance.
Barriers to In Vitro Success in Indica Rice
| Factor | Impact on Breeding Efficiency |
|---|---|
| Genotype Dependency | Success is highly specific to certain lines; many elite indica cultivars show 0% callus induction compared to high rates in japonica (e.g., ‘Taipei 309’). |
| Albinism | High frequency of chlorophyll-deficient (albino) regenerants, often attributed to deletions in the plastid genome, preventing viable plant recovery. |
| Low Recovery Rates | Poor morphogenesis and regeneration during the callus phase lead to insufficient numbers of doubled haploids for commercial breeding. |
| Recalcitrance to Reprogramming | Difficulty in switching microspores from the gametophytic to the sporophytic development pathway in indica backgrounds. |
A prime example is Indica rice, the staple food for more than half the world’s population. While its cousin, Japonica rice, responds well to laboratory (in vitro) breeding, Indica rice has proven notoriously difficult to “hack.” Historically, breeders tried to overcome this by intersubspecific hybridization (crossing Indica with Japonica), but the pure lines remained resistant. Efforts to produce doubled haploids in Indica rice have been stifled by two major failures:
- Poor Callus Induction: The plant tissue refuses to revert to the “blank slate” state required for regeneration.
- Albinism: A frustrating phenomenon where the lab-grown rice produces white shoots lacking chlorophyll, rendering the plants non-viable.
This recalcitrance is a major bottleneck. If we cannot efficiently apply DH technology to the rice that feeds billions, global “zero hunger” goals remain at high risk.
4.2 Watermelon (Citrullus lanatus L.) Protocols
Effective watermelon protocols require high osmotic pressure and specific hormonal triggers. Analysis shows the highest callus induction using MS medium supplemented with 2.0 mg L⁻¹ 2,4-D and 90.0 g L⁻¹ sucrose. To ensure breeding purity, ploidy analysis via Flow Cytometry is a non-negotiable step to confirm haploid status before proceeding with chromosome doubling.
4.3 Brassicaceae: The “Anther-as-Sieve” Innovation
For poorly responsive Brassicaceae, the transition from standard destructive isolation to a “New Isolation Method” (transverse bud cutting and shaking) is transformative.
- Biological Mechanism: The anther tissue acts as a “biological sieve.” By not destroying the anther, larger late-stage microspores and tightly bound early-stage microspores remain trapped in the tissue. This results in a preparation concentrated with the desired mid-stage embryogenic microspores.
- Outcome: This method minimizes the release of somatic toxins, increasing embryo yield by up to 7.5 times in species like mustard.
4.3.1 The Bud Hack: Why Scissors are Mightier than the Blender
In the family Brassicaceae (cabbage, mustard, and rapeseed), scientists have discovered that the secret to success isn’t more power—it’s more precision.
Traditional methods for isolating microspores (the cells that become pollen) involve “destroying” plant buds in a blender or using magnets to grind them down. For poorly responsive genotypes, this mechanical trauma is lethal.
The new “Bud Cutting” technique offers a surgical alternative. Instead of the blender, researchers use a scalpel to individually cut buds in half. These halved buds are then gently shaken on a rotary shaker to release the cells.
The Success Story: In mustard plants, this hack increased embryo yield by 7.5 times. Even more impressively, it allowed scientists to obtain embryos from red cabbage for the first time.
The Result: This “gentle” method minimizes somatic cell destruction and reduces the release of toxins from damaged tissue.
5. The New Frontier: In Vivo Induction and Gene Editing
A strategic shift is underway to circumvent tissue culture requirements through in vivo systems. By using CRISPR/Cas9 to mutate specific centromere and pollen genes, we can create “Inducer Lines” that produce haploid seeds via standard crossing.
5.1 CENH3 and OsMTL Mutation
The next frontier is moving out of the petri dish and into the field. This is the in vivo revolution, where scientists use gene editing to turn a plant’s own reproductive system into a haploid-producing machine. This is the ultimate solution for “stubborn” crops because it completely removes the requirement for tissue culture competence.
The “master switches” for this hack are the CENH3 and MTL genes:
- CENH3 (The Centromere Switch):
- This protein is the anchor for chromosome segregation. Modifying CENH3 creates a “corrupted GPS” effect. When an inducer line is crossed with an elite variety, the inducer’s chromosomes get “lost” during the first cell divisions, leaving behind only the elite parent’s DNA.
The OsCENH3 (Centromere Histone H3) gene provides a strategic, albeit complex, pathway for uniparental genome elimination. OsCENH3 is a universal eukaryotic target responsible for directing the assembly of the kinetochore—the protein structure that attaches chromosomes to the spindle during division. Manipulating this locus allows for the creation of inducer lines that, when crossed with elite wild-type lines, lead to the systematic elimination of the inducer’s own chromosomes from the hybrid embryo.
Structural targets within the OsCENH3 locus are divided into two distinct regions for CRISPR/Cas9 intervention:
- N-terminal Tail: A hypervariable region where amino acid substitutions have been shown to induce haploidy (up to 1% HIR in rice).
- Histone Fold Domain (HFD): A conserved C-terminal region where in-frame deletions or point mutations can impair the loading of the CENH3 protein onto the centromere.
The “So What?” of centromeric dysfunction is the physical elimination of chromosomes. The mechanism is sophisticated: even when the OsCENH3 gene is transcribed, depleted or altered CENH3 activity prevents the protein from being incorporated into the centromeres of the inducer’s chromosomes. During the first mitotic divisions of the hybrid embryo, these “centromere-inactive” chromosomes fail to interact with spindle fibers. Consequently, they lag during division, form micronuclei, and are eventually degraded. The resulting embryo possesses only the chromosomes of the wild-type parent.
Current success rates for CENH3-based induction in rice remain lower than in model systems like Arabidopsis, yielding an HIR of approximately 1%. Despite this, the universal nature of CENH3 makes it a vital tool for genome elimination strategies across species.
The decision between these two targets requires a comparative analysis of their reliability and ease of implementation within a commercial breeding program.
- OsMTL (The Pollen Switch):
- In rice, the OsMTL gene encodes a patatin-like phospholipase. By knocking out this gene with CRISPR/Cas9, scientists create “inducer” lines that naturally trigger the formation of maternal haploids during pollination.
