1. Strategic Introduction to Plant Chromosome Engineering

To address the critical imperatives of global food security, we must move beyond the legacy era of radiation-induced fragmentation and stochastic mutation breeding toward a regime of modular precision. Plant chromosome engineering now leverages CRISPR/Cas systems to transition from random genomic disruption to the “predesigned” orchestration of genetic variation. By inducing site-specific double-strand breaks (DSBs), we can overcome the constraints of natural green biotechnology by reorganizing megabase-scale architecture while maintaining essential genomic collinearity. This strategic shift allows us to move from simple gene editing to the construction of custom genomic platforms, beginning with the development of specialized cargo vectors.
1.1 The Paradigm Shift in Plant Breeding: From Randomness to Precision
To sustain global standards of living and ensure food security for a rising population, our R&D priority must shift from incremental gains to a fundamental overhaul of agricultural productivity. Historically, we have relied on “global mutation enhancement”—a brute-force approach pioneered by Sears (1956) using X-ray irradiation to fragment chromosomes and facilitate the random incorporation of foreign genetic material. While this “shotgun” methodology established the foundations of pre-breeding, it is no longer compatible with modern commercial timelines or precision requirements. The technological landscape now enables a transition to “site-specific chromosomal restructuring.” By leveraging programmable nucleases, specifically the CRISPR/Cas system, we are moving beyond simple gene edits to the architectural design of entire linkage groups, drastically reducing the R&D noise inherent in traditional mutation-based pipelines.
| Dimension | Traditional Pre-Breeding (X-ray Fragmentation) | Modern Engineering (CRISPR/Cas Precision) |
|---|---|---|
| Specificity | Random, genome-wide fragmentation | Site-specific, predesigned targeting |
| Heritability | Unpredictable; requires extensive backcrossing | High; precise changes are stabilized quickly |
| R&D Timelines | Long (years to decades to remove linkage drag) | Accelerated; direct targeting of genomic regions |
| Mechanism | Random DNA lesions via genotoxic agents | Controlled DSBs via programmable nucleases |
This strategic evolution from random fragmentation to genomic surgery necessitates a new architecture for trait delivery: the engineered minichromosome.

2. Engineering Plant Minichromosomes: Developing Next-Generation Cargo Vectors
Minichromosomes represent a top-tier platform for trait stacking and breeding acceleration, functioning as independent, phenotypically neutral vectors that segregate autonomously from the host genome. While “top-down” engineering of standard A chromosomes is possible, they are prone to deleterious aneuploidy and dosage imbalances. Consequently, supernumerary B chromosomes are the superior targets for truncation; they are non-essential by definition and, crucially, become “drive-negative” upon downsizing. While native B chromosomes exhibit “chromosome drive” (transmission >0.5), the Birchler laboratory demonstrated that telomere-mediated truncation in maize often eliminates distal genes required for this drive, resulting in stabilized, predictable inheritance.
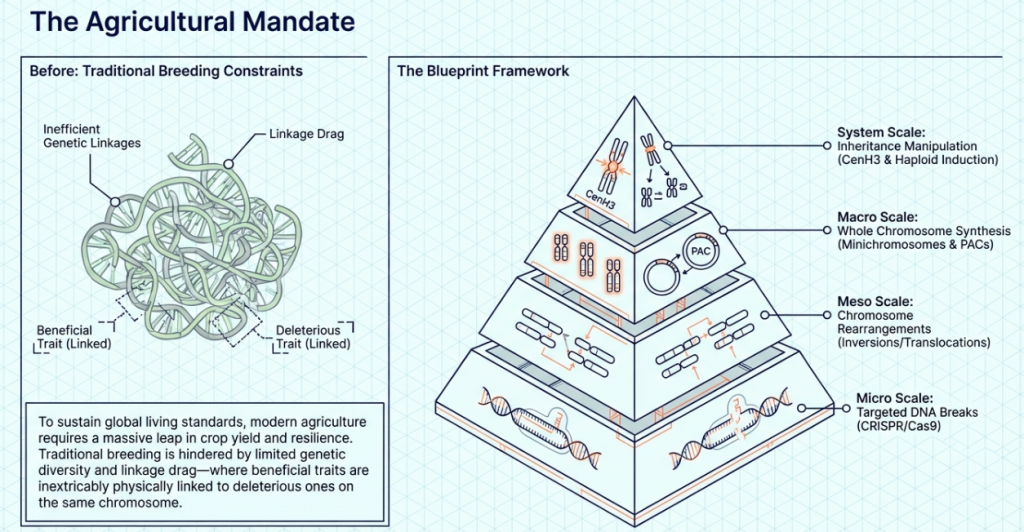
A functional minichromosome requires the following essential components:
- Telomeres: Arrays (such as the Arabidopsis-type repeats) that ensure terminal stability and prevent chromosome deterioration.
- Replication Origins: DNA sequences required for cell-cycle-regulated copying of the vector.
- Centromeres: The structural anchors necessary for faithful mitotic and meiotic segregation.
The Biological Blueprint: Mandatory Components
| Component | Job Description | Stability Role (Mitosis vs. Meiosis) |
|---|---|---|
| Telomeres | Protective DNA-protein caps at the molecule’s termini. | Prevents end-to-end fusion and deterioration; ensures mitotic structural maintenance. |
| Replication Origins | Specific sequences where DNA polymerase initiates duplication. | Ensures the “cargo” is copied accurately before every mitotic and meiotic cycle. |
| Centromere | The epigenetic “anchor” for kinetochore assembly. | Essential for spindle fiber attachment and accurate pulling/segregation during both mitosis and meiosis. |
These vectors provide the boundaries for genetic cargo, but their utility depends entirely on the precision of their functional anchors.
We must evaluate two primary construction methodologies based on species-specific R&D risks:
- Top-Down (Telomere Seeding): Utilizing the Non-Homologous End-Joining (NHEJ) pathway, we truncate endogenous chromosomes by integrating telomeric repeats. This has been successfully demonstrated in maize (Birchler laboratory), wheat, rice, and Arabidopsis. For a truly robust platform, we must utilize the Cre/LoxP system for reporter gene transfer between constructed vectors, as seen in maize.
- Bottom-Up (De Novo Centromeres): This involves assembling functional kinetochores via CENH3-fusion protein tethering (e.g., LexA-CENH3/LexO arrays). While complex, this allows for the creation of synthetic centromeres that drive autonomous segregation across generations.

Strategic Stability Benchmarks:
- Morphological Risk: We must account for species-specific stability. For instance, Ring minichromosomes (like AtARC1) are more stable in Arabidopsis than linear versions, yet they are notoriously unstable in maize. This represents a critical R&D pivot point for different crop programs.
- Telomere Maintenance: Minimum telomere lengths of 1.0–2.3 kb must increase to a 3.0–6.0 kb range within three generations to ensure chromosomal protection.
- Pollen Selection Lever: To overcome transmission issues where engineered chromosomes lag behind endogenous ones, our strategy should include placing pollen selection genes on the minichromosome. This ensures only grains carrying the vector are viable, forcing parity in transmission.
The success of these vectors is inherently linked to our ability to manipulate the underlying mechanisms of centromeric inheritance.
3. The “Top-Down” Strategy: Truncating Native Chromosomes
Minichromosomes serve as independent, high-capacity platforms for trait stacking, allowing us to segregate complex genetic packages independently of the host’s endogenous chromosomes. By functioning as autonomous vectors—complete with telomeres, replication origins, and functional centromeres—these platforms accelerate breeding by bypassing the traditional limitations of genomic balance. This independence is a strategic necessity for managing multi-genic traits without the risk of disrupting the integrity of the primary germplasm.

Strategic “top-down” engineering focuses on the systematic downsizing of endogenous chromosomes. This process utilizes “telomere seeding,” where the integration of cloned telomeric repeats induces truncation at specific sites, removing distal arms while preserving a functional centromere. For subsequent cargo manipulation, we employ site-specific recombination systems like Cre/LoxP. Notably, the complexity of these platforms is increasing; the integration of Ac/Ds transposable elements with Cre/LoxP has enabled the creation of ring minichromosomes. However, a CSO must recognize species-specific stability: while Arabidopsis (AtARC1) ring chromosomes exhibit surprising stability, similar structures in maize are prone to breakage and instability, complicating their use in monocot pipelines.
Supernumerary B chromosomes remain the gold standard for truncation targets. Because they are non-essential and frequently possess “chromosome drive” (exhibiting transmission rates >0.5), they offer a unique advantage for maintaining specialized traits within a population. While drive-loss can be utilized to stabilize transmission—as seen in drive-negative maize minichromosomes where the distal drive-regulating gene is lost during truncation—preserving the drive mechanism remains a powerful tool for rapid trait dissemination.
Critical Technical Hurdles in Minichromosome Stability
- Meiotic Transmission Rates: Engineered platforms often lag behind endogenous chromosomes in transmission efficiency, requiring further optimization of centromere strength.
- Sister Chromatid Cohesion: Mechanical stability is size-dependent; minichromosomes must be large enough to maintain cohesion during the tension of meiosis.
- Size vs. Transmissibility Balance: Strategic sizing is essential—the platform must be small enough to remain phenotypically neutral but large enough to be recognized by meiotic pairing machinery.
- Bivalent Stability and Pairing: Achieving stable bivalent formation remains a challenge; in many species, small engineered chromosomes lack pairing partners, leading to precocious separation and aneuploidy.
The long-term viability of these vectors depends on our ability to engineer functional, autonomous centromeres.
The “Top-Down” methodology involves the strategic downsizing of existing chromosomes. The goal is to retain a functional centromere while removing distal chromosome arms that harbor unnecessary or potentially disruptive genetic information.
The Sequence of Telomere Seeding
Using the Non-Homologous End-Joining (NHEJ) repair pathway, scientists initiate telomere seeding to truncate chromosomes in vivo:
- Construct Introduction: Cloned telomeric repeats (often Arabidopsis-type) are delivered into the cell.
- Repeat Integration: These repeats integrate into distal regions of a target chromosome.
- De Novo End Formation: The integration triggers the formation of a new telomere at the site of the break.
- Acentric Loss: The remaining arm fragment, now lacking a centromere (acentric), is naturally lost during subsequent divisions.
Selection of the Structural Foundation: A vs. B Chromosomes
| Feature | Standard A Chromosomes | Supernumerary B Chromosomes |
|---|---|---|
| Survival Role | Essential for development. | Non-essential; “extra” chromosomes. |
| Engineering Risk | Truncation may result in lethal gene loss. | Modified or lost with zero phenotypic cost. |
| Inheritance | Mendelian patterns. | Exhibit chromosome drive (transmission > 0.5). |
| The “Drive” Paradox | N/A | Preferred Candidates. In maize, truncation often makes them drive-negative by losing distal genes required for drive, which actually stabilizes transmission. |
Biological Shielding and Morphology The stability of these truncated structures is governed by “telomere array length.” In A. thaliana, lengths start at 1.0 to 2.3 kb in the first generation but increase to 3–6 kb by the third generation, providing a natural protection mechanism. Furthermore, architecture matters: while maize ring chromosomes are unstable, Arabidopsis ring minichromosomes (AtARC1) have proven surprisingly more stable than their linear counterparts.
While physical shortening provides the frame, the true challenge of the architect is mastering the epigenetic machinery that grants these vectors their independence.
4. Mastering the Centromere: The Key to Independence
The centromere is the most complex structural element to engineer because it is not defined solely by DNA sequence; it requires epigenetic marking. The histone H3 variant CENH3 serves as the biological signal for kinetochore assembly.
To build a functional centromere from scratch, architects utilize CENH3 tethering:
- Targeting the Site: Integrate a synthetic LexO repeat array into the desired chromosomal location.
- Deploying the Bridge: Introduce a fusion protein—such as LexA-CENH3 or Gal4-CENH3—where the DNA-binding domain (LexA) attaches to the LexO target.
- Recruiting Native Machinery: The tethered CENH3 (often using a GFP-tail swap construct) recruits native CENH3 and other kinetochore proteins, marking the site as a functional centromere.
Epigenetic Landscapes and Risks This process exploits the epigenetic nature of centromere licensing. By “tethering” the protein, we force the cell to recognize a synthetic site. However, if the engineering creates a “dicentric chromosome” (two functional centromeres), it will form anaphase bridges. These occur when the chromosome is pulled toward opposite poles simultaneously, leading to catastrophic breakage and genomic instability.
