1. Strategic Introduction to KASP Genotyping
In the current era of genomic-assisted breeding, Kompetitive Allele-Specific PCR (KASP) has transitioned from a specialized alternative to the global benchmark for targeted SNP genotyping. While high-density arrays and Next-Generation Sequencing (NGS) remain the standards for de novo SNP discovery, KASP provides the necessary scalability and precision for routine Marker-Assisted Selection (MAS). By efficiently replacing labor-intensive, gel-based markers, KASP enables the high-throughput characterization of breeding populations with superior turnaround times. This technology is strategically vital for modern crop improvement, facilitating the rapid conversion of genomic data into actionable selection decisions through its unique biochemical flexibility and cost-efficiency.

Comparison of Genotyping Technologies
| Feature | Gel-Based PCR | Chip-Based (GoldenGate) | KASP Genotyping |
|---|---|---|---|
| Speed | Slow (Manual/Labor-intensive) | Fast (Highly automated) | Ultra-Fast (45x faster than gels) |
| Cost per Data Point | High (Due to labor) | Moderate to High | Low (approx. €0.20 per assay) |
| Flexibility | High (Targeted) | Inflexible (Fixed SNP sets) | High (Any sample/Any SNP) |
| Error Rate | High/Subjective | 2.0% – 2.4% | Low (0.7% – 1.6%) |

Comparison of Genotyping Technologies
| Technology | Speed / Throughput | Cost | Primary Limitation |
|---|---|---|---|
| AS-PCR (Allele-Specific) | Low | Moderate | Dominant marker system; cannot distinguish heterozygotes from homozygotes. |
| CAPS (Cleaved Amplified Polymorphic Sequences) | Low | High | Labor-intensive; relies on expensive restriction enzymes and gel electrophoresis. |
| TS-PCR (Temperature-Switch) | Moderate | Moderate | Requires secondary detection steps like high-resolution melting or gels. |
| SNP Arrays (Affymetrix/Illumina) | Ultra-High | High per sample | Inflexible; optimized for genome-wide scans rather than specific trait selection. |
| KASP (Kompetitive Allele-Specific PCR) | High | Low | Requires distinct primer design for each target locus. |

Technical Performance Comparison: KASP vs. Illumina GoldenGate
| Feature | KASP (Uniplex) | Illumina GoldenGate (Multiplex) |
|---|---|---|
| Genotyping Error Rates | 0.7% – 1.6% (Mean: 1.1%) | 2.0% – 2.4% (Mean: 2.2%) |
| NTC Behavior | Consistent, isolated clustering; rarely exceeds threshold. | Erratic; frequently clusters with genotypes, creating ambiguity. |
| Assay Conversion Success | 98%–100% design success; ~93% working assay rate. | Lower conversion; 81% transferability of existing SNPs in maize. |
| Unscoreable Data Rate | Minimal; sharper plots for single-plant DNA. | 10.9% of SNPs often unscoreable; up to 19.9% discarded. |

Comparative Analysis: Legacy vs. Unified KASP Platform
| Feature | Legacy PCR Markers (AFLP, SCAR, CAPS) | KASP Genotyping Platform |
|---|---|---|
| Throughput | Low; bottlenecked by gel electrophoresis and manual handling. | High; automated for 96-, 384-, and 1,536-well formats. |
| Data Quality | Variable sensitivity; prone to qualitative interpretation errors. | High; quantitative fluorescence-based allele discrimination. |
| Reproducibility | Moderate; sensitive to inter-laboratory protocol variance. | Superior; benchmarked error rates of 0.7–1.6% (vs. 2.0–2.4% for GoldenGate). |
| Flexibility | Inflexible; requires unique chemistry for every locus. | Superior; optimized for “Single-plex” high-throughput screening. |
| Genomic Utility | Primarily linked markers; limited functional precision. | High; enables direct interrogation of functional polymorphisms. |
2. Technical Architecture and Mechanism of Action
KASP is defined as a homogeneous, uniplex, fluorescence-based genotyping technology. Its primary technical advantage lies in its “Kompetitive” nature: the assay utilizes two allele-specific forward primers, each carrying a unique tail (FAM or HEX), which compete for the same target site on the template DNA.
The Biochemical Process
The signal generation is driven by Fluorescence Resonance Energy Transfer (FRET). The reaction mix contains a common reverse primer and two FRET cassettes—one labeled with FAM and the other with HEX. During the initial cycles of PCR, the allele-specific forward primer successfully binds and extends, incorporating its specific tail into the amplicon. In subsequent cycles, the complementary FRET cassette binds to this tail, releasing the fluorophore from its quencher and generating a detectable fluorescent signal. This all-in-one-tube biochemistry ensures a streamlined workflow with minimal risk of contamination.
Operational Specifications
• Reaction Formats: Highly scalable architecture supporting 96-, 384-, and 1,536-well plate formats.
• Template DNA Versatility: Robust performance across diverse sources, including fresh, frozen, lyophilized, and high-value “seed-derived” DNA (seed chips) for non-destructive early-generation screening.
• Assay Design and Nucleotide Diversity: In highly heterozygous crops like potato, the presence of “off-target” flanking SNPs can lead to “null-alleles” by disrupting primer binding. To mitigate this, specialists must characterize flanking diversity and utilize IUPAC ambiguity codes in primer design to ensure robust annealing across diverse germplasm.

The Molecular Toolkit: Essential Components
Your toolkit for SNP detection begins with three essential primers and a specialized Master Mix. The precision of KASP relies on the unique design of these oligonucleotides.
The Three-Primer System
• Two Allele-Specific Forward Primers: Each primer is designed to target one of the two possible nucleotides at the SNP site.
◦ The 3’ End (The Anchor): This is the most critical part. The very last base on the 3’ end corresponds exactly to the SNP. Think of this as the “ignition”—if the 3’ base does not match the target DNA perfectly, DNA polymerase cannot begin extension.
◦ The 5’ Tail (The Handle): These primers have a unique, non-complementary 5’ sequence. One tail is designed to eventually attract a FAM signal, while the other attracts a HEX signal.