The OsMTL gene, located on the qhir1 QTL of rice chromosome 3, is the rice ortholog of the maize ZmMTL (also known as ZmPLA1 or NLD). In rice, this gene is specifically identified as OspPLAIIψ or PLP1. Strategically, OsMTL is a premier target for creating maternal haploid inducer lines because it is a pollen-specific gene. Its manipulation specifically affects the male gamete’s contribution to fertilization without compromising the overall vegetative health or female fertility of the inducer line.
At the molecular level, OsMTL encodes a patatin-like phospholipase protein. Using CRISPR/Cas9, researchers can induce a precise 4-bp insertion at the carboxy-terminal of the fourth exon. This edit is designed to mimic the natural mutations found in high-performing maize inducer lines like “Stock6,” which have served as the backbone for maize DH technology for decades.
The “So What?” of this mutation lies in its functional consequence: a frame-shift mutation that results in a truncated protein. In wild-type rice, the full-length protein is localized in mature pollen membranes and is essential for normal fertilization. The truncated version fails to localize correctly. This mis-localization prevents the male gamete from successfully completing fertilization of the egg cell. When this “defective” pollen is used to pollinate a wild-type female, the failure in gamete fusion triggers the development of the egg cell into a maternal haploid embryo.
Observed performance metrics for OsMTL (OspPLAIIψ) mutants demonstrate their viability:
- Selfed Populations: Observed Haploid Induction Rates (HIR) of approximately 6%.
- Out-crossed Populations: HIR typically ranging between 2% and 5%.
- Genotypic Stability: The system has been successfully streamlined in indica lines such as IR58025B, providing a predictable method for generating haploids.
While OsMTL focuses on fertilization failure, an alternative strategy targets the machinery of the cell itself to eliminate entire genomes through centromeric dysfunction.
Strategic Comparison of Genetic Targets
| Feature | OsMTL (OspPLAIIψ / PLP1) | OsCENH3 (Centromere Histone H3) |
|---|---|---|
| QTL / Locus | qhir1 | Universal Eukaryotic Target |
| Primary Mutation Type | 4-bp Insertion / Frame-shift | Amino Acid Substitution / In-frame Deletion |
| Functional Domain | Carboxy-terminal (Exon 4) | Hypervariable N-tail or Conserved HFD |
| Mechanism of Action | Fertilization failure (pollen-specific) | Post-zygotic chromosome elimination via micronuclei |
| Success Rate (HIR) | 2% – 6% | Approx. 1% |
| Conservation | Highly conserved specifically among monocots/cereals | Universal in eukaryotes |
1. Transferability and Efficiency of Cereal-Based Systems
The OsMTL target is currently the most pragmatic choice for indica rice improvement. Because it is highly conserved among cereals, the successful mechanisms identified in maize translate with high fidelity to rice. This provides significantly higher induction rates (up to 6%) compared to current CENH3 systems, making it the superior candidate for immediate integration into breeding pipelines.
2. Versatility vs. Complexity
While OsCENH3 modification is technically more complex and currently offers lower induction rates, its universal mechanism makes it a valuable alternative for wider genome elimination strategies. However, the requirement for precise tuning of centromeric loading—to maintain inducer fertility while ensuring enough dysfunction to trigger elimination—remains a technical friction point compared to the more straightforward fertilization-failure mechanism of OsMTL.
The successful development of these lines moves the conversation from the laboratory to the regulatory and agricultural landscape.
5.2 The Regulatory Advantage
From a strategic perspective, the “So What?” of in vivo induction is its potential regulatory status. According to the research on indica rice, the final progeny derived from crossing with CRISPR/Cas9 parents “should be exempt from GMO classification as they do not possess the altered genomes.” Because the inducer’s edited chromosomes are eliminated during the first zygotic divisions, the resulting DH plants are biologically indistinguishable from natural homozygous lines.
6. The GMO Loophole: CRISPR Without the Baggage
One of the most powerful benefits of in vivo haploid induction is its “clean” genetic result. While the “parent” inducer line is created using CRISPR/Cas9, the final product—the haploid progeny—is non-transgenic.
The Logic: The genome-edited “inducer” plant is used only as a biological trigger. Because the inducer’s chromosomes are physically eliminated during the development of the embryo, the resulting plant is chromosomally clean, containing only the DNA of the elite, non-edited parent.
This creates a significant regulatory advantage: we can use advanced genomic hacks to skip a decade of breeding, yet the final crop on the consumer’s plate contains no foreign DNA or edited sequences.
7. Conclusion: The Future of Accelerated Breeding
Doubled Haploid technology is no longer a luxury for elite research; it is a baseline requirement for competitive breeding. The convergence of refined in vitro protocols for Brassicaceae and the revolutionary gene-edited in vivo systems for rice represents a “path of least resistance” to commercial release.
Actionable Next Steps for Breeding Programs:
- Conduct an Androgenic Competence Audit: Use morphological markers (internode distance) to categorize elite lines by their responsiveness to existing in vitro protocols.
- Optimize Brassicaceae Isolation: Implement the “cutting and shaking” method to exploit the “sieve effect” for high-purity microspore preparations.
- Establish Flow Cytometry Pipelines: Standardize ploidy verification at the callus stage to eliminate somatic diploids early in the cycle.
- Prototype In Vivo Systems: Invest in CRISPR-mediated OsMTL inducer lines specifically for indica germplasm to bypass the albinism bottlenecks of anther culture.
Image Summary



Questions/Answers
1. How do in vivo and in vitro haploid induction differ?
In vivo and in vitro haploid induction differ primarily in the environment where the induction occurs, the specific biological pathways triggered, and their susceptibility to genetic and technical constraints.
Environmental and Procedural Differences
- In Vitro Induction: This approach involves gametic embryogenesis conducted in a sterile, controlled laboratory environment using tissue culture techniques. Common pathways include anther culture (aseptic inoculation of anthers), microspore culture (isolation and culture of immature pollen), and gynogenesis (culture of unpollinated ovaries or ovules).
- In Vivo Induction: This occurs within the living plant or under field conditions through natural or manipulated biological processes. Key methods include wide hybridization (crosses between different species or genera), the use of haploid inducer lines (genotypes carrying specific mutations like mtl or cenh3), pollen induction via physical or chemical agents, and centromere-mediated genome elimination.