Once a stable, functional centromere is established, the minichromosome is ready to fulfill its primary purpose as a cargo vector.
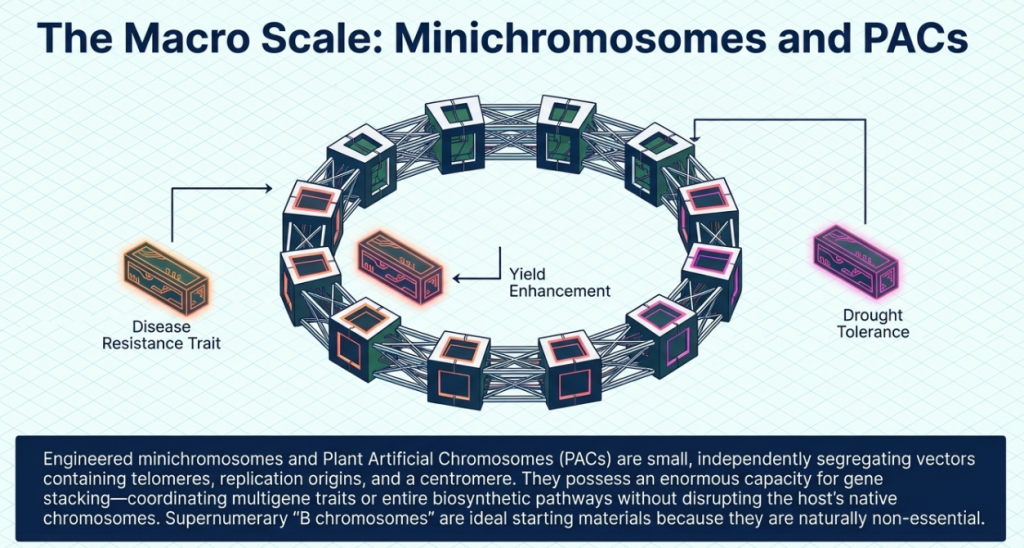
5. The Vanishing Act: A Conceptual Overview of Haploid Induction through Centromere Engineering
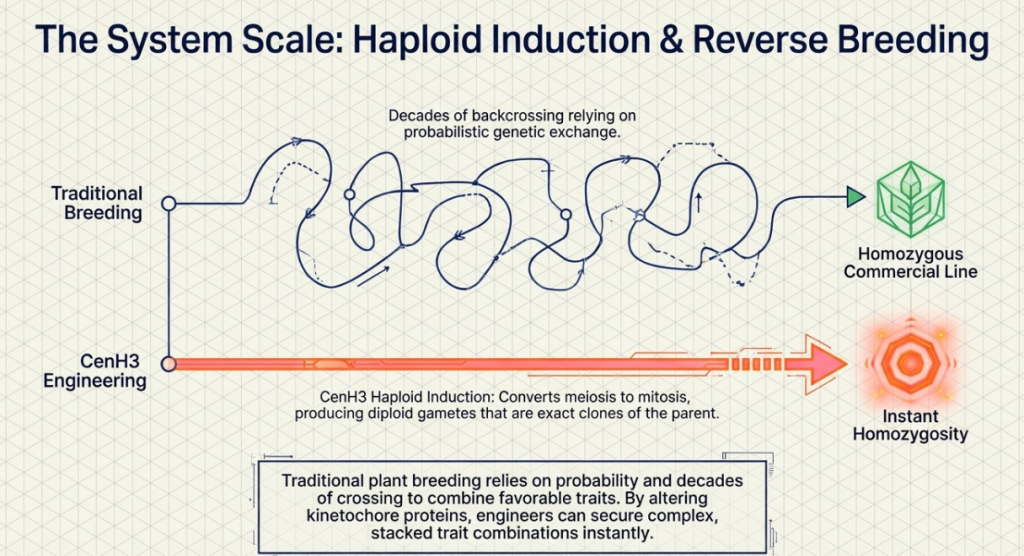
In the pursuit of agricultural productivity, the bottleneck often lies in the slow pace of traditional breeding. For the specialist, the pressure to develop high-yielding, resilient varieties necessitates tools that bypass the multi-generational slog of selfing and selection. Centromere engineering represents a paradigm shift in this field, allowing for the rapid generation of (di)haploids through a process known as uniparental genome elimination. By manipulating the fundamental machinery of chromosome segregation, we can essentially program one parent’s genome to “vanish” post-hybridization, leaving behind a streamlined genetic template.
5.1 The “Why” of Haploids in Modern Breeding
Haploid technology is not merely a laboratory curiosity; it is a critical accelerator for modern crop improvement. For an aspiring breeder, the ability to induce haploidy—and subsequently double the genome to create “doubled haploids” (DH)—provides three transformative benefits:
- Rapid Generation of Inbred Lines: Traditional breeding requires six to eight generations of selfing to achieve near-total homozygosity. DH technology collapses this timeline into just one or two generations, drastically reducing the “time-to-market” for new hybrids.
- Absolute Genetic Stability: Doubled haploids are 100% homozygous at every locus. This fixates beneficial alleles and eliminates the “genetic noise” caused by heterozygous masking of deleterious recessive traits, ensuring high phenotypic uniformity.
- Enhanced Selection Efficiency: In haploids, every allele is expressed. This simplifies the screening for both natural and induced mutations, particularly for recessive traits that would otherwise remain hidden in a diploid population.
While the “so what?” of haploids centers on breeding speed, the “how” requires taking control of the biological steering wheel of inheritance: the centromere.
5.2 The Engine of Segregation: Understanding CENH3 and KNL2
The centromere is the specialized chromosomal domain that serves as the foundation for the kinetochore—the protein complex that attaches to spindle fibers during division. In most plants, centromere identity is maintained through an epigenetic signaling process rather than a specific DNA sequence.
At the heart of this process is CENH3, a centromere-specific histone H3 variant. CENH3 acts as an epigenetic bookmark: its presence on a chromosome prior to replication signals the cell to load “fresh” CENH3 onto the same site after replication. This process is facilitated by centromere licensing factors such as KNL2. If these proteins are modified or depleted, the centromere loses its identity, and the chromosome cannot be successfully maneuvered during mitosis or meiosis.
| Feature | Wild-Type Centromere | Modified/CENH3-Depleted Centromere |
|---|---|---|
| Epigenetic Status | Abundant CENH3/KNL2 signals; clear “bookmarking.” | Reduced, mutated, or missing epigenetic signals. |
| Kinetochore Recruitment | High efficiency; robust protein bridge assembly. | Failure to reassemble functional kinetochores. |
| Fate During Division | Successful segregation to daughter cells. | Eliminated during zygotic or early embryo development. |
By engineering “weakened” centromeres, we can exploit the competitive environment of a hybrid zygote to trigger selective chromosome loss.
5.3 Engineering the “Vanishing Act”: Key Techniques for Genome Elimination
To create haploid inducer lines, researchers target the CENH3 protein and its loading machinery. Four distinct genetic approaches have emerged, each leveraging a unique “primary mechanism of failure” that leads to chromosome loss during hybridization:
- Tail Swaps (GFP-Tail Swap): This involves fusing the N-terminus “tail” of a conventional histone H3 to the histone-fold domain of CENH3.
- Primary Failure: This creates a competitive failure in the zygote. Wild-type CENH3 from the standard parent is preferentially loaded, while the modified inducer centromeres fail to reassemble fresh chromatin, leading to their exclusion.
- Point Mutations and Deletions: Targeted alterations, such as the cenh3-4 mutant or specific splicing disruptions, are utilized.
- Primary Failure: These mutations cause inefficient mRNA splicing or protein malfunctions, resulting in a catastrophic drop in centromeric CENH3 levels.
- Cross-Species Complementation: Native CENH3 is replaced with variants from distant relatives, such as using Brassica variants in Arabidopsis.
- Primary Failure: While functional enough for the inducer’s own survival, the foreign protein is too “divergent” to compete effectively for loading against the wild-type centromere of the crossing partner.
- Targeted Protein Degradation (deGradFP): This system utilizes the ubiquitin-proteasome pathway, employing a GFP-specific nanobody to physically destroy EYFP-tagged CENH3.
- Primary Failure: By targeting the protein specifically in gametes, the centromere is physically absent during the first cell division. This is particularly advantageous for polyploid crops with multiple CENH3 variants.
Recent insights also identify KNL2 as a secondary target. Disruption of this centromere licensing factor results in a 1% haploid induction rate in Arabidopsis, which serves as a foundation for further enhancement. While these genetic designs are the baseline, environmental stressors act as a critical multiplier for induction success.
5.4 The Environmental Multiplier: Heat Stress and Efficiency
Environmental factors, particularly temperature, play a decisive role in the frequency of uniparental genome elimination. Research indicates that heat stress can significantly amplify induction rates:
- Arabidopsis Boost: In cenh3-4 mutants, the induction rate rises from a baseline of 0.2% to 4.1% following short-term heat stress.
- KNL2 Multiplication: The knl2 mutant shows a 10-fold increase in haploid induction effectiveness under temperature stress.
- Barley (H. vulgare x H. bulbosum): Chromosome elimination frequency is significantly higher at temperatures above 20°C compared to those below 17.5°C.
However, specialists must weigh these gains against significant physiological trade-offs. High heat stress is associated with failed endosperm cellularization and a marked reduction in seed germination, necessitating a balanced approach in field applications.
5.5 Real-World Applications: Progress in Major Crops
Translating centromere engineering from the Arabidopsis model to major crops has revealed significant species-specific challenges. The current “state of the art” involves integrating modified constructs into lines with pre-existing genetic weaknesses.
“In Maize, by substituting a tail-altered CENH3 for the full-length protein within a Stock 6 background, researchers achieved induction rates of 16.3%. In Wheat, similar centromere-based strategies have reached 8%, while Switchgrass currently achieves rates between 0.5% and 1.4%.”
A key insight in Maize is the Stock 6 hypothesis: this line is thought to possess inherent centromere flaws. Overexpressing modified CENH3 within Stock 6-derived lines acts as a multiplier, though these results still lag behind the 25–45% efficiencies observed in specialized laboratory models.
5.6 Beyond Simple Haploids: Minichromosomes and Future Horizons
The process of genome elimination occasionally results in “semi-haploids” that carry 3–10 Mb fragments of the inducer genome. These minichromosomes (linear or circular) represent a new frontier in biotechnology.
In Arabidopsis, circular (ring) minichromosomes were found to be more stable than their linear counterparts, a critical finding for long-term transmission. These small, autonomous units serve as ideal biotechnology vectors for “gene stacking.” By placing multiple traits—such as pest resistance and drought tolerance—on a single minichromosome, breeders can insert complex gene packages without disrupting the host plant’s existing genetic architecture.
6. Trait Stacking: The Cargo Chromosome Concept
Minichromosomes serve as “Cargo Ships” for agriculture, enabling Trait Stacking—the simultaneous delivery of multiple complex traits (e.g., drought resistance and nutrient enhancement).
- The Technology Stack: Loading these ships involves site-specific recombinases like Cre/LoxP or Ac/Ds.
- Next-Gen Engineering: Modern architects now use phage-derived large serine recombinases and prime editing to insert specific recognition sites, allowing for the stacking of multi-kilobase DNA fragments.

🌟 Why it Matters: Predictable Architecture
Site-specific recombinases allow for the predictable genomic insertion of massive DNA segments. This ensures that stacked traits remain linked and functional without the random disruption typical of traditional gene insertion methods.
The Size Paradox: Balancing Stability A minichromosome must navigate a delicate size balance:
- The Lower Limit: If a chromosome is too small, it faces the Size Paradox. Specifically, it may suffer from the precocious separation of sister chromatids at meiosis I or a lack of bivalent stability, leading to meiotic loss.
- The Upper Limit: It must remain small enough to be “phenotypically neutral,” ensuring it does not overtax the cell’s metabolic resources.
Mastering this spatial balance is the gateway to a new era of crop domestication through refined chromosomal architecture.