• One Common Reverse Primer: This primer binds downstream from the SNP. It is designed so that the total amplicon length (the resulting DNA fragment) is typically less than 120 base pairs (bp), ensuring a fast and efficient reaction.
The KASP Master Mix
The reaction environment contains specialized FRET cassettes. These consist of a fluorescent Reporter (FAM or HEX) and a Quencher. In their default state, these cassettes form a “hairpin” structure that keeps the reporter and quencher in close proximity, effectively silencing any light signal.
With these components in place, the laboratory process enters a specialized thermal cycling phase where “competition” determines the outcome.
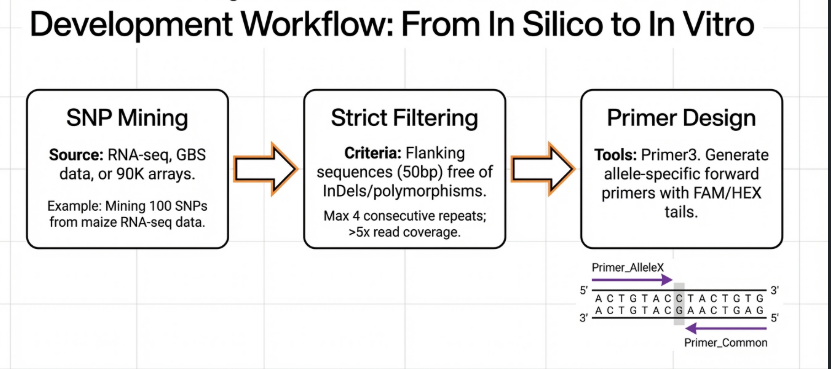
The Mechanism: A Three-Step Reaction Narrative
The “Kompetitive” nature of KASP stems from the fact that both forward primers are present in the same well, vying for the same binding site.
Phase 1: The Competition (Cycle 1)
The two allele-specific primers compete to hybridize to the target SNP. As noted, the 3’ end acts as the anchor; extension only occurs if the primer is a perfect match. If a plant is homozygous for Allele 1, only the FAM-tailed primer will initiate the synthesis of a new DNA strand.
Phase 2: The Complement (Cycle 2)
This is the “missing link” of the reaction. During the second cycle, the Common Reverse Primer binds to the newly synthesized strand and replicates it, including the unique 5’ tail sequence. This creates a double-stranded, complementary binding site for the FRET cassettes found in the Master Mix.
Phase 3: The Signal Generation (Subsequent Cycles)
As PCR continues, the amount of DNA containing the tail-complement grows exponentially. The FRET cassettes in the Master Mix now have a place to bind.
The success of this molecular competition is ultimately revealed through the “magic” of fluorescence resonance energy transfer.
The ‘Magic’ of FRET: From DNA to Light
Fluorescence Resonance Energy Transfer (FRET) is the physics-based “messenger” that tells us which allele was successfully extended.
The Quencher-Reporter Physics
1. Quenched State: In the Master Mix, the FRET cassette is in a hairpin loop. The Quencher absorbs all energy from the Reporter, resulting in no fluorescence.
2. Unquenching through Hybridization: When the cassette encounters the specific tail-complement generated in Phase 2, it opens up and hybridizes to the DNA.
3. Signal Emission: As the hairpin opens, the Reporter is physically separated from the Quencher. It is now “unquenched” and will emit a fluorescent signal (FAM/Blue or HEX/Green) when excited by the plate reader’s laser.
Insight Highlight: Because KASP is a uniplex assay using two distinct dyes, it provides a co-dominant result. This means we can distinguish not just “Yes/No” for a trait, but whether a sample is homozygous for either allele or a hybrid (heterozygous) containing both.
Once the light signals are emitted, they are captured and plotted to reveal the plant’s genetic identity.
Decoding the Data: Signal Clusters and Scatter Plots
KASP data is visualized on a Normalized X-Y Scatter Plot. Normalization is key; researchers use a Passive Reference Dye (such as ROX) in the Master Mix to account for slight well-to-well variations in volume or evaporation, ensuring the data is clean and comparable.

Mapping Cluster Position to Genotype
The software plots the FAM signal on the X-axis and the HEX signal on the Y-axis.
| Cluster Position | Color | Genotype Calling |
|---|---|---|
| X-axis cluster | Blue | Homozygous for Allele 1 (High FAM, Low HEX) |
| Y-axis cluster | Red | Homozygous for Allele 2 (Low FAM, High HEX) |
| Diagonal/Middle | Green | Heterozygous (Signal from both FAM and HEX) |
| Origin (0,0) | Black | Non-Template Control (NTC) / PCR Failure |
3. Comparative Performance: KASP vs. Traditional and Chip-Based Platforms
Selecting a genotyping platform is a critical economic decision; the choice must balance per-sample costs with data reliability. For targeted applications, KASP offers significant advantages over chip-based platforms like Illumina GoldenGate or BeadXpress.
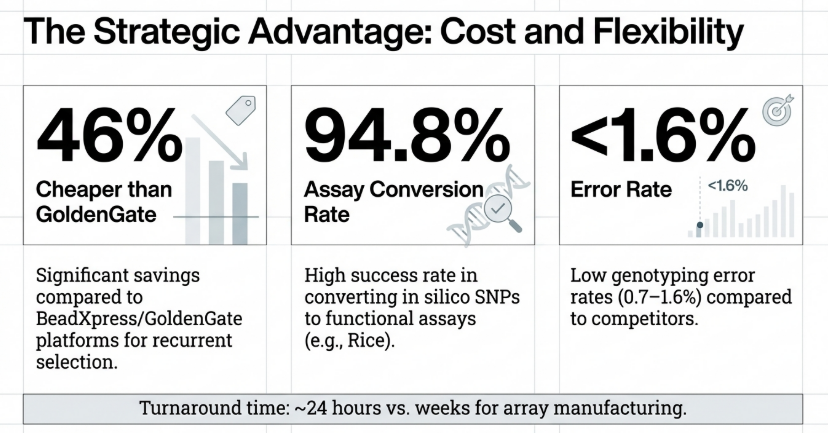

Technical and Economic Benchmarks
• Accuracy: KASP demonstrates a superior accuracy profile, with error rates in positive controls ranging from 0.7% to 1.6%, compared to the 2.0–2.4% observed in GoldenGate platforms.