Genotype Dependency and Success Rates
- Genotype Dependence: In vitro methods are often highly genotype-dependent. For example, in rice, anther culture is well-established for japonica varieties but remains highly recalcitrant for indica types. In vivo wide hybridization is generally considered less genotype-dependent, making it more universal across various cultivars of a species.
- Tissue Culture Competence: A significant advantage of in vivo systems is that they do not require tissue culture competence. This makes them potentially transformative for research on crops that are difficult to regenerate in vitro, such as certain legumes and woody plants.
Technical Challenges and Albinism
- Albinism: A major bottleneck for in vitro induction, particularly in cereals like rice and wheat, is the frequent regeneration of albino plants (chlorophyll-deficient shoots), which cannot survive. In vivo methods, such as those involving wide crosses or inducer lines, do not suffer from albinism, resulting in higher yields of viable green plants.
- Complexity: In vitro techniques require specialized equipment, sterile facilities, and precise manipulation of factors like nutrients, hormones, and temperature. In vivo methods rely more on standard agricultural practices like controlled pollination, though they may still require an embryo rescue step (an in vitro procedure) to preserve the developing haploid embryo.
Emerging Genetic Tools
While anther culture remains the predominant method for many crops, recent advances in CRISPR/Cas9 gene editing have enabled the creation of new in vivo inducer lines. By knocking out genes like MATRILINEAL (MTL) or modifying CENH3, researchers can create plants that naturally induce haploidy in their offspring when crossed, providing an efficient alternative to traditional tissue culture pathways.
2. How do researchers identify effective haploid inducer lines?
Researchers identify effective haploid inducer lines by analyzing natural spontaneous mutations, performing genetic and quantitative trait locus (QTL) mapping to locate induction triggers, and utilizing molecular markers and ploidy analysis to verify the induction rates of candidate genotypes.
1. Identification of Natural and Spontaneous Mutants
Effective inducer lines often originate from the identification of rare spontaneous mutations in existing germplasm.
- Maize Inducers: The maize line ‘Stock6’ was identified as a natural mutant with a maternal haploid induction rate (HIR) of approximately 3%. Similarly, the line Wisconsin-23 (W23) was identified as a paternal inducer due to a spontaneous mutation in the ig1 (indeterminate gametophyte 1) gene.
- Germplasm Screening: Researchers screen diverse germplasm and local cultivars for genotypes that naturally produce a higher-than-average frequency of haploid offspring when crossed.
2. Genetic and QTL Mapping
To understand and improve induction efficiency, researchers use Quantitative Trait Locus (QTL) analysis to pinpoint the specific genomic regions responsible for the trait.
- Major Loci: In maize, genetic mapping identified two primary QTLs: qhir1 (accounting for 66% of genetic variation) and qhir8 (accounting for 20%).
- Locus Dissection: Through fine mapping, researchers identified specific genes within these QTLs that govern induction: MATRILINEAL (MTL) on qhir1 and DOMAIN OF UNKNOWN FUNCTION 679 MEMBRANE PROTEIN (DMP) on qhir8.
- Subspecies Variation: In rice, researchers use doubled haploid (DH) and recombinant inbred line populations to map QTLs associated with induction efficiency, identifying specific additive and epistatic loci.
3. Molecular and Sequence Analysis
Once loci are identified, researchers compare sequences between inducer and non-inducer lines to find functional mutations.
- Frameshift Mutations: Analysis of the MTL gene revealed that a 4-bp insertion at the carboxy-terminal of the fourth exon causes a frameshift mutation, leading to a truncated protein that triggers haploid induction.
- CENH3 Variants: Researchers identify effective inducers by looking for variants in the CENH3 (centromere-specific histone H3) protein. Mutations in either the conserved C-terminal domain or the hypervariable N-terminal tail can weaken centromere function, leading to genome elimination and haploid formation upon outcrossing.
4. Verification of Induction Effectiveness
After identifying or creating a potential inducer line, researchers must rigorously verify its effectiveness through progeny testing.
- Ploidy Analysis: Researchers use flow cytometry to rapidly screen large populations of offspring from the candidate inducer. This is often supplemented by chromosome counting in root-tip cells to provide definitive proof of haploidy.
- Molecular Marker Validation: Simple Sequence Repeat (SSR) markers are used to confirm whether the offspring are truly haploid-derived and homozygous (possessing only one parent’s alleles) rather than being products of somatic mutations or unintended self-pollination.
- Morphological Selection: Researchers observe the offspring for characteristics typical of haploids, such as reduced vigor, dwarfism, or smaller leaves, which provide an early indication of successful induction.
5. Artificial Creation via Gene Editing
Modern researchers now identify effective “trigger” genes in model species and then target orthologs (functionally similar genes) in recalcitrant crops using CRISPR/Cas9.
- Targeting Orthologs: Researchers have successfully created new inducer lines in crops like rice and wheat by knocking out the OsMTL or TaMTL genes based on their knowledge of the maize MTL gene.
- CENH3 Engineering: By modifying the CENH3 gene, researchers can artificially engineer lines that selectivey eliminate their own chromosomes during fertilization, thereby inducing haploidy in the resulting embryo.
3. How can gene editing improve in vivo haploid induction?
Gene editing, particularly through the CRISPR/Cas9 system, improves in vivo haploid induction by allowing for the precise creation and optimization of haploid inducer lines. By targeting specific conserved genes, researchers can bypass the technical constraints and genotype dependencies of traditional in vitro tissue culture methods.
Creation of Novel Inducer Lines
Gene editing allows for the creation of inducer lines by knocking out or modifying specific genes responsible for normal fertilization and chromosome segregation:
- MATRILINEAL (MTL): CRISPR/Cas9 has been used to knock out the MTL gene (also known as ZmPLA1 or NLD) in crops like maize, rice, and wheat. This triggers maternal haploid induction by causing a failure in paternal fertilization or chromosome elimination. For instance, triple-knockout mutations in wheat have achieved induction frequencies of up to 31.6%.
- CENH3 (Centromere Histone H3): Editing this gene causes centromere dysfunction. When a CENH3 mutant is crossed with a wild-type plant, its “weak” centromeres lead to the selective elimination of non-parental chromosomes in the hybrid embryo, producing haploids. This has been successfully implemented in Arabidopsis, maize, and wheat.
- DMP (Domain of Unknown Function 679 Membrane Protein): Mutations in the DMP gene have been shown to enhance the induction efficiency of MTL mutants by five- to six-fold when both mutations are present together.