7. Redirecting Meiotic Recombination: Overcoming Inversion-Induced Suppression
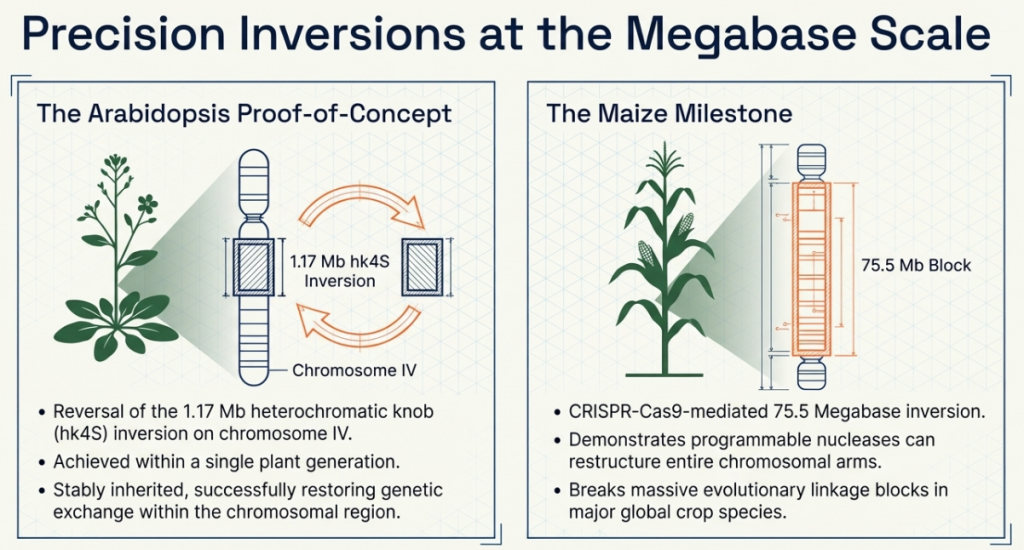
Natural chromosomal inversions, such as the 1.1Mb knob in Arabidopsis, act as “biological safety locks” that suppress genetic exchange. These inversions prevent the recombination of desirable alleles between cultivars, effectively locking elite haplotypes and stalling progress.
Using CRISPR/Cas-mediated site-specific Double-Strand Breaks (DSBs), we can revert these Mb-sized barriers. This has been validated at scale in maize, where a 75.5Mb inversion—spanning a third of a chromosome—was successfully reverted to unlock recombination-cold regions.
Conversely, we can utilize the “Knock Up” approach for regulatory-friendly trait modification. By inducing a 0.9Mb inversion, we can swap endogenous promoters to modulate gene expression without the introduction of foreign DNA, significantly streamlining the path through global GMO regulatory frameworks. Furthermore, purposefully induced inversions are a powerful tool for Trait Bundling; by suppressing local recombination, we can protect Intellectual Property and ensure that elite trait packages remain linked and do not segregate in future crosses. CRISPR-mediated rearrangements allow us to unlock regions of the genome that have been “recombination-cold” for millennia.
- Inversions: Using SaCas9 to revert natural inversions can restore genetic exchange. A landmark example is the 1.1Mb knob (hkS4) in Arabidopsis, which was reverted to “unlock” a region that had been suppressed for five millennia (5,000 years).
- Translocations: NHEJ-based reciprocal translocations allow for the exchange of chromosome arms, effectively unlinking traits from different genetic backgrounds.
- Promoter Swaps: The “knock up” approach uses a targeted inversion to move a distal, strong promoter into proximity with a gene of interest, increasing expression levels significantly without introducing transgenic DNA.
7.1 Directing Meiotic Recombination via Targeted Inversions
Natural chromosomal inversions act as “recombination cold-spots,” locking favorable and unfavorable traits together in repulsion. Breaking these linkage groups is essential for unlocking genetic potential in elite germplasm.
Commercial Scale and the “Knock-Up” Strategy: The feasibility of megabase-scale restructuring is proven; Corteva’s successful reversion of a 75.5Mb inversion in maize (spanning one-third of chromosome 2) demonstrates that we can “unlock” vast genomic regions for genetic exchange.
Strategically, the “Knock-Up” approach offers a significant commercial and regulatory advantage. By inducing a 0.9Mb inversion, we can reposition a strong native promoter to drive a gene of interest.
The “So What?” Layer: Because this modulates expression using native promoters and no foreign DNA, these crops may be exempt from “GMO” classification under emerging PBO/NBT frameworks in multiple jurisdictions. This provides a clear, high-speed regulatory path for increasing trait expression.
7.2 Breaking Genetic Linkages through Reciprocal Translocations
The challenge of “linkage drag”—where deleterious traits are physically tethered to agronomic benefits—is most acute in heterochromatic regions where natural crossovers are rare.
NHEJ Pathway Optimization: To facilitate reciprocal translocations (exchanging chromosome arms), we must optimize the repair of CRISPR-induced double-strand breaks (DSBs). Our strategy involves the temporary inhibition of the classical NHEJ (cNHEJ) pathway. By blocking cNHEJ, which normally tethers “correct” ends together, we shift the repair mechanism toward aNHEJ (alternative NHEJ/micro-homology mediated end joining). This favors the “mis-joining” of ends from different chromosomes, drastically increasing translocation frequencies.
Commercial and Biosafety Impact:
Genetic Isolation: By altering chromosome numbers or structures, we can create Genetic Isolation. This prevents fertile outcrossing with wild relatives, serving as a powerful tool for IP protection and environmental biosafety.
Trait Separation: Breaking linkages in recombination-cold regions to decouple yield from susceptibility.
7.3 Breaking Genetic Linkages via CRISPR-Mediated Chromosomal Translocations
Genetic linkages represent the primary physical obstacle to decoupling beneficial traits from adverse “drag.” In heterochromatic regions where natural crossovers are nearly non-existent, the induction of chromosomal translocations in somatic cells provides a robust alternative for restructuring.

The strategic challenge lies in the repair pathway. Classical Non-Homologous End Joining (cNHEJ) acts as a “proximity guard,” keeping broken ends together to prevent the very mis-joining required for translocations. To increase translocation efficiency, we must temporarily block the cNHEJ pathway. This allows the cellular machinery to bypass the guard mechanism, facilitating the reciprocal exchange of chromosomal arms needed to break persistent linkages.
Comparative Analysis of Linkage Management Techniques
| Methodology | Mechanism of Action | Impact on Linkage Group | Primary Breeding Outcome |
|---|---|---|---|
| Inversion Reversal | CRISPR DSBs at inversion borders | Restores collinearity | Unlocks “cold” regions for recombination |
| Reciprocal Translocation | Induced DSBs on heterologous chromosomes; cNHEJ block | Reorganizes chromosome arms | Decouples linkage drag; breaks “unbreakable” linkages |
| Induced Inversion | Purposeful 180° segment rotation | Suppresses local recombination | Trait Bundling; Protecting IP/Elite Haplotypes |
| “Knock Up” Inversion | CRISPR-mediated promoter swapping | Repositions regulatory elements | Regulatory-friendly expression modulation |
These surgical interventions represent the final step toward the design of entirely synthetic karyotypes.
8. Advanced Applications and The Road Ahead
Chromosome engineering is evolving beyond simple vectors into the discipline of Spatial Architecture, where we reorganize the 3D organization and epigenetic context of the nucleus.
- Breaking Genetic Linkages: Using CRISPR/Cas-mediated reciprocal translocations to separate beneficial traits from adverse ones. This redefines linkage groups, uncoupling genes that have been stuck together for millennia.
- Haploid Induction: By manipulating CENH3 variants (such as the GFP-tail swap) or targeting the centromere licensing factor KNL2, architects can trigger uniparental genome elimination. This produces plants with a single set of chromosomes, accelerating breeding cycles by years.
- Genetic Isolation: We can reorganize a crop’s genome to prevent outcrossing with wild relatives, creating a “built-in” genetic barrier that ensures engineered traits remain contained.
Conceptual Synthesis We are moving from “gene editing” to a world of Spatial Architecture. By rearranging the 3D structure of the nucleus, we are not just editing individual instructions; we are redesigning the entire biological “building.” This approach allows us to place genes in entirely new epigenetic contexts, influencing expression and recombination in ways that were previously impossible.
9. The Future Landscape: Integration, Isolation, and 3D Nuclear Architecture
The convergence of CRISPR/Cas, prime editing, and novel recombinases is transitioning biotechnology from gene-level modifications to whole-chromosome engineering. We are now moving toward the predictable construction of synthetic chromosomes and the management of entire karyotypes.
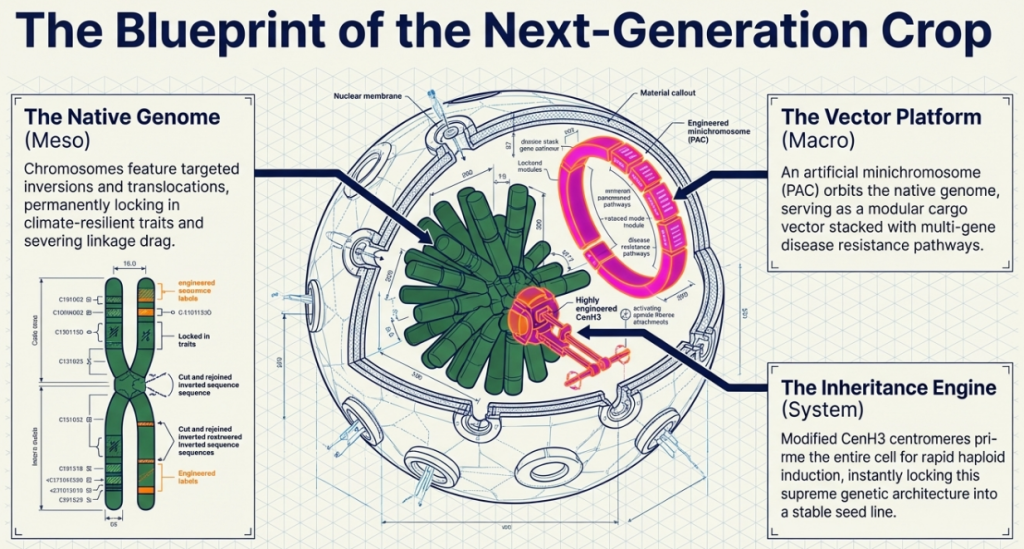
The long-term strategic future is defined by:
- Genetic Isolation and Biological Safety Locks: By inducing consecutive chromosomal rearrangements, we can create varieties that are genotypically identical to wild relatives but chromosomally incompatible. This effectively prevents outcrossing and the uncontrolled spread of specialized traits.
- KaryoCreate and Engineered Aneuploidy: Leveraging KaryoCreate (karyotype CRISPR-engineered aneuploidy technology) allows for the production of chromosome-specific aneuploidies. This provides an unprecedented tool for evaluating the dosage effects of entire chromosomes on complex traits.
- 3D Nuclear Architecture and Positional Dynamics: We are moving toward a functional understanding of the 3D nucleus. By repositioning megabase-sized segments, we can determine whether a gene’s expression is driven primarily by its physical position in the nuclear space or its epigenetic state, opening new avenues for designer gene regulation.
These technologies represent the next frontier in germplasm optimization. By mastering the art of chromosomal restructuring, we are no longer just editing genes—we are rewriting the structural foundations of agriculture to ensure global food security.