• Assay Success Rate: As the source context notes: “KASP achieves higher assay design success rates (98–100%) and conversion to successful working assays (93–94%) than TaqMan (72 and 61%, respectively).”
• Cost and Speed Efficiency: For applications such as Marker-Assisted Recurrent Selection (MARS), KASP is 7.9% to 46.1% more cost-effective. Furthermore, in wheat breeding, KASP assays have proven to be 45 times faster than traditional gel-based PCR systems.
• Control Reliability: KASP provides superior signal-to-noise ratios. Its No-Template Controls (NTCs) cluster consistently together and rarely cross false-positive thresholds, whereas GoldenGate NTCs often cluster erratically, complicating automated allele calling.
4. High-Throughput Screening in Bread Wheat (Triticum aestivum)
The development of KASP has enabled the first-of-its-kind report on the high-throughput screening of a vast library of functional genes in a major cereal crop. By converting Functional Markers (FMs) into a validated 70-assay KASP toolkit, breeders can now characterize crossing parents and advanced lines with unprecedented technical depth.
Validated Functional Gene Assays
The toolkit targets specific gene symbols underpinning economically critical traits:
• Adaptability: Photoperiod sensitivity (Ppd-D1, Ppd-B1) and Vernalization (Vrn-A1, Vrn-B1, Vrn-D1).
• Grain Yield and Quality: Plant height (Rht-B1b, Rht-D1b), Thousand Kernel Weight (TaGS-D1, TaSus2-2B, TaCwi-5D, TaTGW6-A1), and Grain Hardness (Pina-D1, Pinb-D1).
• Biotic/Abiotic Stress: Resistance genes for leaf and stripe rust (Lr14a, Yr15) and drought tolerance markers (TaDreb-B1).
These KASP assays provide a robust alternative to inflexible high-density arrays, allowing for the precise movement of specific alleles into elite backgrounds.

5. Implementation in Potato Breeding: Disease Resistance and PoolSeq
Genotyping the autotetraploid potato is complicated by high heterozygosity and complex dosage effects. To address this, the “PoolSeq” approach is utilized to convert diverse marker types (AFLPs, SCARs) into the KASP platform.
The PoolSeq Methodology
The logic of PoolSeq involves constructing DNA pools based on the presence or absence of a diagnostic marker (e.g., the 57R marker for H1). These pools undergo whole-genome resequencing and are aligned to the potato reference genome (v4.04). This allows for the identification of candidate SNPs unique to the marker-positive pool while simultaneously characterizing flanking variation to avoid “off-target” binding failures.
Strategic Deployment and Dosage Determination
KASP has been successfully deployed for critical resistance genes:
• H1 Gene: Resistance to the nematode Globodera rostochiensis (Ro1 and Ro4).
• R2 Gene: Resistance to Phytophthora infestans (Late Blight).
The Strategic Layer: KASP’s ability to generate distinct clusters facilitates dosage determination (distinguishing simplex from duplex states). Identifying duplex parents is a strategic priority; these parents ensure a significantly higher inheritance frequency of the resistance gene in progeny, directly reducing the population size a breeder must maintain to find resistant individuals.
6. Operational Applications: QC, QTL Mapping, and MARS
KASP serves as the operational backbone for multiple breeding logistics:
• Quality Control (QC): Essential for assessing genetic identity and purity. A subset of 50–100 SNPs is typically used to ensure inbred lines are homogeneous and F1 hybrids are true-to-type.

• QTL Mapping: KASP facilitates the mapping of complex traits in bi-parental populations, typically requiring 200 to 400 polymorphic markers.
• Marker-Assisted Recurrent Selection (MARS): The high speed of KASP enables “pre-flowering selection.” This allows for rapid cycle recombination of polygenic traits, such as drought tolerance, within a single growing season.
• Allele Mining: Screening large germplasm collections for favorable functional polymorphisms identified via GWAS.
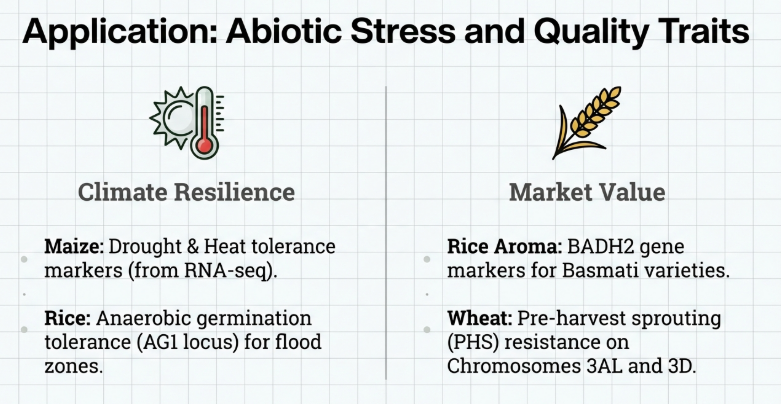


Laboratory Best Practices
To minimize scoring errors, specialists utilize “touchdown PCR” protocols (e.g., 65–57°C or 61–55°C) and “KlusterCaller” software. Manual inspection via “SNPviewer” is mandatory to resolve ambiguous clusters or unassigned calls before finalizing selection data.
7. Conclusion and Strategic Takeaways
KASP genotyping represents a transformative shift in agricultural molecular biology, providing a robust, scalable, and cost-effective solution for targeted marker selection. The conversion of Functional Markers (FMs) into KASP assays—particularly the 70-assay toolkit for wheat and the PoolSeq-derived markers for potato—allows breeding programs to maximize genetic gains with high precision. Integrating KASP into early-generation breeding cycles is essential for accelerating selection efficiency and ensuring the rapid deployment of disease resistance and yield-enhancing alleles.