Overcoming Technical Bottlenecks
- Recalcitrance: A major bottleneck for in vitro methods is genotype dependence; for example, indica rice is highly recalcitrant to anther culture. Gene editing circumvents this by creating in vivo inducer lines that can be crossed with any elite cultivar, regardless of its tissue culture competence.
- Eliminating Albinism: Unlike in vitro induction, which frequently produces non-viable albino plantlets in cereals like rice and wheat, in vivo induction via gene-edited lines does not suffer from this issue, resulting in higher yields of viable green plants.
- Accelerated Breeding Cycles: While traditional anther culture might require three years to develop a mapping population, gene-edited in vivo systems can expedite this process to 2–3 years within controlled environmental systems.
Advanced Breeding Applications
- HI-Edit (Transgene-Free Editing): This strategy uses a gene-edited haploid inducer to deliver CRISPR components via pollen to a recipient’s egg cell. During the subsequent genome elimination, the inducer’s chromosomes (and the CRISPR transgene) are lost, leaving behind a haploid embryo that is edited but transgene-free.
- Integration with Speed Breeding: By combining gene-edited inducer lines with “speed breeding” (manipulating light and temperature), researchers can stack multiple edited traits into homozygous lines in just one generation.
- Regulatory Advantages: It has been argued that the progeny of gene-edited haploid-inducing parents should be exempt from GMO classification because the resulting haploid plant consists only of chromosomes from the non-inducer parent and does not contain the altered inducer genome.
4. Are there ways to overcome albinism in plant breeding?
Albinism—the regeneration of chlorophyll-deficient plants—is a major bottleneck in plant breeding, particularly during in vitro haploid induction methods like anther and microspore culture. Since albino plantlets are incapable of photosynthesis, they cannot survive to produce seed.
Breeders use several strategies to overcome or mitigate this phenomenon, ranging from biochemical modifications to alternative induction pathways.
1. Optimization of Culture Media and Chemicals
Adjusting the chemical composition of the induction and regeneration media is one of the most common ways to reduce albinism rates.
- Carbon Source Substitution: Replacing sucrose with maltose in the induction medium has been shown to significantly reduce the frequency of albino plants in rice and other cereals.
- Nutrient and Hormone Ratios: Reducing the concentrations of inorganic salts and manganese in the induction medium can lower albinism frequency. Additionally, optimizing the ratio of plant growth regulators, such as 2,4-D and Kinetin (KT), helps improve green plant differentiation.
- Supplementing with Metal Ions: Adding copper sulfate (CuSO4) to the induction medium can increase the number of green plantlets and decrease albinos by improving microspore survival and synchronization. Silver nitrate (AgNO3) and silver thiosulphate have also been utilized to enhance green plantlet regeneration.
- Synthetic Polymers and Additives: The use of Ficoll (a synthetic polymer of sucrose) as a gelling agent enhances the achievement of green plants over albinos. Adding Dimethyl Sulfoxide (DMSO) to mannitol pretreatment media has also yielded higher numbers of green plantlets.
2. Refined Procedural and Environmental Controls
Precise control over the timing of laboratory procedures can prevent the physiological changes that lead to albinism.
- Pretreatment Timing: While cold stress is necessary to trigger haploid induction, prolonged cold pretreatment (longer than 11 days in rice or 6–8 weeks in oats) can lead to a marked increase in albino regenerants.
- Incubation Conditions: Research in oats and wheat suggests that incubating embryo-like structures (ELS) in darkness for the first two weeks of the regeneration phase can improve the frequency of green plants.
- Epigenetic Modifiers: Applying inhibitors like Trichostatin A (TSA), which affects histone acetylation and chromatin accessibility, has been shown to increase the yield of green wheat plantlets.
3. Genetic Selection and Crossing Strategies
Since albinism is a highly genotype-dependent recessive trait, breeders can address it through genetic selection.
- Responsive Genotype Selection: Breeders can select for “amenable” genotypes that naturally have lower albinism rates; for example, winter barley is generally less prone to albinism than spring wheat due to evolutionary adaptations to low-temperature stress.
- Intersubspecific Hybridization: Crossing recalcitrant cultivars (like indica rice) with highly responsive varieties (like japonica rice) can introduce favorable alleles that improve the predisposition for successful green plant regeneration.
- Marker-Assisted Selection (MAS): Identifying Quantitative Trait Loci (QTLs) associated with albino differentiation—such as a major QTL found on chromosome 9 in rice—allows breeders to use MAS or gene editing to target and reduce albinism.
4. Utilizing In Vivo Induction Systems
The most definitive way to overcome albinism is to bypass the in vitro tissue culture phase entirely. In vivo induction methods, such as wide hybridization or the use of haploid inducer lines (e.g., mtl or cenh3 mutants), do not suffer from albinism. These systems occur within the living plant or under field conditions, resulting in high yields of viable green plants even from species that are otherwise recalcitrant to tissue culture.
5. How can gene editing technologies advance rice breeding techniques?
Gene editing technologies, particularly CRISPR/Cas9, significantly advance rice breeding by shortening development cycles, creating efficient haploid induction systems, and enabling the fixation of hybrid vigor.
Accelerating Breeding and Homozygosity
Traditional rice breeding requires 8 to 10 generations of selfing to achieve genetically stable homozygous lines. Gene editing combined with doubled haploid (DH) technology can reduce this process to just one generation. By leveraging CRISPR/Cas9, breeders can bypass labor-intensive procedures and low efficiency in selecting elite cultivars.
Development of In Vivo Haploid Inducer Lines
A transformative advancement is the creation of in vivo haploid inducer lines.
- MTL Gene Editing: Knocking out the rice ortholog of the MATRILINEAL (MTL) gene (also known as OsMATL or OspPLAIIψ) using CRISPR/Cas9 enables the production of maternal haploids. These inducer lines, when crossed with elite cultivars, produce offspring with the haploid chromosome complement of the elite parent, which can then be doubled to create fertile, homozygous lines.
- Indica Rice Improvement: This is especially critical for indica rice, which remains largely recalcitrant to traditional in vitro anther culture methods. Because in vivo induction does not require tissue culture competence, it offers a pragmatic approach to improving these cultivars.