Key Terminology Summary
| Term | Definition (Source Context Application) |
|---|---|
| NHEJ | Non-Homologous End-Joining: The primary repair pathway used in plants for transgene insertion and the creation of new telomeres during seeding. |
| Telomere Seeding | The intentional integration of telomeric repeats to truncate a native chromosome and stabilize the newly created end. |
| Epigenetic Marking (CENH3) | The process of using CENH3 proteins to license a specific DNA site as a centromere, independent of the underlying sequence. |
| Linkage Groups | Sets of genes inherited together; these are now being physically redefined through CRISPR/Cas-mediated translocations. |
Image Summary

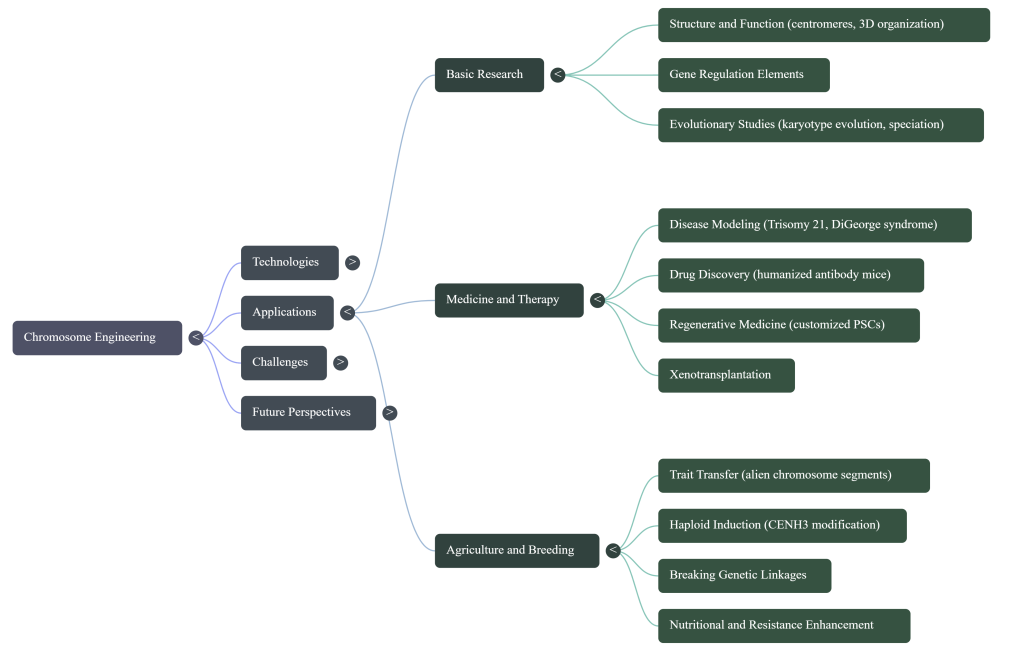


Questions/Answers
1. How does CRISPR/Cas enable the creation of plant minichromosomes?
CRISPR/Cas technology enables the creation of plant minichromosomes (also known as engineered or artificial chromosomes) primarily by providing a precise tool for targeted chromosomal downsizing and karyotype restructuring. Unlike earlier random methods, CRISPR/Cas allows researchers to induce double-strand breaks (DSBs) at specific genomic locations to manipulate chromosome structure.
The specific ways CRISPR/Cas facilitates this process include:
- Targeted Telomere-Mediated Truncation (TMCT): Traditional telomere seeding occurs randomly when telomeric repeats integrate into the genome, causing the distal portion of a chromosome arm to be lost. CRISPR/Cas enhances this by inducing a DSB at a preselected location, allowing telomere repeat DNA to be targeted precisely to that site. This results in a stable, truncated chromosome of a predefined size and content.
- Inducing Large-Scale Deletions: By creating two DSBs on the same chromosome, CRISPR/Cas can be used to remove massive intervening segments of DNA. This “top-down” approach aims to retain essential components like the centromere and telomeres while eliminating non-essential gene-containing arms to create a phenotypically neutral minichromosome.
- Centromere Engineering and Fission: CRISPR/Cas can be combined with “centromere seeding” strategies. For example, a de novo centromere can be formed by tethering CENH3 proteins to designed repeat arrays. Subsequent CRISPR-induced breaks or spindle-induced tension during cell division (chromosome fission) can release these synthetic centromeres as independent, self-sustaining neochromosomes or minichromosomes.
- Chromosome Arm Transfer via Translocations: Researchers have demonstrated that CRISPR/Cas-mediated translocations can be used to fuse independent chromosomes or transfer entire arms. This process can result in the reduction of a karyotype while generating a separate, stable minichromosome that can serve as a vector for trait stacking.
- By-products of Genome Elimination: When CRISPR/Cas is used to edit centromeric proteins (like CENH3) to create “haploid inducer” lines, the resulting uniparental chromosome elimination sometimes produces small, linear, or circular minichromosomes derived from the inducer’s (peri)centromeric regions.
- Retrofitting and Trait Stacking: Once a minichromosome is established, CRISPR/Cas (often in combination with site-specific recombinases) is used as a “power tool” to precisely stack multiple genes or edit specific loci on the minichromosome. This allows the minichromosome to act as a high-capacity “cargo” vector for complex agricultural traits without interfering with the rest of the host genome.
2. Explain how engineering centromeres can produce haploid inducer plants.
Engineering centromeres to produce haploid inducer plants is a revolutionary method in plant breeding that facilitates the rapid creation of 100% homozygous “doubled haploid” lines in a single generation. This process primarily targets the centromeric histone protein 3 (CENH3), an essential variant of histone H3 that defines the identity and function of the centromere by mediating chromosome segregation during cell division.
The standard mechanism for generating these inducer lines involves the following steps:
1. Weakening the Centromere
To create a haploid inducer, scientists subtly disable the centromeres of a plant while ensuring the plant remains viable and fertile. This is typically achieved through two main approaches:
- The “Tailswap” Method: This involves using a cenh3 null mutant (where the native gene is inactivated) and complementing it with a modified transgene. A common modification is the “GFP-tailswap,” where the hyper-variable N-terminal tail of CENH3 is replaced with the tail of conventional histone H3 and fused with a green fluorescent protein (GFP).
- Direct Modification: More recent “one-step” methods use CRISPR/Cas9 or chemical mutagenesis (EMS) to induce specific point mutations or small deletions in the conserved C-terminal histone fold domain (HFD) of the endogenous CenH3 gene. These mutations impair the protein’s ability to bind to the centromere or load properly, effectively “weakening” its competitive ability during cell division.
2. Crossing and Uniparental Chromosome Elimination
Once the haploid inducer plant is established, it is crossed with a wild-type plant. The following biological events occur in the resulting hybrid zygote:
- Kinetochore Competition: During the first mitosis of the hybrid zygote, the chromosomes from the wild-type parent (which have functional CENH3) compete with the chromosomes from the inducer parent (which have modified/weakened centromeres) for attachment to spindle fibers.
- Missegregation: The “weak” inducer centromeres fail to load kinetochore proteins efficiently or fail to bind properly to microtubules. Consequently, the inducer parent’s chromosomes lag behind during anaphase, form micronuclei, and are eventually torn into fragments and degraded.
- Genome Elimination: This process, known as uniparental genome elimination, leaves an embryo containing only the set of chromosomes from the wild-type parent, resulting in a haploid plant.
3. Restoration of Fertility
Because haploid plants have only one set of chromosomes, they are unable to undergo normal meiosis and are sterile. Fertility is restored through spontaneous or chemically induced (e.g., using colchicine) chromosome doubling. This creates a doubled haploid (DH) plant that is completely homozygous at every locus, removing the need for 8–10 generations of traditional backcrossing to achieve true-breeding lines.
Key Advantages and Challenges
- Non-GMO Progeny: A significant benefit is that the resulting haploid plants are technically non-transgenic, as the chromosomes carrying the engineered CENH3 transgenes are entirely eliminated during the process.
- Versatility: This method allows for the production of both maternal and paternal haploids, facilitating rapid cytoplasm exchange and the development of cytoplasmic male sterility (CMS) lines.
- Challenges: Efficiencies can be low in many crops (e.g., 0.5% to 8% in maize and wheat), and incomplete elimination can sometimes lead to aneuploid progeny (plants with an irregular number of chromosomes). Additionally, many crops like sunflower remain difficult to engineer due to their resistance to standard tissue culture and transformation protocols.
3. What are the future benefits of stacking genes on minichromosomes?
Stacking genes on engineered minichromosomes (also referred to as Plant Artificial Chromosomes or PACs) offers a transformative “super vector” platform for plant biotechnology that overcomes many limitations of traditional genetic engineering.
The future benefits of gene stacking on these platforms include:
1. Management of Highly Complex Traits
Traditional transgenesis is often limited to a few genes, but minichromosomes allow for the coordinated transfer and expression of an almost unlimited number of genes. This enables:
- Simple Stacking: Combining multiple genes for herbicide resistance, insect prevention, and disease resistance in a single package.
- Compound Gene Complexes: Engineering sophisticated genetic networks for complex agricultural traits such as increased water and fertilizer utilization, metabolic engineering for nutritional or medicinal purposes, and high biomass production for biofuels.
- Orthogonal Integration: In polyploid crops, where genome redundancy makes traditional stacking difficult, PACs provide dedicated integration sites with predictable inheritance and defined copy numbers.
2. Elimination of Linkage Drag
A major hurdle in conventional breeding is linkage drag, where a desired gene is physically linked to deleterious genes that cannot be easily separated by backcrossing.
- Minichromosomes act as independent linkage groups that do not recombine with the host plant’s native chromosomes.
- This allows a whole package of beneficial genes to be transferred to a new cultivar in a single cross without introducing unwanted genetic material from the donor’s genome.
3. Preservation of Genomic Integrity and Stability
Minichromosomes offer a safer way to introduce new traits without disrupting the host’s existing genetic blueprint:
- Endogenous Gene Protection: Unlike traditional methods that integrate transgenes randomly, PACs prevent the disruption of native genes and the associated unpredictable “position effects” on gene expression.
- Phenotypic Neutrality: Because they are designed to contain only essential components (centromeres, telomeres, and replication origins) and lack essential host genes, they have minimal interference with the plant’s normal growth and development.
- Faithful Transmission: Minichromosomes produced through chromosomal truncation are stable during both mitosis and meiosis, ensuring traits are faithfully inherited across generations.
4. Advanced Precision and Modularity
Minichromosomes are designed to be highly “retrofittable” using modern molecular tools:
- Precise Gene Editing: They facilitate the use of site-specific recombination (SSR) systems (like Cre-lox) and CRISPR/Cas technology to precisely add, delete, or replace genes in vivo at different times.
- Gene Replacement: Researchers can design systems where older transgenes are replaced with improved versions on the same minichromosome vector.
5. Streamlined Breeding and Regulatory Pathways
The unique properties of PACs could simplify the commercialization of new crop varieties:
- Ease of Transfer: Once a package of genes is established on a stable minichromosome, it can be transferred between compatible varieties through sexual hybridization or haploid breeding.
- Deregulation Efficiency: Since the entire array of genes acts as a single unit, it may be possible to perform concurrent environmental and health impact assessments for the whole package, potentially speeding up regulatory approval compared to assessing dozens of individual random insertions.
- Specialized Production: PACs could be used to engineer plants as bio-factories to produce pharmaceutical or metabolic products in contained environments, utilizing the ability to express entire biological pathways.
4. What is the difference between top-down and bottom-up engineering?
In the context of chromosome engineering and the creation of artificial chromosomes, the primary difference between top-down and bottom-up engineering lies in whether the process begins with an existing native chromosome or assembles a new one from individual components.
Top-Down Engineering
The top-down approach involves the modification and downsizing of existing chromosomes already present in the host organism.
- Mechanism: It typically utilizes telomere-mediated chromosomal truncation (TMCT) or large-scale deletions to remove non-essential, gene-containing arms while retaining the essential functional components, such as the original endogenous centromere, telomeres, and origins of replication.
- Goal: The aim is to create phenotypically neutral minichromosomes that do not interfere with the plant’s normal growth but can serve as stable vectors for carrying new genes.
- Status in Plants: This method is currently considered the more robust and successful approach for plants.
Bottom-Up Engineering
The bottom-up approach involves the de novo assembly of new chromosomes from scratch using cloned or synthetic genetic parts.
- Mechanism: Researchers assemble the three essential elements of a chromosome—centromeres, telomeres, and replication origins—in vitro or via cell-mediated processes. One specific method involves centromere seeding, where a synthetic centromere is formed by tethering CENH3 proteins to designed repeat arrays.
- Goal: This strategy aims to build entirely synthetic chromosomes that can be customized with specific features and gene cargos.
- Challenges: While well-established in yeast and some animal cells, this approach faces significant hurdles in plants, largely due to the complexity of plant centromeres and the strong epigenetic requirements for their function.
Summary Table of Differences
| Feature | Top-Down Engineering | Bottom-Up Engineering |
|---|---|---|
| Starting Point | Existing native chromosomes | Cloned or synthetic components |
| Primary Method | Truncation and arm removal | De novo assembly and synthesis |
| Centromere Source | Retains original endogenous centromere | Requires synthetic or seeded centromere |
| Flexibility | Dependent on existing architecture | Offers maximum design flexibility |
| Plant Success | Proven and robust | Experimental and more challenging |
5. How does centromere seeding allow for synthetic chromosome creation?
Centromere seeding allows for the creation of synthetic chromosomes (such as minichromosomes, neochromosomes, or Plant Artificial Chromosomes) by establishing functional centromere activity on non-centromeric DNA sequences. This process bypasses the need to chemically synthesize and transform the massive, highly repetitive DNA arrays found in natural centromeres, which has historically been a major obstacle in synthetic genome biology.