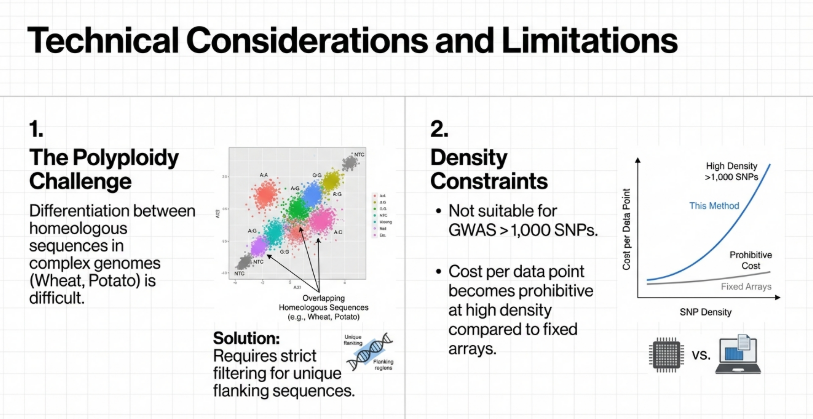
KASP is not just a lab exercise; it is the “workhorse” of modern food security. In studies of wheat and potato, three advantages stand out:
1. Throughput & Error Management: KASP’s Non-Template Controls (NTCs) cluster much more tightly than those in chip-based systems. This precision leads to significantly lower error rates (as low as 0.7%), which is vital when screening thousands of breeding lines for disease resistance.
2. Extreme Speed for MAS: KASP is 45 times faster than traditional gel-based PCR markers. In Marker-Assisted Selection (MAS), this speed allows breeders to genotype and select the best plants within a single growing season, drastically accelerating the release of new varieties.
3. Dosage Determination in Polyploids: In complex crops like the tetraploid potato, the “dosage” of a gene matters immensely. For example, the cultivar FONTANE is duplex for the H1 nematode resistance gene (carrying two copies). Using KASP to identify duplex parents is a game-changer for breeders because a duplex parent passes the resistance gene to a much higher percentage of its offspring than a “simplex” (single-copy) parent.
Final Insight: KASP’s flexibility—genotyping many samples for a few high-value loci at a cost of only €0.20 per assay—perfectly complements high-density SNP arrays. While arrays map the entire “territory” of a genome, KASP is the precision tool used to select the specific “winners” that will feed the next generation.

8. From Gels to Gold: 5 Surprising Ways KASP is Transforming the Future of Food
8.1 Introduction: The High-Stakes Race for Better Crops
For decades, the front lines of agricultural science were a messy affair. Breeders spent years in the field, waiting for seasons to turn, or in labs peering at dripping, translucent gels to catch a glimpse of a plant’s genetic potential. But as the global population climbs toward 10 billion, that analog pace is no longer sustainable. We are in a high-stakes race to build more resilient, higher-yielding crops, and traditional breeding—reliant on physical observations and labor-intensive molecular tools—is simply too slow.
The solution is the digitization of the genome. Leading this “silent revolution” is KASP (Kompetitive Allele Specific PCR), a high-throughput technology that allows scientists to unmask the genetic secrets of a seed before it even hits the soil. By shifting the focus from the unpredictable field to the fluorescent precision of the lab, KASP is transforming the global food supply chain into a streamlined data pipeline.
8.2 Takeaway 1: The Need for Speed (45x Faster Than Traditional Methods)
In the legacy era of biotechnology, identifying a specific gene required a series of grueling steps. Scientists had to rely on “gel-based” markers, a process involving restriction enzyme digestion and gel electrophoresis—essentially separation by hand that acted as a massive bottleneck.
KASP has shattered that ceiling. By moving to an automated, high-throughput platform, research into bread wheat genes has shown that KASP assays are 45 times faster than those traditional methods. Because KASP eliminates the need for manual gel separation and enzyme digestion, it transforms a process that once took weeks into a real-time decision-making tool. In a single growing season, breeders can now characterize parents and advance lines with a speed that was once unthinkable.
“Such assays could greatly accelerate the characterization of crossing parents and advanced lines for marker-assisted selection and can complement the inflexible, high-density SNP arrays.” — Rasheed et al.
8.3 Takeaway 2: Precision at Scale (Beating the “Gold Standard”)
While massive “multiplex” chips—which test thousands of genetic points at once—generate headlines, they often produce messy, erratic datasets. KASP is a “uniplex” platform, meaning it focuses on specific genetic points with surgical precision. When compared to the Illumina GoldenGate platform, the former “gold standard,” KASP consistently wins on accuracy.
KASP’s average genotyping error rate sits between 0.7–1.6%, significantly cleaner than the 2.0–2.4% error rate seen in GoldenGate. This precision is driven by KASP’s superior signal-to-noise ratio:
• Reliable Controls: In KASP assays, the “no-template” controls (NTCs) consistently cluster together, rarely producing false signals. In contrast, GoldenGate NTC signals can be highly erratic, showing a mismatch rate as high as 44.6% in some tests.
• High Transferability: KASP is remarkably versatile; roughly 81% of SNPs from other custom assays can be successfully migrated to the KASP platform without loss of data quality.
• Reduced False Calls: Because the chemistry is more stable, the “no-template” failures that plague other platforms are virtually non-existent, ensuring that a “null” result actually means the gene is absent, not that the test failed.
8.4 Takeaway 3: The “Cents-per-Sample” Economy
Advanced genetics was once the exclusive domain of massive multinational corporations. KASP is democratizing the field by triggering an “economy of scale” that brings costs down to a few cents. By outsourcing both DNA extraction and genotyping, regional programs can now operate at a professional tier.
The numbers are startling: MAS (Marker-Assisted Selection) costs can drop to just €0.20 ($0.22) per assay. Logistically, this means a small team of just five people can sample thousands of plant lines in a single day. This efficiency makes it economically viable for smaller labs to use advanced genetics to protect regional food staples like potatoes and maize.
KASP Cost Savings over BeadXpress and GoldenGate:
• 7.9% to 46.1% lower total genotyping costs.
8.5 Takeaway 4: The Universal Translator for Genetic Data
One of the most ingenious aspects of KASP is its role as a “universal translator.” For years, valuable agricultural data was locked in old, “anonymous” marker formats like AFLP or SCAR markers—data that was hard to read and even harder to integrate into modern pipelines.