Fixing Heterosis for Clonal Seed Production
Gene editing allows for the creation of clonal seeds from hybrid rice. By simultaneously editing the OsMTL gene and three genes crucial for meiosis (the MiMe technology), researchers can replace meiosis with mitosis-like division. This process results in diploid clonal gametes that retain the hybrid’s genetic makeup, effectively fixing heterosis and allowing hybrids to be propagated by seed without losing their superior traits.
Improving Tissue Culture Efficiency
For crops still using in vitro methods, gene editing can resolve technical bottlenecks:
- Fate Reprogramming: Research has shown that the BBM-BAR1 gene module can be used to reprogram the fate of microspores in rice anthers, significantly increasing the efficiency of asexual embryo formation.
- Overcoming Genotype Dependence: Future research aims to employ reverse genetics to edit or knockout genes that limit anther culturability, helping to achieve broader technical universality across different rice varieties.
Precision Trait Modification and Genetic Research
- Targeted Improvement: CRISPR/Cas9 provides precise control for the targeted modification of specific genes associated with yield, quality, and stress resistance.
- QTL Mapping: DH mapping populations generated through gene-edited inducer lines are invaluable for constructing genetic linkage maps and performing high-precision Quantitative Trait Locus (QTL) analysis.
- Speed Breeding Integration: Integrating gene editing with “speed breeding” (manipulating light and temperature) further accelerates trait modification and the release of new varieties.
Regulatory Advantages
It has been noted that the progeny of CRISPR-edited haploid-inducing parents may be exempt from GMO classification in some frameworks. Since the final haploid progeny consists only of chromosomes derived from the non-inducer parent, they do not possess the altered genome or the transgene.
6. How can gene editing improve in vivo haploid induction?
Gene editing, particularly the CRISPR/Cas9 system, improves in vivo haploid induction by allowing for the precise creation of haploid inducer lines through the modification of conserved genes essential for fertilization and chromosome segregation.
Creation of Targeted Inducer Lines
Gene editing enables researchers to bypass the need to find rare spontaneous mutants by artificially inducing haploid-triggering mutations in diverse germplasm:
- MATRILINEAL (MTL) Knockout: CRISPR/Cas9 is used to knock out the MTL gene (also known as PLA1 or NLD), which triggers maternal haploid induction in monocots. In wheat, triple-knockout mutations of TaMTL achieved haploid induction frequencies as high as 31.6%. In rice, editing the OsMTL gene has successfully produced inducer lines with induction rates between 2% and 6%.
- CENH3 Modification: The temporal alteration or excision of the centromere histone H3 (CENH3) protein disrupts centromere function. When these modified lines are outcrossed, their chromosomes are selectively eliminated during early embryogenesis, resulting in haploid progeny.
- DMP Enhancement: Mutations in the DMP (Domain of Unknown Function 679 Membrane Protein) gene have been shown to enhance the induction efficiency of mtl mutants by five- to six-fold when both modifications are present.
Overcoming Technical and Biological Barriers
- Bypassing Recalcitrance: A major advantage of gene-edited in vivo induction is that it does not require tissue culture competence. This is transformative for improving indica rice, which is highly recalcitrant to traditional in vitro anther culture.
- Eliminating Albinism: Unlike in vitro induction, which frequently results in non-viable albino plantlets in cereals like rice and wheat, in vivo induction via edited inducer lines produces viable green plants.
- Accelerated Timelines: Traditional breeding takes 8–10 generations to reach homozygosity, whereas gene-edited haploid induction combined with chromosome doubling achieves complete homozygosity in one generation. This can reduce the time required to develop doubled haploid mapping populations from three years to approximately two years.
Advanced Breeding Applications
- Transgene-Free Editing (IMGE/HI-Edit): This approach uses an edited haploid inducer to deliver CRISPR components to a recipient cell. As the inducer’s chromosomes (and the CRISPR transgene) are eliminated during the subsequent induction process, the resulting haploid embryo is edited but transgene-free. Progeny derived this way may be exempt from GMO classification in some jurisdictions because they do not contain the altered inducer genome.
- Fixing Heterosis: By simultaneously editing the OsMTL gene and genes governing meiosis (MiMe technology), researchers can replace meiosis with a mitosis-like division. This allows for the fixation of hybrid vigor, enabling hybrid rice to be propagated via clonal seeds without losing superior traits.
- Integration with Speed Breeding: Gene editing can be integrated with speed breeding (manipulating photoperiods and temperature) to further expedite the modification of traits and the stabilization of new varieties.
7. How do CRISPR/Cas9 systems improve in vivo haploid induction?
CRISPR/Cas9 systems improve in vivo haploid induction by allowing researchers to precisely create and optimize haploid inducer lines through the modification of conserved genes involved in fertilization and chromosome segregation. This technology bypasses the need for finding rare natural mutants and overcomes the technical recalcitrance of certain crop subspecies.
Creation of Targeted Inducer Lines
CRISPR/Cas9 is used to knock out or modify “trigger” genes that naturally induce haploidy when mutated:
- MATRILINEAL (MTL) Knockout: Researchers use CRISPR to knock out the MTL gene (also known as ZmPLA1 or NLD), a pollen-specific phospholipase. While naturally found in maize, CRISPR has enabled the creation of mtl inducer lines in rice, wheat, and foxtail millet. For instance, triple-knockout mutations in wheat achieved a haploid induction rate (HIR) of up to 31.6%.
- CENH3 Modification: By using CRISPR to modify the Centromere Histone H3 (CENH3) gene, researchers can create plants with “weak centromeres”. When these modified lines are outcrossed, their chromosomes are selectively eliminated in the hybrid embryo, resulting in haploid progeny. This has been successfully implemented in Arabidopsis, maize, and wheat.
- DMP Enhancement: CRISPR-mediated mutations in the DOMAIN OF UNKNOWN FUNCTION 679 MEMBRANE PROTEIN (DMP) gene can enhance the induction efficiency of mtl mutants by five- to six-fold when both modifications are present.
Bypassing Technical and Biological Barriers
- Overcoming Recalcitrance: In vitro methods like anther culture are often highly genotype-dependent; for example, indica rice is notoriously recalcitrant. CRISPR-edited in vivo inducer lines offer a pragmatic alternative because they do not require tissue culture competence, allowing them to be crossed with any elite cultivar to produce haploids.