The mechanism involves several key steps and technological components:
- Tethering CENH3 Fusion Proteins: The primary method for seeding involves the use of a DNA-binding system, such as the bacterial LacO-LacI or LexO-LexA systems. A fusion protein is created combining a DNA-binding domain (like LexA or LacI) with CENH3 (the centromere-specific histone H3 variant) or a related assembly factor.
- Recruiting Kinetochore Proteins: When this fusion protein is expressed in a plant carrying the corresponding target DNA array (e.g., LexO or LacO repeats), it binds to those sites and recruits native CENH3 and other proteinaceous kinetochore complexes. This effectively reconstitutes a functional centromere at an ectopic (non-native) location.
- Inducing Chromosome Fission or Truncation: To transform this ectopic site into an independent synthetic chromosome, the DNA fragment must be “released” from the host chromosome:
- Chromosome Fission: In maize, tethering LexA-CENH3 to repeat arrays on a chromosome arm creates a dicentric chromosome (a chromosome with two centromeres). During cell division, spindle-induced tension can cause chromosome fission, breaking the chromosome and resulting in self-sustaining, heritable neochromosomes.
- Targeted Truncation: Alternatively, centromere seeding can be combined with telomere-mediated chromosomal truncation. By inducing double-strand breaks (DSBs) proximal to the seeded centromere using CRISPR/Cas, new telomeres can be established, releasing the fragment as a prototype minichromosome.
- Transmissibility and Heritage: Once seeded and released, these synthetic centromeres can drive autonomous chromosome segregation and are faithfully transmitted across multiple generations, even in the absence of the original activator protein, due to the epigenetic nature of centromere maintenance.
This “bottom-up” or “native-centromere-bypass” approach enables the design of synthetic chromosomes with predefined lengths and content, providing expandable genomic space for high-capacity gene stacking of complex agricultural traits.
6. Could minichromosomes be used to stack many genes at once?
Yes, engineered minichromosomes (also called Plant Artificial Chromosomes or PACs) are specifically designed to serve as high-capacity “super vectors” or “cargo chromosomes” for stacking many genes at once.
While traditional genetic engineering is often limited to transferring a few genes at a time due to random integration and capacity constraints, minichromosomes offer a revolutionary platform for complex trait management.
Why Minichromosomes are Ideal for Gene Stacking
Minichromosomes provide several unique advantages for stacking multiple genes:
- Coordinated Transfer and Inheritance: They enable the coordinated transfer and expression of an almost unlimited number of genes. This allows researchers to install entire biosynthetic pathways or coordinate complex multigene traits, such as increased water and fertilizer utilization or metabolic engineering for biofuels and medicine.
- Elimination of Linkage Drag: Because minichromosomes act as independent linkage groups, they do not recombine with the plant’s native chromosomes. This allows a whole package of beneficial genes to be transferred to a new cultivar in a single cross without bringing along unwanted “drag” from the donor genome.
- Genomic Integrity: Unlike traditional methods that integrate transgenes randomly, PACs prevent the disruption of native genes and avoid unpredictable “position effects” on gene expression.
- Orthogonal Integration in Polyploids: In polyploid crops where genome redundancy makes traditional stacking difficult, minichromosomes provide dedicated integration sites with predictable inheritance and defined copy numbers.
Tools Used to Stack Genes on Minichromosomes
Several precision molecular tools are used to add, delete, or replace genes on these platforms:
- Site-Specific Recombination (SSR) Systems: Tools like Cre-lox, FLP-frt, and phiC31-att allow for the precise insertion and “retrofitting” of minichromosomes with multiple genes.
- Recombinase-Mediated Cassette Exchange (RMCE): This method facilitates the exchange or sequential stacking of gene cassettes at the same locus on the minichromosome in vivo over time.
- CRISPR/Cas and Genome Editing: CRISPR technology is used as a “power tool” for in vivo gene stacking and precise editing of specific loci on established minichromosomes.
- Advanced Insertion Systems: Newer platforms like PrimeRoot (capable of precise insertions up to 11.1 kb) and Programmable Chromosome Engineering (PCE) are being developed specifically for these high-capacity applications in crops like rice and maize.
Once a package of genes is established on a stable minichromosome, it can be efficiently transferred between varieties through sexual hybridization or haploid breeding.
7. How does the tailswap method weaken centromeres?
The tailswap method weakens centromeres by altering the essential centromeric protein CENH3 (a variant of histone H3) so that it functions well enough to keep the parent plant viable but fails to compete effectively during cell division when paired with wild-type chromosomes.
This weakening is achieved through the following molecular modifications:
- Replacing the N-Terminal Tail: CENH3 consists of two domains: a conserved C-terminal histone fold domain (HFD) and a highly variable N-terminal tail. In the tailswap method, the native hyper-variable N-terminal tail of CENH3 is replaced with the tail of a conventional histone, such as histone H3.3.
- Adding a GFP Tag: This modified protein is typically fused with a Green Fluorescent Protein (GFP) tag at the N-terminus (termed “GFP-tailswap”).
- Functional Complementation: This chimeric protein is introduced into a cenh3 null mutant (a plant where the native CENH3 gene is inactive). Because the modified protein retains a functional histone fold domain, it can still localize to centromeres and direct chromosome segregation well enough for the inducer plant to grow, though it often suffers from meiotic defects and reduced fertility.
The Mechanism of Weakness in Hybrid Zygotes
The “weakness” of these engineered centromeres becomes apparent when the inducer plant is crossed with a wild-type plant:
- Preferential Removal and Poor Loading: Studies in Arabidopsis zygotes show that modified CENH3 is selectively removed from the centromeres of the inducer parent. The wild-type centromeres (from the other parent) then receive preferential loading of functional CENH3.
- Kinetochore Malfunction: Because the tailswap protein fails to recruit or stabilize centromere complex proteins effectively, the chromosomes from the inducer parent develop defective kinetochores.
- Missegregation and Elimination: During the first mitosis of the hybrid zygote, these “weak” centromeres fail to bind properly to spindle microtubules. Consequently, the inducer’s chromosomes lag behind during anaphase, form micronuclei, and are eventually torn into fragments and degraded—a process known as uniparental genome elimination.
This process leaves an embryo containing only the chromosomes from the wild-type parent, resulting in a haploid plant that can later be doubled to create a homozygous line.
8. What role does temperature play in haploid induction?
Temperature plays a significant role in determining the efficiency and frequency of centromere-mediated genome elimination and haploid induction in plants. Research indicates that environmental factors, specifically heat, can be used as a tool to drastically increase the yield of haploid progeny.
The influence of temperature on haploid induction is characterized by the following:
- Enhancement of Elimination Frequency: High temperatures generally increase the frequency of uniparental genome elimination. For example, in wide crosses between Hordeum vulgare and Hordeum bulbosum, chromosome elimination was much higher at temperatures above 20°C compared to those below 17.5°C.
- Synergy with Centromeric Mutations: Temperature stress acts synergistically with specific mutations in centromere-related proteins:
- In cenh3-4 mutants, which typically produce a very low frequency of haploids (0.2%), a short heat stress treatment increased the frequency to 4.1%.
- In knl2 mutants (defective in a centromere licensing factor), short-term temperature stress increased haploid induction effectiveness tenfold.
- Haploid inducer lines such as CENH3 GFP-tailswap and CENH3 G83E also showed increased haploid frequencies when combined with heat stress.
- Biphasic Temperature Manipulation: Temperature can be used strategically to overcome biological hurdles like male sterility in inducer lines. Lowering the temperature can dramatically enhance the pollen quality of sterile lines like the GFP-tailswap. A subsequent increase in temperature after crossing these lines with wild-type plants has been shown to induce maternal haploids at a rate of approximately 25%.
- Trade-offs and Risks: While high temperatures can boost haploid rates, they can also have deleterious effects on seed development. Heat-treated crosses often result in endosperm failure and seeds that fail to germinate. Furthermore, the sensitivity of the elimination process to environmental fluctuations can be a disadvantage if stable, predictable seed production is required.
In summary, temperature manipulation is an emerging and highly efficient strategy to induce both paternal and maternal haploids by increasing the frequency of genome elimination during early zygotic development.
9. How are haploid plants restored to fertile diploids?
Haploid plants are sterile because they carry only a single set of chromosomes (n), which prevents them from undergoing normal meiosis. To restore fertility, the chromosome number must be doubled to create doubled haploid (DH) lines (2n).
The restoration process occurs through the following methods:
- Spontaneous Chromosome Doubling: In some instances, fertility is restored naturally without human intervention when chromosomes double on their own during plant development.
- Chemically Induced Doubling: The most common method for restoring fertility involves treating haploid material with chemical agents, primarily colchicine.
- Mechanism of Action: Colchicine acts as a microtubule inhibitor. It disrupts microtubule synthesis and inhibits spindle formation during cell division. By preventing the proper separation of chromosomes into daughter cells, the chemical forces the cell to retain a doubled set of chromosomes, effectively converting the haploid state into a diploid state.
Benefits of Restoring Fertility via Doubled Haploidy
The restoration of fertility through chromosome doubling is a critical shortcut in modern plant breeding:
- Instant Homozygosity: Doubled haploids are 100% homozygous at every genetic locus in a single generation.
- Time Efficiency: This technology eliminates the need for the 8–10 generations of repetitive backcrossing and selection required by conventional breeding to establish true-breeding lines.
- Non-GMO Progeny: When haploids are produced through centromere engineering (such as editing the CENH3 gene), the resulting doubled haploids are technically non-transgenic, as the engineered chromosomes from the “inducer” parent are entirely eliminated during the initial cross.
10. What is linkage drag and how do PACs avoid it?
Linkage drag is a significant hurdle in conventional plant breeding where a desired gene is physically linked to deleterious or undesired genes on the same chromosome. Because these genes are located in close proximity, especially in recombination-poor regions, they tend to be inherited together as a package.
Plant Artificial Chromosomes (PACs), also known as engineered minichromosomes, avoid linkage drag through the following mechanisms:
1. Independent Segregation
PACs act as independent linkage groups or independent chromosomes. Unlike traditional transgenes that integrate randomly into native chromosomes, PACs do not physically link to the host plant’s existing genome. This independence ensures that the beneficial traits carried on the PAC are not “dragged” along with any unfavorable alleles from the host’s native chromosomes.
2. Lack of Recombination with Native Chromosomes
Minichromosomes are specifically designed to avoid pairing or recombining with the plant’s progenitor (native) chromosomes during meiosis.
- Because they do not engage in recombination with the host genome, they remain as autonomous vectors.
- The entire package of genes installed on a PAC acts as a single, stable unit of inheritance.
3. Efficiency in Trait Transfer
In conventional breeding, removing unwanted linked genetic material requires many rounds of tedious and time-consuming backcrossing. In contrast, a whole set of beneficial traits stacked on a PAC can be transferred to a new cultivar in a single cross. This allows breeders to introduce complex traits—such as simultaneous resistance to multiple diseases or improved environmental tolerance—without the risk of introducing “drag” from the donor’s genome.
4. Protection of Genomic Integrity
By functioning as independent platforms, PACs prevent the disruption of endogenous gene functions that often occurs during the random integration of traditional transgenes. They also avoid unpredictable “position effects” where the location of a gene on a native chromosome might negatively impact its expression.
11. Can minichromosomes be used to turn plants into bio-factories?
Yes, engineered minichromosomes (or Plant Artificial Chromosomes, PACs) can be used to turn plants into bio-factories by serving as high-capacity “super vectors” for complex metabolic and industrial engineering.
The technical capabilities of minichromosomes facilitate this application in several specific ways:
- Engineering Entire Biosynthetic Pathways: Unlike traditional genetic engineering, which is often limited to a few genes, minichromosomes can accommodate an almost unlimited number of genes. This allows researchers to install coordinated compound gene complexes and entire biological pathways to produce high-value products.
- Production of Pharmaceuticals and Medicine: PAC technology has significant potential for industrial applications to produce pharmaceutical products and metabolic products with biomedical activities. Plants can be engineered to act as contained, efficient production systems for these complex molecules.
- Metabolic Engineering for Nutrition and Biofuels: Minichromosomes can be used to redirect a plant’s metabolism for nutritional improvement or to adapt plants to produce high amounts of biomass specifically for biofuel feedstock.
- Independent and Stable Expression: Because minichromosomes function as independent linkage groups, the resident genes are faithfully expressed and transmitted without interfering with the plant’s native growth and development. This allows the plant to maintain its standard physiological functions while simultaneously operating as a factory for specific industrial or medical compounds.