Using a methodology called PoolSeq, researchers can now “reverse-engineer” this legacy data. By constructing DNA pools based on the presence or absence of a specific marker, scientists use KASP to hunt for “candidate nucleotide polymorphisms” in regions where they previously only had messy, outdated info. This process takes the “legacy codebase” of our food supply and converts it into clean, digital SNP-based markers that are compatible with 21st-century high-throughput pipelines.
8.6 Takeaway 5: Seeing in “Dosage” (The Power of Three Colors)
A KASP result is visualized as a scatter plot with dots of red, blue, and green. The X-axis (FAM fluorescence) and Y-axis (HEX fluorescence) represent the different versions of a gene. While most tests only tell you if a plant has a gene, KASP reveals the “dosage.”
In complex crops like the potato, this is a game-changer. Breeders can look at the clusters on the plot and see not just a “simplex” (one copy) of a resistance gene, but a third cluster indicating a “duplex” or “triplex” (multiple copies). For example, the potato cultivar Fontane was identified as a duplex carrier for resistance. Identifying these “high-dosage” plants allows breeders to select parents that will pass on stronger, more resilient traits to nearly 100% of their offspring.
Image Summary





Questions/Answers
1. How does KASP technology accelerate marker-assisted breeding programs?
Kompetitive Allele-Specific PCR (KASP) technology accelerates marker-assisted breeding (MAB) programs by providing a high-throughput, cost-effective, and highly precise platform for single nucleotide polymorphism (SNP) genotyping. It streamlines the selection process by allowing breeders to screen thousands of samples for desirable traits quickly, bypassing the need for time-consuming and inconsistent phenotypic evaluations.
Several specific ways KASP technology drives this acceleration:
1. High-Throughput Efficiency and Speed
KASP is significantly faster than traditional molecular methods. One study noted that KASP assays offer 45 times superiority in speed compared to traditional gel-based PCR markers.
• Gel-Free System: Unlike traditional simple sequence repeat (SSR) markers, KASP is a homogeneous, closed-tube, and gel-free system. This eliminates the labor-intensive fragment analysis and gel electrophoresis steps that often create bottlenecks in breeding programs.
• Rapid Turnaround: Genotyping results can be obtained in just a few days, allowing for real-time decision-making during a single growing or crossing season.
2. Cost-Effectiveness
KASP is one of the most affordable genotyping solutions, particularly when screening a moderate number of markers across a large population.
• Lower Consumable Costs: The technology uses universal fluorescent primers, which avoids the high cost of synthesizing expensive labeled probes for every individual SNP.
• Scalability: The cost per data point decreases as the number of samples increases, making it ideal for large-scale genetic studies. In some programs, KASP reduced total genotyping costs by up to 44% to 46% compared to other high-throughput platforms like Illumina GoldenGate.
3. Precision and Reliability
The technology offers high accuracy and reproducibility, which is critical for identifying favorable alleles in diverse genetic backgrounds.
• Robustness: KASP consistently identifies true genotypes with low error rates (typically 0.7–1.6%) and produces distinct clusters that are easier to interpret than the erratic signals often found in chip-based assays.
• Handling “Off-Target” Variation: KASP design can incorporate information on neighboring SNPs/InDels to prevent “null alleles” that might otherwise disrupt primer binding in highly heterozygous crops like potato.
4. Codominant Scoring for Early Selection
KASP is a codominant platform, meaning it can distinguish between homozygous and heterozygous individuals in a single reaction. This is vital for early-generation selection, as it allows breeders to identify plants that carry two copies of a desired gene (homozygotes) versus those carrying only one (heterozygotes) without needing to wait for the next generation.
5. Flexibility and Customization
MAB programs often need to switch between different populations where different markers might be polymorphic.
• Assay Design: KASP is highly flexible; breeders can pick specific SNPs relevant to a particular cross and design assays for them overnight.
• Broad Application: It has been successfully used across a wide range of crops—including rice, wheat, maize, potato, and tomato—to identify resistance to pests like the brown planthopper, diseases like Fusarium head blight, and abiotic stresses like heat and drought.
6. Enhanced Quality Control (QC)
Acceleration is also achieved by reducing errors within the breeding pipeline. KASP is routinely used for QC analysis, such as verifying the purity of inbred lines, confirming hybridity, and identifying mislabeled genotypes at key stages of the breeding cycle. This ensures that resources are only spent on verified genetic material.
2. How does KASP compare to other platforms for disease resistance?
Compared to other genotyping platforms used for disease resistance, Kompetitive Allele-Specific PCR (KASP) is frequently cited as a global benchmark technology because it offers a unique balance of high throughput, flexibility, and cost-effectiveness. While other platforms like SNP arrays or sequencing are superior for initial gene discovery, KASP is preferred for the routine screening of large breeding populations.
The following sections detail how KASP compares to specific alternative platforms:
1. Comparison with Traditional Gel-Based Markers (SSR, SCAR, RFLP)
• Speed and Efficiency: KASP is a homogeneous, gel-free system, which eliminates the labor-intensive and time-consuming steps of gel electrophoresis and fragment analysis required by Simple Sequence Repeat (SSR) and Sequence Characterized Amplified Region (SCAR) markers. One study reported that KASP assays are 45 times faster than gel-based PCR markers.
• Cost and Scalability: KASP is significantly more affordable for large-scale screening. For example, the cost of marker-assisted selection (MAS) with low-throughput gel assays is approximately €3 per assay, whereas SNP-based KASP assays can cost as little as €0.20 per assay when outsourced.
• Reliability: Traditional markers often suffer from low polymorphism or a lack of tight linkage to the target gene, leading to false positives. In contrast, KASP markers can be designed to be gene-specific “perfect markers,” such as the PFT_KASP marker for Fusarium head blight, which eliminates errors caused by recombination.
2. Comparison with High-Density SNP Arrays (GoldenGate, Infinium)
• Flexibility for MAS: SNP arrays are “inflexible” because they require a fixed set of thousands of markers to be run on every sample, which is not cost-effective for evaluating just a few disease-resistance genes across tens of thousands of plants. KASP allows breeders to pick and choose specific SNPs relevant to their current crosses.