- Eliminating Albinism: Unlike in vitro induction, which frequently results in non-viable albino plantlets in cereals, in vivo induction via CRISPR-edited lines does not suffer from this issue, resulting in higher yields of viable green plants.
- Accelerated Breeding Cycles: Traditional breeding takes 8–10 generations to reach homozygosity. CRISPR-edited in vivo systems, combined with chromosome doubling, can reduce the time required to develop doubled haploid (DH) mapping populations from three years to 2–3 years.
Advanced Breeding Strategies
- Transgene-Free Editing (HI-Edit/IMGE): This strategy uses a CRISPR-edited haploid inducer to deliver Cas9 and guide RNA to a recipient’s egg cell. During the subsequent genome elimination process, the inducer’s chromosomes (and the CRISPR transgene) are lost, leaving behind a haploid embryo that is edited but transgene-free. Such plants may be exempt from GMO classification in some jurisdictions because they do not contain the altered inducer genome.
- Fixing Heterosis: In rice, CRISPR has been used to simultaneously edit OsMTL and three genes crucial for meiosis (MiMe technology). This replaces meiosis with a mitosis-like division, creating diploid clonal gametes that retain hybrid vigor, effectively fixing heterosis for propagation via clonal seeds.
- Integration with Speed Breeding: CRISPR/Cas9 and in vivo induction can be integrated with speed breeding (manipulating light and temperature) to expedite trait modification and the stabilization of new varieties within controlled environments.
8. How do in vivo methods avoid the albinism problem?
In vivo methods avoid the albinism problem primarily by conducting haploid induction within the natural environment of the living plant, thereby bypassing the stressful in vitro tissue culture phase that disrupts plastid development.
The Nature of the Albinism Problem
Albinism is a phenomenon where regenerated plantlets are chlorophyll-deficient, appearing white or yellow, and are incapable of photosynthesis. It is the most significant limiting factor in in vitro induction methods like anther and microspore culture, particularly in cereals such as rice and wheat. This occurs because the laboratory environment—specifically the artificial media, hormone ratios, and prolonged stress treatments—can cause large-scale deletions or rearrangements in the plastid genome, preventing proplastids from developing into functional chloroplasts.
Why In Vivo Methods Avoid Albinism
- Natural Physiological Environment: In vivo techniques, such as wide hybridization (e.g., crossing wheat with maize or Imperata cylindrica), occur within the recipient plant’s embryo sac. Because the developing haploid embryo remains in its native biological setting, it is not subjected to the “epigenetic stress memory” or chemical imbalances of tissue culture that typically trigger albinism.
- Bypassing Tissue Culture Competence: A primary advantage of in vivo systems—including the use of haploid inducer lines (like maize mtl or cenh3 mutants)—is that they do not require tissue culture competence. While in vitro methods often fail for recalcitrant genotypes like indica rice by producing high frequencies of albinos, in vivo methods consistently yield viable green plants regardless of the cultivar’s tissue culture response.
- Stable Plastid Biogenesis: In in vivo genome elimination, the maternal chromosomes (and their associated plastids) are preserved in their natural state while the paternal chromosomes are eliminated. This avoids the developmental blockages in plastid transition that occur when microspores are forced to switch from a gametophytic to a sporophytic pathway under laboratory conditions.
Comparison of Methods
According to the sources, the presence of albinism differs sharply between the two induction pathways:
- In Vitro (Anther/Microspore Culture): Albinism is a frequent and serious drawback.
- In Vivo (Wide Hybridization/Ovary Culture): Albinism is generally absent, resulting in 100% green plantlet regeneration in many reported cases.
9. Can CRISPR make in vivo haploid induction more efficient?
CRISPR/Cas9 technology significantly improves the efficiency of in vivo haploid induction by enabling the precise creation and optimization of haploid inducer lines through the targeted modification of conserved genes,,,.
Creation of Targeted Inducer Lines
CRISPR/Cas9 allows researchers to bypass the reliance on rare natural mutants by artificially inducing haploid-triggering mutations in elite germplasm,,.
- MATRILINEAL (MTL) Knockout: Knocking out the MTL gene (also known as ZmPLA1 or NLD) using CRISPR has successfully triggered maternal haploid induction in maize, rice, and wheat,,. In wheat, triple-knockout mutations of TaMTL achieved induction frequencies as high as 31.6%. In rice, researchers have created inducer lines with an efficiency of 12.4%.
- CENH3 Manipulation: Editing the Centromere Histone H3 (CENH3) gene disrupts centromere function,,. When these modified “haploid inducer” lines are outcrossed with wild-type plants, their chromosomes are selectively eliminated during early embryogenesis, resulting in haploid progeny,,. This has been implemented in Arabidopsis, maize, and wheat,.
- DMP Enhancement: CRISPR-mediated mutations in the Domain of Unknown Function 679 Membrane Protein (DMP) gene can enhance the induction efficiency of MTL mutants by five- to six-fold when both modifications are present together.
Overcoming Biological and Technical Constraints
- Bypassing Recalcitrance: A major advantage of CRISPR-edited in vivo induction is that it does not require tissue culture competence,. This is transformative for indica rice, which remains largely recalcitrant to traditional in vitro anther culture methods,,.
- Eliminating Albinism: Unlike in vitro induction, which frequently results in non-viable albino plantlets in cereals, in vivo induction via gene-edited lines does not suffer from this issue, resulting in higher yields of viable green plants,.
- Expediting Breeding Cycles: Doubled haploid (DH) mapping populations, which traditionally require three years to generate via anther culture, can be expedited to 2–3 years using CRISPR-mediated in vivo systems within controlled environments,.
Advanced Precision Breeding
- IMGE/HI-Edit (Transgene-Free Editing): This strategy uses a CRISPR-edited haploid inducer to deliver genome-editing components to a recipient cell,. During the subsequent genome elimination process, the inducer’s chromosomes (and the CRISPR transgene) are lost, leaving behind a haploid embryo that is edited but transgene-free,.
- Fixing Heterosis: By simultaneously editing the OsMTL gene and genes governing meiosis (MiMe technology), CRISPR allows for the fixation of hybrid vigor, enabling hybrid rice to be propagated via clonal seeds without losing superior traits,.
- Integration with Speed Breeding: CRISPR/Cas9 can be integrated with speed breeding (manipulating light, temperature, and photoperiod) to further accelerate the production of DH lines and streamline the QTL mapping process in controlled environments,.