- Contained Environments for Industrial Use: For biomedical and pharmaceutical production, these engineered plants could be grown in contained environments to ensure biosafety while fulfilling specialized industrial needs.
By overcoming the “linkage drag” and capacity limits of conventional breeding and transgenesis, PACs offer a modular platform for precisely stacking the traits necessary for diverse bio-factory applications.
12. Tell me more about the differences between ‘top-down’ and ‘bottom-up’ chromosome engineering.
The primary difference between top-down and bottom-up chromosome engineering lies in the starting point: the top-down approach modifies and downsizes existing native chromosomes, while the bottom-up approach assembles new chromosomes from scratch using individual genetic components.
1. Top-Down Engineering
This approach is currently considered the more robust and successful method for plants.
- Mechanism: It involves the large-scale deletion of non-essential, gene-containing chromosome arms while retaining the essential functional elements required for stability.
- Key Tool: The most common method is Telomere-Mediated Chromosomal Truncation (TMCT), where transformed telomere repeats “seed” new telomeres at the site of integration, causing the distal portion of the chromosome arm to be lost.
- Centromere Source: It utilizes the original endogenous centromere of the host chromosome, which ensures high mitotic and meiotic stability because the epigenetic landscape is already established.
- Applications: It is used to create phenotypically neutral minichromosomes (also called Plant Artificial Chromosomes or PACs) that act as independent linkage groups for gene stacking without interfering with the rest of the genome. Researchers often target B chromosomes for truncation because they are supernumerary and non-essential.
2. Bottom-Up Engineering
This strategy aims to build entirely synthetic chromosomes from individual parts.
- Mechanism: Researchers attempt to assemble the three “pillars” of a chromosome—centromeres, telomeres, and replication origins—either in vitro or through cell-mediated processes.
- Centromere Source: Unlike the top-down approach, this requires the de novo assembly or “seeding” of a synthetic centromere. One method involves tethering CENH3 (the centromere-defining protein) to designed repeat arrays, such as the bacterial LacO-LacI system, to initiate functional centromere activity at a new genomic location.
- Challenges in Plants: While bottom-up methods are successful in yeast and animal cells, they face significant hurdles in plants due to the complexity of plant centromeres and strong epigenetic requirements for their function. For example, simply transforming centromeric DNA sequences into plants is often insufficient to create a functional centromere.
- Future Potential: This approach offers maximum design flexibility, allowing for the creation of chromosomes with predefined lengths, specific gene cargos, and “megachunks” of functional open reading frames.
Comparison Summary
| Feature | Top-Down Approach | Bottom-Up Approach |
|---|---|---|
| Starting Point | Existing native chromosomes. | Cloned or synthetic parts. |
| Primary Method | Chromosomal truncation (TMCT) or large deletions. | De novo assembly and centromere seeding. |
| Centromere | Retains endogenous host centromere. | Requires synthetic or “seeded” centromere. |
| Plant Success | Proven, robust, and stable. | Experimental and challenging. |
| Design Flexibility | Limited by the host’s existing architecture. | Offers maximum customizability. |
Recent breakthroughs have even moved toward karyotype restructuring, where consecutive CRISPR/Cas-mediated translocations are used to fuse independent chromosomes (reducing chromosome number) or induce chromosome fission (increasing chromosome number), effectively blurring the lines between standard engineering and total genome reconstruction.
13. What role does CRISPR/Cas play in targeted chromosomal translocation?
CRISPR/Cas plays a critical role in targeted chromosomal translocation by providing the precision necessary to induce simultaneous double-strand breaks (DSBs) on different, non-homologous chromosomes. This allows researchers to move large genomic segments between chromosomes, a feat previously difficult to control in plants.
The mechanisms and applications of CRISPR-mediated translocations include:
1. Mechanism: Harnessing Somatic DNA Repair
Targeted translocations are achieved by inducing DSBs at specific locations on two different chromosomes. In somatic plant cells, these breaks are typically repaired via non-homologous end joining (NHEJ).
- Classical NHEJ (cNHEJ): This pathway is responsible for the majority of precise translocations. It tends to keep the “correct” ends of a DSB together, which actually suppresses the frequency of translocations.
- Alternative NHEJ (aNHEJ): Researchers have found that blocking the cNHEJ pathway (e.g., by knocking out the KU70 gene) significantly increases the frequency of translocations. However, this comes at the cost of precision, often resulting in larger deletions or inversions at the junction sites.
2. Management of Genetic Linkages
A primary application in plant breeding is the manipulation of traits that are physically linked on chromosomes:
- Breaking Linkage Drag: CRISPR can decouple desirable traits from deleterious or unfavorable genes by moving them to different chromosomes.
- Fixing Traits Together: Conversely, translocations can be used to move two favorable genes from different chromosomes into close proximity on a single chromosome, ensuring they are inherited together as a stable unit.
3. Karyotype Restructuring and Chromosome Fusion
Recent breakthroughs have demonstrated that CRISPR/Cas can be used to fundamentally alter a plant’s genome structure:
- Reducing Chromosome Numbers: In a landmark study, researchers used CRISPR/Cas9 to induce sequential translocations in Arabidopsis thaliana, fusing independent chromosomes to reduce the karyotype from 10 to 8 chromosomes.
- Mimicking Evolution: These techniques allow scientists to imitate natural evolutionary processes, such as chromosome fissions and fusions, to study their impact on gene expression and genome stability.
4. Experimental Successes and Limitations
- Heritability: By using egg cell-specific promoters to drive Cas9 expression, scientists have successfully generated translocations that are stably inherited by subsequent generations.
- Efficiency Challenges: Translocations generally occur at a much lower frequency than intrachromosomal inversions or deletions. For instance, in Arabidopsis, reciprocal translocations of 0.5 Mb and 1 Mb fragments were achieved at frequencies of approximately 2.5% to 3.75%.
- Reproductive Isolation: Large-scale chromosomal restructuring can create reproductive barriers, resulting in plants that are phenotypically normal but unable to produce fertile offspring with wild-type relatives, potentially preventing the uncontrolled spread of engineered traits.
14. Can minichromosomes be used to prevent crop outcrossing?
Yes, chromosome engineering—which includes the creation of minichromosomes—can be used to prevent crop outcrossing by establishing genetic isolation between an engineered crop and its wild relatives.
While minichromosomes are primarily designed as high-capacity “cargo vectors” for gene stacking, the broader techniques of chromosome engineering enable the creation of reproductive barriers that prevent the uncontrolled spread of traits through the following mechanisms:
- Large-Scale Karyotype Restructuring: Researchers can use CRISPR/Cas to induce consecutive translocations or chromosome fusions, fundamentally changing the number of chromosomes or the arrangement of linkage groups in a crop.
- Establishment of Reproductive Isolation: These major structural changes can result in plants that are phenotypically normal but unable to produce fertile offspring when they cross with wild-type relatives or original varieties. This “reprogramming” of the genome at the chromosomal level makes the individuals genetically incompatible with their wild counterparts.
- Minichromosomes as Independent Units: Because minichromosomes are engineered to be stable and independent linkage groups, they can carry beneficial traits in a genomic background that has been restructured to be reproductively isolated.
- Pollen Selection Systems: It is also envisaged that pollen selection genes could be placed on engineered chromosomes. In such a system, only the pollen grains containing the specific engineered chromosome (and its associated traits) would be viable, which could be used to control how traits are transmitted and maintained within a specific population.
By combining these tools, scientists can develop crops that are almost identical to their wild relatives at the gene level but are genetically isolated, effectively avoiding the risks associated with trait escape via outcrossing.
15. What are the regulatory benefits of using minichromosomes?
The use of engineered minichromosomes (or Plant Artificial Chromosomes, PACs) offers several unique regulatory and commercial advantages compared to traditional genetic engineering. These benefits stem primarily from their nature as stable, independent units of inheritance that do not integrate randomly into the host genome.
1. Concurrent Regulatory Assessment
A major bottleneck in biotechnology is the time and cost of deregulation, which can range from $1 million to $15 million per individual insertion event.
- Trait Stacking as a Single Unit: While traditional methods require separate assessments for each random transgene insertion, a package of multiple genes stacked on a single minichromosome can be assessed concurrently.
- Efficiency: Because the entire array of genes acts as a single unit, the increased number of genes does not necessarily prolong the regulatory process.
2. Reduced Regulatory Barriers via “Non-Transgenic” Progeny
In many instances, the techniques used to create or transfer minichromosomes can produce plants that are technically non-GMO, which allows them to bypass certain regulatory frameworks.
- Genome Elimination: When minichromosomes are used in conjunction with CENH3-mediated haploid induction, the progeny contain only the desired set of chromosomes from one parent. Because the engineered “inducer” chromosomes are entirely lost during uniparental genome elimination, the resulting haploid plants are technically non-transgenic.
- Public and Global Acceptance: Producing non-transgenic progeny makes these varieties much easier to market in regions with high public resistance to GMOs, such as Europe.
3. Ease of Cross-Lineage Transfer
Once a package of genes on a stable minichromosome has been deregulated, it offers long-term commercial efficiency.
- Reusable Regulatory Packages: A minichromosome can be transferred from one cultivar to another through sexual hybridization or haploid breeding without needing a complete repeated assessment for each new line.
- Stable Linkage: Because minichromosomes do not recombine with the host’s native chromosomes, the package of genes remains a single, stable unit across generations, ensuring the regulatory profile of the “cargo” remains consistent as it is moved into different genetic backgrounds.
4. Avoidance of “Position Effects”
Traditional transgenesis involves random integration into the genome, which can disrupt native genes or lead to unpredictable “position effects” where the surrounding DNA negatively impacts transgene expression.
- Predictability: Minichromosomes avoid these issues by remaining autonomous. This increased predictability in gene expression may simplify the safety and stability data required by regulatory agencies during the risk assessment process.
- Genomic Integrity: By not disrupting endogenous genes, minichromosomes are less likely to cause unintended phenotypic changes that would trigger more intensive regulatory scrutiny.
16. Could you explain the difference between linear and ring minichromosomes?
The primary differences between linear and ring minichromosomes (also called plant artificial chromosomes) relate to their physical structure, stability during inheritance, and the molecular tools used to create them.
1. Structural Differences
- Linear Minichromosomes: These mimic the structure of natural chromosomes, possessing a centromere and two arms that must be capped by telomeres to prevent DNA degradation or fusion.
- Ring Minichromosomes: These are circular DNA molecules. Because they have no ends, they do not require telomeres for stability.
2. Stability and Transmission
The stability of these structures varies significantly depending on the host species:
- In Arabidopsis thaliana: Ring minichromosomes (such as the 2.85 Mb AtARC1) have been found to be more stably transmitted through sexual reproduction than comparable linear minichromosomes.
- In Maize: The behavior is the opposite. Small ring chromosomes in maize are typically unstable, do not follow Mendelian inheritance, and their transmission is often impaired.
- Meiotic Pairing: Both types often struggle to find pairing partners during meiosis because of their small size, which can lead to reduced transmission rates compared to full-sized native chromosomes.
3. Methods of Creation
- Linear minichromosomes are most commonly produced through Telomere-Mediated Chromosomal Truncation (TMCT). In this “top-down” approach, telomere repeats are integrated into a native chromosome (often a non-essential B chromosome), causing the distal portion of the arm to be lost and leaving a stable, truncated linear fragment.
- Ring minichromosomes are typically created using site-specific recombination systems, such as Cre-loxP. Researchers place two loxP sites on a native chromosome; when the Cre recombinase is introduced, it triggers a recombination event that releases the intervening DNA as a circular fragment (a “plant artificial ring chromosome” or PARC).
4. Summary Comparison
| Feature | Linear Minichromosome | Ring Minichromosome |
|---|---|---|
| Shape | Linear string | Circular loop |
| Telomeres | Required to cap ends | Not required |
| Primary Method | Telomere-mediated truncation (TMCT) | Site-specific recombination (Cre-lox) |
| Arabidopsis Stability | Less stable than rings | High mitotic/meiotic transmission |
| Maize Stability | More stable than rings | Highly unstable; impaired transmission |
17. What is the minimum size for a stable minichromosome?
The exact minimum size for a stable minichromosome is currently unknown and is likely species-specific. While there is a well-defined upper limit for chromosome size (based on the length of the spindle axis during mitosis), the lower limit is less clear.
According to the sources, several factors and observations define our current understanding of minichromosome stability:
- Meiotic Requirements: The potential lower size limit may be determined by the biological requirements for proper sister chromatid cohesion or bivalent stability during meiosis. If a chromosome is too small, it may struggle to find a pairing partner or fail to segregate faithfully.