• Accuracy: KASP has a lower average genotyping error rate (0.7–1.6%) compared to the Illumina GoldenGate platform (2.0–2.4%). It also handles negative controls more reliably; negative controls in KASP consistently cluster together and rarely produce false allele calls, unlike the erratic signals often seen in array-based assays.
• Cost Reduction: For applications like marker-assisted recurrent selection, KASP can be 44% to 46% cheaper than chip-based platforms.
3. Comparison with TaqMan and rhAmp
• Reagent Costs: KASP is more cost-effective than TaqMan because it uses universal fluorescent probes instead of requiring expensive, custom-labeled probes for every individual SNP site.
• Success Rates: KASP reports a higher assay design success rate (98–100%) and a higher conversion rate to working assays (93–94%) compared to TaqMan, which averages 72% and 61%, respectively.
4. Comparison with Next-Generation Sequencing (NGS) and LAMP
• Operational Focus: NGS and GBS are excellent for discovery and mapping but generate massive amounts of data that can be expensive and time-consuming to process for routine selection. KASP is designed for rapid turnaround, allowing results in as little as 14 days, which is sufficient for real-time breeding decisions.
• Field vs. Lab: While Loop-mediated isothermal amplification (LAMP) is highly sensitive and can be read by the naked eye in the field, KASP is superior for high-throughput laboratory screening of thousands of samples simultaneously.
5. Technical Advantages in Data Interpretation
• Dosage Determination: Unlike many gel-based methods, KASP is a codominant platform that can accurately distinguish between homozygous and heterozygous individuals. It even allows for dosage determination in polyploid crops like potato, enabling breeders to identify “duplex” parents that will pass resistance to a higher proportion of their progeny.
• Handling Off-Target Variation: In highly heterozygous crops, KASP design can incorporate information on neighboring SNPs/InDels to prevent “null alleles” that might otherwise disrupt primer binding and cause false-negative results.
3. Can KASP markers be used to verify hybrid purity in rice?
Yes, KASP markers can be used to verify hybrid purity and genetic identity in rice. According to the sources, KASP technology is a recognized tool for quality control (QC) analysis, which includes assessing genetic purity and verifying parentage.
• Purity Testing in Rice: KASP-based SNP analysis has been successfully implemented in rice for several applications, including genetic purity tests, diversity analysis, and quality control.
• Verification of Resistant Lines: In a study involving brown planthopper resistance in rice, KASP markers were used to validate the presence of specific stable resistance genes (Bph17 and Bph32) in original donors, which confirmed the purity of the lines maintained in the breeding program.
• Quality Control (QC) Definition: The sources define QC analysis as the process of identifying errors throughout the breeding pipeline, specifically mentioning identity control and hybrid purity as key components.
• High-Throughput Capabilities: Because KASP is fast and cost-effective, it is considered ideal for large-scale quality control and genetic identity assessments in diverse tropical crops, including rice and maize.
• Codominant Advantage: A critical feature of KASP for purity testing is that it is a codominant platform, meaning it can distinguish between homozygous and heterozygous individuals in a single reaction. This allows breeders to easily identify “off-type” plants or confirm that a hybrid carries the expected alleles from both parents.
While KASP is highly efficient for checking a small to moderate number of specific markers across many samples for purity, it is less competitive for applications requiring more than 100 SNPs per sample, where higher-density platforms might be preferred.
4. What defines a ‘perfect’ KASP marker for disease?
Based on the sources, a “perfect” KASP marker is primarily defined as one that is gene-specific and diagnostic, meaning it is derived directly from the DNA sequence of the causal gene responsible for the trait rather than just being “linked” to it.
The key attributes that define a perfect KASP marker for disease resistance include:
1. Gene-Specificity and Functional Relevance
• Direct Alignment with Causal Genes: A perfect marker is based on the underlying gene for resistance, such as the PFT_KASP marker which targets the pore-forming toxin-like gene underlying the Fhb1 resistance locus in wheat.
• Targeting Functional Mutations: These markers focus on functional SNPs or InDels that directly contribute to phenotypic variation, effectively making them “causal markers”.
• Haplotype Accuracy: It must distinguish between resistant and susceptible haplotypes even when they are very similar, such as the one-SNP difference between the R and S3 haplotypes in the Fhb1 region.
2. Elimination of Recombination Errors
• No Recombination Risk: Because the marker is within or represents the functional region of the gene, it eliminates the risk of recombination between the marker and the trait. Traditional “linked” markers can separate from the gene over multiple selection cycles, leading to breeding errors.
• Reducing False Positives: Perfect markers avoid the false positives or false negatives often produced by linked markers that rely on assumptions of tight linkage rather than functional identity.
3. High Diagnostic Reliability
• Broad Applicability: A perfect marker remains diagnostic across diverse genetic backgrounds, whereas linked markers often fail to be polymorphic or accurate when introgressed into new cultivars.
• Conclusive Call Rates: It must demonstrate high sensitivity and specificity, ensuring that the presence of the marker allele always corresponds to the presence of the resistance gene.
• Validation through Concordance: For a marker to be considered “perfect,” its genotyping results must be in complete agreement with known resistant pedigrees and phenotypes, even in cases where established linked markers disagree.
4. Technical Robustness for the KASP Platform
• Distinct Clustering: The marker must produce distinct, tight clusters on a Cartesian graph, allowing for easy manual or automated allele calling.
• Codominant Scoring: It should clearly categorize homozygous resistant, heterozygous, and homozygous susceptible lines in a single assay, which is vital for early-generation selection.
• Suitability of Flanking Regions: The genomic region surrounding the target SNP must meet the technical criteria for KASP design, avoiding excessive “off-target” variation or InDels that could disrupt primer binding and cause null alleles.
5. How does KASP perform dosage determination in polyploids?
Kompetitive Allele-Specific PCR (KASP) technology performs dosage determination in polyploid organisms by analyzing the fluorescence signal intensities generated during the PCR reaction and visualizing them as distinct clusters on a scatter plot,. While KASP is standard for bi-allelic scoring in diploids, it can be adapted to identify the number of copies of a specific allele in more complex genomes,.