10. How do researchers identify candidate genes for haploid induction?
Researchers identify candidate genes for haploid induction through a combination of genetic mapping, multi-omics analysis, and the study of conserved biological pathways across species.
1. Genetic Mapping and QTL Analysis
The primary method for identifying candidate genes involves Quantitative Trait Locus (QTL) mapping to pinpoint genomic regions associated with haploid induction efficiency.
- Major Loci Identification: In maize, researchers located two critical QTLs: qhir1 (responsible for 66% of genetic variation) and qhir8 (accounting for 20%).
- Fine Mapping: Once broad QTLs are identified, fine mapping is used to narrow these regions down to specific genes. This process led to the identification of MATRILINEAL (MTL) (also known as ZmPLA1 or NLD) within the qhir1 locus and DOMAIN OF UNKNOWN FUNCTION 679 MEMBRANE PROTEIN (DMP) within qhir8.
- Anther Culture Efficiency: In rice, QTL analysis detected loci for callus induction on chromosomes 6, 7, 8, 10, and 12, as well as major loci for green and albino plant differentiation on chromosome 9.
2. Sequence Comparison and Mutational Analysis
Researchers identify candidate genes by comparing the genetic sequences of inducer lines versus non-inducer lines to find functional mutations.
- Functional Triggers: For the MTL gene, a 4-bp insertion at the carboxy-terminal was identified as the mutation causing a truncated protein, which triggers the haploid-inducing phenotype.
- Centromere Manipulation: Candidate genes like CENH3 (centromere-specific histone H3) are identified because modifications to their conserved domains lead to “weak centromeres” and subsequent chromosome elimination during fertilization.
3. Multi-Omics and Transcriptomic Analysis
Modern researchers utilize genomics, transcriptomics, and proteomics to identify genes that are upregulated or downregulated during the induction process.
- Fate Reprogramming: Transcriptomics and chromatin immunoprecipitation sequencing (ChIP-seq) were used in rice to identify the BBM-BAR1 gene module, which can reprogram microspores toward an embryogenic pathway.
- Trait Markers: RNA-seq data analysis has suggested that specific candidate genes, such as those encoding High Mobility Group (HMG) proteins, are associated with higher callus induction rates.
- Stress and Metabolism: Transcriptomic investigations in crops like Brassica napus have identified stress-associated genes, cell wall remodeling proteins, and hormone signaling pathways (such as auxin and gibberellin synthesis) that regulate microspore embryogenesis.
4. Reverse Genetics and Orthology
Because many induction genes are functionally conserved, researchers identify candidates by looking for orthologs in other species.
- Conserved Pathways: Since MTL is highly conserved among monocots and CENH3 is found in most eukaryotes, researchers target these genes in recalcitrant crops based on their known roles in model species like maize or Arabidopsis.
- Gene Knockouts: Future strategies involve using reverse genetics to edit or knockout genes with similar functions in other crops to construct mutant libraries and validate them through forward genetics.
5. Studying Specific Physiological Triggers
Researchers also look at genes involved in the plant’s physiological response to external stimuli used during induction.
- Photoperiod and Temperature: Studies suggest a correlation between induction frequency and the photoperiod response or hormonal status of the parent plants, leading researchers to examine genes regulating these environmental interactions.
- Ovary Conditioning: In wheat, the identification of candidate genes like FLA26 and FER was linked to their activation in ovary tissues, which act as “nurse factors” to stimulate microspore division.
11. What makes indica rice recalcitrant to in vitro induction?
The recalcitrance of indica rice to in vitro induction, particularly through anther culture, is primarily attributed to its inherent genetic background, which results in significantly lower efficiency compared to japonica subspecies.
1. Genetic and Hereditary Determinants
- Genotype Dependence: Genotype is considered the most critical internal determinant governing induction efficiency. Studies indicate that over 90% of the variability in anther culture response is due to genetic effects.
- Quantitative Trait Inheritance: Anther culture efficiency is a complex quantitative trait governed by the synergistic action of multiple genes.
- Low Induction and Regeneration Rates: Indica varieties typically exhibit callus induction rates below 5% (with a mean as low as 0.5%) and green plantlet differentiation rates averaging only 0.5%. In many cases, the response is 0% compared to japonica varieties that can exceed 40% efficiency.
- Discordant Morphogenesis: Genotypes that respond well to callus induction often fail during the subsequent stage of morphogenesis due to the distinct genetic factors controlling these two events.
2. Physiological and Biological Barriers
- Early Anther Necrosis: A significant constraint in indica rice is early anther wall necrosis or cell death, which occurs very early during culture and represses the pollen from forming calli.
- Recalcitrance to Tissue Culture Competence: Unlike japonica, indica rice has poor “tissue culture competence,” meaning it is fundamentally less able to undergo dedifferentiation and redifferentiation outside the living plant.
- Pollen Abortion: Failures are often caused by the early abortion of pollen under in vitro conditions.
3. The Albinism Bottleneck
- High Frequency of Albinos: Indica cultivars are significantly more prone than japonica to albinism—the regeneration of chlorophyll-deficient plantlets that cannot survive.
- Plastid Genome Deletions: This phenomenon in rice has been attributed to large-scale deletions and rearrangements in the plastid genome, preventing proplastids from developing into functional chloroplasts.
- Delayed Plastid Transition: Genotypes that fail to regenerate green plants often suffer from an accelerated or blocked transition of proplastids to amyloplasts during normal gametogenesis.
4. Sensitivity to Culture Environment
Carbon Source Preference: While japonica often responds well to sucrose, indica induction is generally more effective when maltose is used as the carbohydrate source, which satisfies different sugar metabolism requirements during development.
Nutritional Sensitivity: Indica rice is highly sensitive to the composition of culture media; for instance, it performs better when ammonium (NH4+) ion concentrations are reduced to half-strength or replaced by organic nitrogen sources.
Hormonal Imbalance: The direction of development is highly regulated by the balance of auxins and cytokinins, and indica varieties often require lower levels of 2,4-D because high concentrations, while inducing callus, hurt subsequent regeneration.