- Host Genome Influence: There is evidence that the minimum size for stable segregation may depend on the size of the host plant’s genome.
- Typical Size Ranges: Most engineered minichromosomes created to date typically range from a few hundred kilobases (kb) to a few megabases (Mb) in size.
- Species Variations in Stability:
- Arabidopsis thaliana: Researchers successfully generated a stable 2.85 Mb artificial ring chromosome (AtARC1) that demonstrated substantial mitotic and meiotic transmission.
- Maize: In contrast to Arabidopsis, small ring chromosomes in maize are generally unstable and their transmission is often impaired.
- Replication Control: Very small chromosomes can sometimes lose replication control that is normally tied to the cell cycle, as has been observed in some maize B minichromosomes.
To date, the majority of synthetic efforts involving artificial chromosomes have resulted in DNA structures of less than 1 Mb when additional sequences were not added to existing chromosomes. To overcome transmission issues in smaller engineered chromosomes, researchers have suggested potentially adding pollen selection genes to ensure only those gametes carrying the minichromosome remain viable.
18. How does temperature stress increase haploid induction frequency?
Temperature stress, particularly high heat, functions as a powerful catalyst for centromere-mediated genome elimination, significantly boosting the frequency of haploid induction across various plant models and crossing strategies.
Research highlights several ways temperature impacts this process:
1. Synergy with Centromere Mutations
High temperatures act synergistically with specific mutations in centromere-related proteins to increase haploid yields:
- cenh3-4 Mutants: In Arabidopsis, these mutants typically produce a very low haploid frequency (0.2%), but a short heat stress treatment increases this to 4.1%.
- knl2 Mutants: The inactivation of the centromere licensing factor KNL2 generally produces a 1% haploid induction rate, but applying short-term temperature stress makes the process 10 times more effective.
- Inducer Lines: Heat stress also increases haploid frequencies when combined with established inducer lines like CENH3 GFP-tailswap and CENH3 G83E.
2. Enhancement in Wide Crosses
The effect of temperature is well-documented in wide hybridization (crosses between different species). In crosses between Hordeum vulgare and Hordeum bulbosum, chromosome elimination is much higher at temperatures exceeding 20°C than at temperatures below 17.5°C. This indicates that heat facilitates the selective loss of the unwanted parental genome.
3. Biphasic Temperature Manipulation
Scientists have developed a “biphasic” strategy to use temperature to overcome biological barriers like male sterility in inducer lines:
- Initial Cooling: Lower temperatures are first used to improve pollen quality in otherwise sterile lines like the GFP-tailswap.
- Subsequent Heating: After crossing these improved pollen lines with wild-type plants, a temperature increase is used to trigger maternal haploids at a rate of approximately 25%.
4. Risks and Limitations
While higher temperatures boost induction rates, they can negatively impact seed viability. Heat-treated crosses often lead to endosperm failure (where the endosperm fails to cellularize) and result in seeds that fail to germinate. Furthermore, some modifications, such as CENH3 RNAi transformants, do not show any increase in haploid production when combined with heat stress, despite having reduced centromeric CENH3.
In summary, temperature is an emerging tool to enhance the frequency of uniparental genome elimination, though its success depends on the specific genetic background of the plant and must be balanced against the risk of reduced seed viability.
19. What are the meiotic stability issues with engineered chromosomes?
The meiotic stability of engineered chromosomes is a significant challenge in plant biotechnology, as these structures often exhibit reduced transmission rates and unpredictable inheritance compared to natural chromosomes. The primary issues stem from physical size constraints, defective pairing, and the consequences of structural rearrangements.
1. Size Constraints and Transmission
The exact minimum size for a stable engineered chromosome is unknown and likely species-specific, but it is heavily influenced by meiotic requirements.
- Transmission Rates: Engineered chromosomes generally have lower meiotic transmission rates than endogenous ones. For example, maize minichromosomes produced through truncation often deviate from Mendelian inheritance patterns.
- Gamete-Specific Efficiency: In Arabidopsis, transmission efficiency can vary significantly depending on the genetic background and whether the chromosome is passed through the male or female gametophyte.
- Replication Control: Extremely small chromosomes can sometimes lose replication control that is normally synchronized with the cell cycle, leading to further instability.
2. Defective Pairing and Segregation
The most common meiotic issue with minichromosomes is their inability to behave like standard chromosomes during division:
- Pairing Failure: Because of their small size, engineered chromosomes often struggle to find pairing partners (homologs) during meiosis.
- Sister Chromatid Cohesion: Small engineered chromosomes frequently exhibit precocious separation of sister chromatids during meiosis I. This leads to mis-segregation and the production of aneuploid progeny.
- Structural Vulnerability: In maize, circular or ring minichromosomes are notably unstable and suffer from impaired transmission, whereas in Arabidopsis, ring chromosomes like AtARC1 have been found to be more stable than their linear counterparts.
3. Stability Issues with Structural Rearrangements
Engineering large-scale changes like inversions introduces specific stability problems when the plant is in a hemizygous state (carrying one engineered and one wild-type chromosome):
- Reduced Fertility: Hemizygous inversion lines often show a significant reduction in viable seeds (e.g., a 1/3 reduction in Arabidopsis).
- Crossover Lethality: While pairing can occur, a single crossover within a large inversion leads to chromosomes with unbalanced genetic information (e.g., identical telomeric ends or massive deletions), resulting in non-viable gametes. Only double crossovers within the inversion or crossovers at the unchanged telomeric ends produce viable progeny.
- Recombination Suppression: Inversions effectively suppress genetic exchange across the rearranged region, which is useful for stabilizing traits but limits further natural breeding within that genomic area.
4. Centromere-Related Instability
Engineered centromeres, particularly those modified to create haploid inducers, are inherently unstable during meiosis to achieve their purpose:
- Aneuploidy: Incomplete genome elimination in these systems often results in a percentage of aneuploid progeny (plants with an irregular number of chromosomes).
- Meiotic Defects: Modifications to essential proteins like CENH3 often cause meiotic defects in the inducer parent, which can lead to sterility or reduced fitness before the cross even occurs.
Potential Solutions
To overcome these transmission and stability issues, researchers have suggested incorporating pollen selection genes onto engineered chromosomes. This would ensure that only the pollen grains carrying the engineered chromosome are viable, effectively forcing the stable inheritance of the cargo across generations.
20. What is the upper limit for chromosome size?
The upper limit for chromosome size is determined by mechanical constraints during cell division. Specifically, the longest chromosome arm must not exceed half the length of the spindle axis during mitotic telophase.
According to the sources, this limit is critical for the normal development of an organism. If a chromosome arm were to exceed this length, it would likely interfere with proper chromosome segregation and the stable maintenance of the genome during mitosis.
In contrast to this well-defined upper limit, the lower size limit for a stable minichromosome is currently unknown and is believed to be species-specific. It is likely determined by biological requirements such as:
- Sister chromatid cohesion and bivalent stability during meiosis.
- The size of the host plant’s genome.
- The ability to maintain replication control tied to the cell cycle.
21. What species-specific factors influence minichromosome stability?
Species-specific factors significantly influence the stability and transmission of engineered minichromosomes, with notable differences observed between model plants like Arabidopsis and major crops like maize. The primary factors include physical structure, biological size requirements, centromere epigenetics, and the host’s genetic background.
1. Physical Structure (Linear vs. Ring)
The shape of a minichromosome has a species-dependent impact on its stability:
- Arabidopsis thaliana: Ring minichromosomes (such as the 2.85 Mb AtARC1) appear to be more stably transmitted through sexual reproduction than comparable linear minichromosomes.
- Maize: Conversely, small ring chromosomes in maize are typically unstable, do not follow Mendelian inheritance, and their transmission is often impaired.
2. Minimum Size and Mechanical Limits
While there is a well-defined upper limit for chromosome size—where the longest arm must not exceed half the length of the spindle axis during mitosis—the lower limit is less clear and believed to be species-specific.
- Meiotic Requirements: The potential lower size limit is likely determined by species-specific requirements for proper sister chromatid cohesion or bivalent stability during meiosis.
- Host Genome Influence: Evidence suggests that the minimum size for stable segregation may depend on the overall size of the host plant’s genome.
- Replication Control: In maize, very small minichromosomes (particularly those derived from B chromosomes) have been observed to lose replication control that is normally synchronized with the cell cycle, leading to instability.
3. Centromere Epigenetics and Composition
The stability of minichromosomes depends heavily on functional centromeres, which are governed by both DNA sequence and epigenetic factors:
- Sequence Specificity: Plant centromeric repeats are often species-specific, which limits the ability to use cloned centromere components from one species to direct stable chromosome assembly in another.
- CENH3 Loading: Centromere identity is defined by the protein CENH3. In hybrid centromeres (such as the wheat-rye 1BL/1RS translocation), wheat CENH3 only incorporates into the rye-derived portion of the centromere, illustrating complex interactions that affect chromosome stability in hybrids.
- Epigenetic Influence: Simply transforming natural centromere sequences into plants is often insufficient to create function due to a strong epigenetic requirement for centromere maintenance that varies between species.
4. Host Genetic Background and Ploidy
The genomic environment of the host plant determines whether a minichromosome can be successfully maintained:
- Buffer Capacity: Truncated chromosomes that cause lethal gene dosage imbalances in diploid plants can often be stably inherited in autotetraploid, trisomic, or polyploid backgrounds because backup copies of essential genes are present. This has been successfully demonstrated in maize, Arabidopsis, and barley.
- Haploid Induction Efficiency: The efficiency of producing and recovering minichromosomes as by-products of uniparental genome elimination varies greatly by species. For instance, the CENH3 tailswap method is highly efficient in Arabidopsis (25–45%) but shows much lower rates in crops like maize, tomato, and rice (0.06% to 2.3%).
5. Environmental Factors
Environmental stress can impact the stability of chromosome inheritance during induction:
- Temperature: In wide crosses (e.g., Hordeum vulgare × H. bulbosum) and cenh3 mutants, high temperatures (above 20°C) can drastically increase the frequency of chromosome elimination, which affects the stability and recovery of the desired chromosome set.
22. What is telomere seeding and how does it work?
Telomere seeding (also known as telomere-mediated chromosomal truncation or TMCT) is a “top-down” chromosome engineering technique used to truncate or shorten existing native chromosomes to create stable, gene-poor minichromosomes. It works by integrating cloned telomeric repeats into a specific genomic site, which then “seeds” or nucleates the formation of a new, functional telomere.
How Telomere Seeding Works
The biological mechanism involves several key steps:
- Introduction of Telomeric Repeats: Researchers introduce a DNA construct—typically via Agrobacterium-mediated transformation or particle bombardment (gene gun)—that contains a block of cloned telomeric repeats. In plants, these repeats usually follow the conserved Arabidopsis-type sequence (5′-TTTAGGG-3′).
- Nucleation of a New Telomere: When this construct integrates into a chromosome arm, the cell’s telomerase enzyme recognizes the DNA end carrying the repeats and adds more repeat units to synthesize a functional telomere array. This effectively “caps” the chromosome at the new integration site.
- Chromosomal Truncation: Because a new telomere has been established, the original distal portion of the chromosome (the part further away from the centromere) is “released” as an acentric fragment. Lacking a centromere, this fragment cannot segregate properly and is eventually lost during subsequent cell divisions.
- Formation of a Truncated Chromosome: The remaining portion of the chromosome, which retains its original endogenous centromere, becomes a stable truncated chromosome. These structures are often engineered to contain selection markers and site-specific recombination sites (like Cre-loxP) to allow for the future “retrofitting” or stacking of new genes.
Applications and Precision
- Creating Minichromosomes: The primary goal is to produce phenotypically neutral minichromosomes. By removing large arms that contain essential host genes, researchers avoid gene dosage imbalances, though this often requires using polyploid or trisomic backgrounds that can “buffer” the loss of essential genetic material.
- Targeting B Chromosomes: Supernumerary B chromosomes are ideal targets for telomere seeding because they are non-essential and do not cause deleterious effects when truncated.
- Success Across Species: This method has been successfully applied to major crops and model plants, including maize, Arabidopsis, barley, rice, wheat, and rapeseed.
- Emerging Precision with CRISPR: While traditional telomere seeding occurs at random integration sites, modern advancements suggest that CRISPR/Cas9 can be used to induce targeted double-strand breaks (DSBs) at preselected locations. This would allow for the precise deletion of chromosome arms while retaining specific desired elements.