The following details the process and importance of dosage determination in polyploids:
1. Clustering of Allelic Ratios
In autotetraploid crops like potato, KASP can potentially distinguish between five different genotypes based on the ratio of the two target alleles: nulliplex, simplex, duplex, triplex, and quadruplex,.
• Visual Grouping: On a Cartesian graph where the X-axis represents one allele (e.g., FAM) and the Y-axis represents the other (e.g., HEX), these genotypes form separate clusters.
• Cluster Interpretation: For instance, in an evaluation for disease resistance, the nulliplex breeding lines cluster at the bottom (low HEX signal), followed by a simplex cluster containing controls with one copy of the gene, and a third cluster identifying duplex individuals carrying two copies.
2. Software and Data Analysis
Because polyploid calls are more complex than simple diploid scoring, specialized bioinformatics tools are required to interpret the raw fluorescence intensity data.
• Specialized Packages: Researchers use software such as the R package fitTetra to perform tetraploid allele dosage calling from raw KASP intensity data.
• Increased Resolution: By including tetraploid allele dosage information, breeders can achieve a higher level of heterozygosity discrimination, allowing them to separate samples that would otherwise appear as a single “heterozygous” group in a diploid model.
3. Practical Breeding Applications
Determining the dosage of an allele is a significant advantage for marker-assisted selection (MAS).
• Parental Selection: Identifying duplex parents is highly valuable because using a parent with two copies of a resistance gene ensures that a higher proportion of its progeny will inherit the desired trait compared to using a simplex parent.
• Cross Optimization: Knowledge of dosage levels helps breeders refine their crossing blocks to maximize genetic gain for traits like resistance to late blight or potato cyst nematodes,.
4. Technical Challenges
Despite its capabilities, dosage determination in polyploids faces specific technical hurdles:
• Cluster Separation: In some autopolyploid genomes, the different heterozygote levels (simplex, duplex, triplex) can sometimes merge into a single cluster, making them extremely difficult to separate accurately.
• Genome Complexity: The presence of homeologous and paralogous sequences in polyploid crops requires extensive biological and technical validation to ensure that the KASP primers are targeting the correct locus without interference from other parts of the genome.
• ** karakterization of Donors:** characterized donors with known dosage are often used as positive controls to “anchor” the clusters and ensure accurate calls across different sample plates,.
6. What is the success rate for SNP-to-KASP conversion?
The success rate for converting Single Nucleotide Polymorphisms (SNPs) to Kompetitive Allele-Specific PCR (KASP) assays varies depending on the crop species and the complexity of its genome, but it is generally high compared to other platforms.
The “SNP conversion rate” as being composed of two distinct components:
1. Assay Design Success Rate: The ratio of SNP sites where primers can be successfully designed relative to the total number of target SNPs.
2. Work Success Rate: The number of SNP sites that actually generate reliable genotype calls once the primers are manufactured.
The specific success rates reported in the sources are as follows:
1. General and Manufacturer Benchmarks
• LGC Genomics Benchmark: The manufacturer reports an assay design success rate of 98–100% and a work success rate of 93–94%.
• Comparison to TaqMan: KASP is noted for having significantly higher success rates than TaqMan, which averages a 72% design success rate and a 61% work success rate.
2. Success Rates by Crop
• Wheat: One study reported an overall conversion rate of over 95% for functional markers. However, another study cited a lower conversion rate of 67%.
• Maize: In a genome-wide study for heat stress tolerance, researchers achieved a 100% assay design success rate and a 92% work success rate. Another study successfully converted 1,242 out of 1,536 SNPs from an Illumina GoldenGate chip, resulting in an 80.9% final working assay rate.
• Rice: Success rates appear variable; one source reports a 94.8% design success rate and 93.6% work success rate, while another recorded a 49.9% design success rate but a very high 96.6% work success rate.
• Potato: Success rates are notably lower due to the complexity of the autotetraploid genome. One study found that only 9 out of 20 (45%) KASP markers were found to work well, while the others were non-callable or not polymorphic.
• Other Crops: Conversion rates have been reported at approximately 80.6% for chickpea.
3. Factors Influencing Success and Failure
The sources identify several technical reasons why a SNP-to-KASP conversion might fail (estimated at a 29% failure rate in one maize study):
• Paralogous sequences: Similar sequences elsewhere in the genome can interfere with specific primer binding.
• Primer design issues: Incorrect design too close to the SNP site or unsuitable flanking regions.
• False SNP identification: SNPs identified in silico that are not present in the actual biological samples.
• Complex polyploidy: In crops like potato, the presence of homeologous and paralogous sequences requires extensive validation and makes it difficult to distinguish between different heterozygous clusters.
• Off-target variation: In highly heterozygous crops, high SNP frequency in the regions surrounding the target (estimated at 1 per 20 bp in potato) can disrupt primer binding.
7. How is KASP used to combat brown planthopper?
KASP technology is used to combat the brown planthopper (BPH)—one of the most destructive rice pests capable of causing up to 80% yield loss—by providing a high-throughput and precise platform for marker-assisted breeding (MAB) of resistant rice varieties,. Because many BPH resistance genes do not confer broad-spectrum resistance to different biotypes, KASP allows breeders to rapidly identify and transfer specific, effective resistance genes into popular high-yielding varieties,.
KASP is utilized in the following ways to address BPH:
1. Identification and Validation of Diagnostic Markers
KASP is used to convert single nucleotide polymorphisms (SNPs) identified through discovery methods like Genome-Wide Association Studies (GWAS) and Genotyping-by-Sequencing (GBS) into reliable diagnostic markers,.
• Efficient Selection: Researchers identified six robust KASP markers (snpOS00912, snpOS00915, snpOS00922, snpOS00923, snpOS00927, and snpOS00929) that efficiently distinguish between BPH-resistant and susceptible rice genotypes,.
• Top Performer: Among these, snpOS00922 on chromosome 6 was found to be the most efficient marker for identifying resistance.
• Gene Targeting: These KASP markers are associated with defense-related genes involving pathways such as GDSL-like lipase/acylhydrolase and pentatricopeptide repeat proteins, which help the plant detect insect effectors and activate defense mechanisms,,.