References
Arabzai, M. G., Huang, D., Mohammadi, N. K., Gao, J., Wang, X., Zheng, P., Qin, Y., & Wang, L. (2025). Techniques and advantages of microspore culture for crop improvement. In Plant Growth Regulation (Vol. 105, Number 4, pp. 903–918). Springer Science and Business Media B.V. https://doi.org/10.1007/s10725-025-01312-8
Bajpai, R., & Chaturvedi, R. (2023). In vitro production of doubled haploid plants in Camellia spp. and assessment of homozygosity using microsatellite markers. In Journal of Biotechnology (Vol. 361, pp. 89–98). Elsevier B.V. https://doi.org/10.1016/j.jbiotec.2022.11.019
Cardoso, J. C., Abdelgalel, A. M., Chiancone, B., Latado, R. R., Lain, O., Testolin, R., & Germanà, M. A. (2016). Gametic and somatic embryogenesis through in vitro anther culture of different Citrus genotypes. Plant Biosystems, 150(2), 304–312. https://doi.org/10.1080/11263504.2014.987847
Cardoso, J. C., Martinelli, A. P., Germanà, M. A., & Latado, R. R. (2014). In vitro anther culture of sweet orange (Citrus sinensis L. Osbeck) genotypes and of a C. clementina × C. sinensis “Hamlin” hybrid. Plant Cell, Tissue and Organ Culture, 117(3), 455–464. https://doi.org/10.1007/s11240-014-0456-x
Chen, X., Li, S., Zha, W., Li, C., Zhou, L., You, A., & Wu, Y. (2025). Advances in Anther Culture-Based Rice Breeding in China. In Plants (Vol. 14, Number 11). Multidisciplinary Digital Publishing Institute (MDPI). https://doi.org/10.3390/plants14111586
Eliby, S., Bekkuzhina, S., Kishchenko, O., Iskakova, G., Kylyshbayeva, G., Jatayev, S., Soole, K., Langridge, P., Borisjuk, N., & Shavrukov, Y. (2022). Developments and prospects for doubled haploid wheat. In Biotechnology Advances (Vol. 60). Elsevier Inc. https://doi.org/10.1016/j.biotechadv.2022.108007
Feng, Z., Chu, H., Cao, L., Wang, R., & Zhang, A. (2025). A Review of Research Progress in Rice Anther Culture. Current Issues in Molecular Biology, 48(1), 18. https://doi.org/10.3390/cimb48010018
Germanà, M. A. (2011). Anther culture for haploid and doubled haploid production. In Plant Cell, Tissue and Organ Culture (Vol. 104, Number 3, pp. 283–300). Kluwer Academic Publishers. https://doi.org/10.1007/s11240-010-9852-z
Grela, M., Czyczyło-Mysza, I., & Skrzypek, E. (2024). Crucial Factors Influencing the Efficiency of Androgenesis in Oat (Avena sativa L.) Through Anther and Microspore Cultures. In Agronomy (Vol. 14, Number 10). Multidisciplinary Digital Publishing Institute (MDPI). https://doi.org/10.3390/agronomy14102394
Islam, M. T., Arif, M. R., Hasan, M. T., & Robin, A. H. K. (2023). Anther Culture in Crop Plants: Progress and Perspectives. In Plant Breeding and Biotechnology (Vol. 11, Number 2, pp. 69–96). Korean Society of Breeding Science. https://doi.org/10.9787/PBB.2023.11.2.69
Jin, S. B., Kim, M. J., Choi, C. W., Park, S. M., & Yun, S. H. (2022). Anther Culture-Derived Haploids of Citrus aurantium L. (Sour Orange) and Genetic Verification of Haploid-Derived Regenerated Plants. Plants, 11(22). https://doi.org/10.3390/plants11223022
Kozar, E. V., Kozar, E. G., & Domblides, E. A. (2022). Effect of the Method of Microspore Isolation on the Efficiency of Isolated Microspore Culture In Vitro for Brassicaceae Family. Horticulturae, 8(10). https://doi.org/10.3390/horticulturae8100864
Mayakaduwa, D. M. R. G., & Silva, T. D. (2018). Anther Culture as a Supplementary Tool for Rice Breeding. In Rice Crop – Current Developments. InTech. https://doi.org/10.5772/intechopen.76157
Mayakaduwa, R., & Silva, T. (2023). Haploid Induction in Indica Rice: Exploring New Opportunities. In Plants (Vol. 12, Number 17). Multidisciplinary Digital Publishing Institute (MDPI). https://doi.org/10.3390/plants12173118
Nyirahabimana, F., Adıgüzel, P., & Solmaz, İ. (2025). Current state of anther and ovary cultures in watermelon haploid induction. In In Vitro Cellular and Developmental Biology – Plant. Springer. https://doi.org/10.1007/s11627-025-10584-8
Nyirahabimana, F., & Solmaz, İ. (2025). Prevailing situation of anther and ovary cultures in cucumber haploid induction. In Vegetos (Vol. 38, Number 4, pp. 1309–1317). Springer. https://doi.org/10.1007/s42535-024-01055-3
Oleszczuk, S., Podskarbi, M., & Michalski, K. (2025). First regeneration of doubled haploids via anther culture in Triticum timopheevii. Plant Cell, Tissue and Organ Culture, 163(1). https://doi.org/10.1007/s11240-025-03216-x
Ramlal, A., Mehta, S., Nautiyal, A., Baweja, P., Shivam, Sharma, D., Lal, S. K., Vijayan, R., Raju, D., Subramaniam, S., & Rajendran, A. (2024). Androgenesis in soybean (Glycine max (L.) Merr.): a critical revisit. In In Vitro Cellular and Developmental Biology – Plant (Vol. 60, Number 1, pp. 1–15). Springer. https://doi.org/10.1007/s11627-023-10402-z
Sharma, A., Pandey, H., Misra, V., Kumar, R., Vashishth, A., Devadas, V. S., Mall, A. K., Ashutosh, Raghvan, M., Kesavan, A. K., & Chaturvedi, V. D. (2024). CRISPR/Cas9 and Anther Culture for Precision Double Haploid Line Production in Controlled Environments. In Plant Breeding. John Wiley and Sons Inc. https://doi.org/10.1111/pbr.13227
Yaman, H., & Sanyürek, N. K. (2023). Obtaining haploid plants by irradiated pollen culture in oil seed crops. In Biotech Studies (Vol. 32, Number 1, pp. 10–16). Field Crops Central Research Institute. https://doi.org/10.38042/biotechstudies.1273824