Once established, these truncated chromosomes are stable during mitosis and meiosis, acting as independent linkage groups that can carry large packages of beneficial genes into new crop varieties.
23. What is telomere seeding and how does it work?
Telomere seeding, also known as telomere-mediated chromosomal truncation (TMCT), is a “top-down” chromosome engineering technique used to create minichromosomes by shortening existing native chromosomes. The process works by introducing cloned telomeric repeats into a plant cell, which then “seed” or nucleate the formation of a new, functional telomere at the site of DNA integration.
The mechanism involves several specific biological steps:
- Introduction of Repeats: Researchers introduce a DNA construct containing tandem arrays of cloned telomeric repeats, such as the conserved Arabidopsis-type sequence (5′-TTTAGGG-3′), into the cell via methods like Agrobacterium-mediated transformation or particle bombardment.
- Telomere Nucleation: When this construct integrates into a chromosome arm, the cell’s endogenous telomerase enzyme recognizes the DNA end carrying these repeats and adds more units to synthesize a functional telomere cap.
- Chromosomal Truncation: Because a new telomere has been established at an internal site, the original distal portion of the chromosome arm (the section further from the centromere) is released as an acentric fragment. Lacking its own centromere, this fragment cannot segregate properly and is eventually lost during subsequent cell divisions.
- Formation of a Stable Vector: The remaining part of the chromosome, which retains its original endogenous centromere, becomes a stable truncated chromosome. These engineered chromosomes are often designed to include selection markers and site-specific recombination sites (like Cre-loxP) for future gene stacking.
While traditional telomere seeding occurs at random integration sites, modern advancements using CRISPR/Cas9 allow researchers to target double-strand breaks at preselected locations to make the truncation process precise and predictable. This technique has been successfully applied to produce stable minichromosomes in diverse species, including maize, rice, Arabidopsis, barley, wheat, and rapeseed. To ensure these truncated chromosomes are not lost due to the deletion of essential genes, they are often maintained in polyploid or trisomic genetic backgrounds that provide “buffer” copies of the missing genetic material.
24. How does colchicine induce chromosome doubling in haploids?
Colchicine induces chromosome doubling in haploids by acting as a microtubule inhibitor that disrupts the physical machinery required for cell division.
The specific biological mechanism works as follows:
- Inhibition of Spindle Formation: During normal cell division (mitosis), spindle fibers bind to the kinetochores of chromosomes and pull sister chromatids to opposite poles of the cell. Colchicine disrupts microtubule synthesis, which prevents these spindle fibers from forming correctly.
- Prevention of Chromosome Segregation: Because the spindle apparatus is inhibited, the chromosomes cannot be separated and moved into two distinct daughter cells.
- Doubling the Chromosome Set: Instead of dividing into two cells with one set of chromosomes each, the cell retains the entire doubled set of genetic material within a single nucleus. This effectively converts the sterile haploid state (n) into a fertile diploid state (2n).
Significance in Breeding
The use of colchicine is a critical step in creating doubled haploid (DH) lines. This process allows breeders to achieve 100% homozygosity in a single generation, bypassing the 8–10 generations of backcrossing and selection required in traditional breeding to stabilize traits. Once the chromosome set is doubled, the previously sterile haploid plant becomes fertile and can be used to produce true-breeding progeny.
25. Can CRISPR/Cas9 be used to target telomere seeding precisely?
Yes, CRISPR/Cas9 can be used to target telomere seeding precisely, transforming what was once a random process into a predictable tool for chromosome engineering.
The Shift from Random to Precise Seeding
Traditional telomere seeding (also known as telomere-mediated chromosomal truncation or TMCT) relies on the random integration of telomeric repeats into the genome. While effective for shortening chromosomes, this method requires screening a large number of transgenic plants to find one where the truncation occurred at a desirable location.
CRISPR/Cas9 introduces precision through the following mechanism:
- Targeted Double-Strand Breaks (DSBs): CRISPR/Cas9 is used to induce a site-specific DSB at a preselected genomic location.
- Integration at the Break Site: A DNA construct containing telomeric repeats is introduced and integrated specifically at this DSB during the cell’s natural DNA repair process.
- Nucleation and Truncation: Once integrated, these repeats serve as a “seed” for telomerase to form a new, functional telomere, effectively capping the chromosome at the target site and releasing the distal arm as an acentric fragment that is subsequently lost.
Applications of Targeted Telomere Seeding
This precision allows researchers to “sculpt” chromosomes with high accuracy:
- Customized Minichromosomes: Breeders can create minichromosomes of predefined lengths by selecting the exact site of truncation.
- Controlled Deletion of Arms: It enables the targeted removal of specific chromosome arms while retaining essential elements like the endogenous centromere.
- Trait Stacking Platforms: These precisely engineered chromosomes can then be equipped with site-specific recombination sites (like Cre-lox) to serve as stable “cargo” vectors for stacking multiple beneficial genes.
While the sources note that these targeted approaches are still being optimized for widespread use in complex crops, the combination of CRISPR/Cas9 with telomere seeding is a key advancement toward precise karyotype restructuring.
26. What is telomere-mediated chromosomal truncation?
Telomere-mediated chromosomal truncation (TMCT), also frequently referred to as telomere seeding, is a “top-down” chromosome engineering technique used to shorten existing native chromosomes to create stable, gene-poor minichromosomes. It is a powerful tool for generating plant artificial chromosomes (PACs) that can serve as high-capacity vectors for gene stacking.
Mechanism of Action
The process works by exploiting the natural function of telomeres, which are the tandemly repeated sequences that cap the ends of linear chromosomes to protect them from deterioration or fusion.
- Introduction of Repeats: Researchers introduce a DNA construct—often via Agrobacterium-mediated transformation or biolistics—that contains a block of cloned telomeric repeats. In most plants, these repeats follow the conserved Arabidopsis-type sequence (5′-TTTAGGG-3′).
- Nucleation of a New Telomere: When this construct integrates into a chromosome arm, the cell’s telomerase enzyme recognizes the repeats and adds more units to synthesize a functional telomere array. This effectively “seeds” a new chromosome end at the site of integration.
- Chromosomal Breakage: Because a new telomere cap has been established at an internal site, the original distal portion of the chromosome (the part further from the centromere) is released as an acentric fragment.
- Loss of Genetic Material: Lacking its own centromere, this fragment cannot attach to spindle microtubules and is eventually lost during subsequent cell divisions.
Resulting Structure
The remaining portion of the chromosome, which retains its original endogenous centromere, becomes a stable truncated chromosome. These engineered minichromosomes are often designed to include selection markers and site-specific recombination sites (such as Cre-loxP or FLP-frt) to allow for the future addition or “retrofitting” of beneficial genes.
Applications and Advantages
- Creating Minichromosomes: The primary goal is to produce phenotypically neutral vectors that act as independent linkage groups, segregating independently of the host’s native chromosomes.
- Avoiding Linkage Drag: Because they are independent, the stacked genes on a minichromosome can be transferred to new varieties in a single cross without bringing along unwanted genetic material from the donor genome.
- Targeting B Chromosomes: Researchers often target B chromosomes for truncation because they are supernumerary and non-essential, meaning their truncation does not cause the deleterious effects or lethality that often occurs when essential A chromosomes are shortened.
Precision with CRISPR/Cas9
While traditional telomere seeding occurs at random integration sites, modern advancements allow for precise, targeted truncation. By using CRISPR/Cas9 to induce a double-strand break at a preselected genomic location, researchers can direct the telomere-seeding construct to integrate at that exact spot, allowing for the creation of minichromosomes with predefined lengths and gene content.
Challenges
The main hurdle for TMCT in diploid plants is gametophyte lethality; removing large sections of a standard chromosome often results in the loss of essential genes. To overcome this, the technique is typically performed in polyploid or trisomic genetic backgrounds, which provide “buffer” copies of the missing genetic information.
References
Chan, S. W. L. (2010). Chromosome engineering: Power tools for plant genetics. In Trends in Biotechnology (Vol. 28, Number 12, pp. 605–610). https://doi.org/10.1016/j.tibtech.2010.09.002
Gehrke, F., Schindele, A., & Puchta, H. (2022). Nonhomologous end joining as key to CRISPR/Cas-mediated plant chromosome engineering. Plant Physiology, 188(4), 1769–1779. https://doi.org/10.1093/plphys/kiab572
Knox, O., Backhouse, D., & Gupta, V. (2020). A comparative study of field nematode communities over a decade of cotton production in Australia. Agronomy, 10(1). https://doi.org/10.3390/agronomy10010122
Kwiatek, M. T., Noweiska, A., Bobrowska, R., Czapiewska, A., Aygün, M., Munyamahoro, F. d’Assise, Mikołajczyk, S., Tomkowiak, A., Kurasiak-Popowska, D., & Poślednik, P. (2023). Novel Tetraploid Triticale (Einkorn Wheat × Rye)—A Source of Stem Rust Resistance. Plants, 12(2). https://doi.org/10.3390/plants12020278
Liu, Y., Liu, Q., Yi, C., Liu, C., Shi, Q., Wang, M., & Han, F. (2025). Past innovations and future possibilities in plant chromosome engineering. In Plant Biotechnology Journal (Vol. 23, Number 3, pp. 695–708). John Wiley and Sons Inc. https://doi.org/10.1111/pbi.14530
Mabuza, L. M., Mchunu, N. P., Crampton, B. G., & Swanevelder, D. Z. H. (2023). Accelerated Breeding for Helianthus annuus (Sunflower) through Doubled Haploidy: An Insight on Past and Future Prospects in the Era of Genome Editing. In Plants (Vol. 12, Number 3). MDPI. https://doi.org/10.3390/plants12030485
Mette, M. F., & Houben, A. (2015). Engineering of plant chromosomes. Chromosome Research, 23(1), 69–76. https://doi.org/10.1007/s10577-014-9449-1
Ojha, A., Zhang, F., & Patil, G. B. (2023). Genome editing and chromosome engineering in plants. In Plant Genome (Vol. 16, Number 2). John Wiley and Sons Inc. https://doi.org/10.1002/tpg2.20352
Pershina, L. A. (2014). Plant chromosome engineering is an area of biotechnology. Russian Journal of Genetics: Applied Research, 4(4), 311–317. https://doi.org/10.1134/S207905971404008X
Puchta, H., & Houben, A. (2024). Plant chromosome engineering – past, present and future. In New Phytologist (Vol. 241, Number 2, pp. 541–552). John Wiley and Sons Inc. https://doi.org/10.1111/nph.19414
Rönspies, M., Schindele, P., & Puchta, H. (2021). CRISPR/Cas-mediated chromosome engineering: Opening up a new avenue for plant breeding. In Journal of Experimental Botany (Vol. 72, Number 2, pp. 177–183). Oxford University Press. https://doi.org/10.1093/jxb/eraa463
Rönspies, M., Schmidt, C., Schindele, P., Lieberman-Lazarovich, M., Houben, A., & Puchta, H. (2022). Massive crossover suppression by CRISPR–Cas-mediated plant chromosome engineering. Nature Plants, 8(10), 1153–1159. https://doi.org/10.1038/s41477-022-01238-3
Sairam Nagalla, L., Ojha Kshetry, A., Pagare, R., Janga, M. R., & Patil, G. B. (n.d.). Chromosome engineering and Prospectives in Plant Breeding. In Next Generation Plant Breeding (21).
Yaşar, S., Gehrke, F., Capdeville, N., & Puchta, H. (2026). Recent progress in plant genome engineering: from large insertions to chromosome number changes. In Current Opinion in Biotechnology (Vol. 97). Elsevier Ltd. https://doi.org/10.1016/j.copbio.2025.103426
Yu, W., Yau, Y. Y., & Birchler, J. A. (2016). Plant artificial chromosome technology and its potential application in genetic engineering. In Plant Biotechnology Journal (Vol. 14, Number 5, pp. 1175–1182). Blackwell Publishing Ltd. https://doi.org/10.1111/pbi.12466
Zhou, J., Liu, Y., Guo, X., Birchler, J. A., Han, F., & Su, H. (2022). Centromeres: From chromosome biology to biotechnology applications and synthetic genomes in plants. In Plant Biotechnology Journal (Vol. 20, Number 11, pp. 2051–2063). John Wiley and Sons Inc. https://doi.org/10.1111/pbi.13875