2. Pyramiding and Tracking Stable Resistance Genes
KASP assays have been successfully developed for major, stable BPH resistance genes, specifically Bph17 and Bph32, which are highly effective against the biotype 4 BPH common in the Indian subcontinent,,.
• Digenic Donors: KASP was used to identify “digenic” lines—such as MAGIC lines M201, M272, and M344—that carry favorable alleles for both Bph17 and Bph32,,.
• Gene Pyramiding: These lines serve as valuable donors for “pyramiding” (combining) multiple resistance genes into popular rice varieties to achieve long-term and broad-spectrum resistance,.
3. Quality Control (QC) and Purity Testing
KASP is used for identity control and hybrid purity assessments within BPH breeding programs,.
• Verifying Donors: In recent studies, KASP gene-specific markers for Bph17 and Bph32 were used to validate original donor lines like PTB33 and RathuHeenati, confirming the genetic purity of the materials maintained in the breeding pipeline,.
• Early Generation Selection: Because KASP is a codominant platform, it can distinguish between homozygous and heterozygous individuals in a single reaction, allowing breeders to select plants with two copies of the resistance gene early in the breeding cycle.
4. Operational Advantages
The sources note that KASP is significantly more effective than traditional gel-based markers for BPH resistance due to its speed and cost-effectiveness,. KASP-based genotyping services, such as those offered through the High-Throughput Genotyping Project (HTPG), facilitate industrial-scale screening, enabling results in just a few days to inform real-time breeding decisions,.
8. Can KASP help identify resistance to rice blast?
Yes, KASP technology can help identify resistance to rice blast, which is considered one of the most important rice diseases globally.
KASP provides a reliable and efficient way to detect genes and quantitative trait loci (QTLs) associated with blast resistance:
• Pi9 Gene Identification: A specific KASP marker, Pi9-KASP (also identified as snpOS0007), has been developed and validated to identify the Pi9 gene, which confers resistance to blast pathogens. This marker was successfully used to screen 278 rice lines to identify germplasm resources and support marker-assisted breeding.
• Pita Gene Detection: Another KASP marker (snpOS0006) has been validated for identifying the Pita gene, another significant source of blast resistance in rice.
• Pi63 Marker: KASP technology has also been applied to the Pi63 locus (associated with marker Pi63-KASP), which confers blast resistance at both the seedling and tillering stages.
• Efficiency in Breeding: Using these KASP-based molecular markers improves the efficiency and accuracy of rice breeding programs by allowing researchers to quickly screen thousands of samples for these resistance alleles without needing time-consuming phenotypic evaluations.
The sources conclude that these markers are valuable tools for the marker-assisted breeding (MAB) of rice blast-resistant lines and the identification of useful genetic resources.
9. How does KASP compare to SSR for characterization?
Kompetitive Allele-Specific PCR (KASP) technology and Simple Sequence Repeat (SSR) markers are both used for genetic characterization, but they differ significantly in terms of efficiency, technical complexity, and the depth of information they provide. While SNPs (the basis of KASP) are replacing SSRs in many modern breeding programs, SSRs still hold certain advantages for specific types of characterization.
The comparison between KASP and SSR for characterization can be broken down into the following key areas:
1. Throughput and Speed
• KASP’s Superiority: KASP is significantly faster and more amenable to high-throughput screening. One study reported that KASP assays offer 45 times superiority in speed compared to traditional gel-based PCR markers like SSRs.
• Bottlenecks of SSR: SSR markers are “gel-based,” requiring labor-intensive fragment analysis and electrophoresis steps after the PCR reaction. This makes SSRs impractical for the real-time, large-scale characterization required in modern marker-assisted selection (MAS).
2. Cost-Effectiveness
• Consumable Costs: KASP is generally more affordable for characterization, especially when processing large numbers of samples. Reagent costs for KASP can be as low as $0.05 per data point, which is significantly less than traditional systems like SSR or AFLP.
• Operational Costs: When outsourcing, the cost of applying low-throughput gel assays (like SSR) is approximately €3 per assay, whereas SNP-based KASP assays can cost as little as €0.20 per assay.
3. Accuracy and Reliability
• Error Rates: KASP provides lower genotyping error rates, typically between 0.7% and 1.6%, compared to older systems. SSR markers can be “error-prone” due to the subjective nature of scoring gel bands, which can be faint or fail to separate properly.
• Diagnostic Stability: SSR markers often suffer from a lack of polymorphism across diverse genetic backgrounds or the risk of recombination between the marker and the target gene. KASP markers can be designed as “perfect markers” derived directly from the gene sequence, ensuring they remain diagnostic regardless of the cultivar’s pedigree.
4. Resolution in Germplasm Characterization
• SSR Advantage in Diversity: Interestingly, the sources note that for germplasm characterization (such as identifying population structure or measuring genetic distance), SSRs can provide more information than SNPs on a per-marker basis. Because SSRs are highly polymorphic, they are often better at clustering germplasm into distinct populations.
• SNP Requirement: It is estimated that 7 to 11 times more SNPs are required to achieve the same resolution as SSRs for analyzing population structure and genetic diversity, particularly in highly heterogeneous or outcrossing species.
5. Technical Requirements and Scoring
• Gel-Free vs. Gel-Based: KASP is a homogeneous, closed-tube, and gel-free system. This eliminates post-PCR handling and the need for hazardous materials like ethidium bromide.
• Codominant Scoring: Both KASP and SSRs are codominant, meaning they can distinguish between homozygous and heterozygous individuals. However, KASP’s fluorescence-based clustering (FAM vs. HEX signals) allows for more automated and objective “allele calling” than visual gel inspection.
Summary Comparison Table
| Feature | KASP (SNP-based) | SSR (Fragment-based) |
|---|---|---|
| Speed | High (45x faster) | Low (Labor-intensive) |
| Throughput | High-throughput | Low-throughput |
| Detection | Gel-free/Fluorescence | Gel-based |
| Cost | Low (~€0.20/assay) | High (~€3.00/assay) |
| Precision | High (Gene-specific) | Variable (Linked) |
| Diversity Info | Requires many markers | Highly informative per locus |
10. How can KASP verify if a rice hybrid is genetically pure?
Kompetitive Allele-Specific PCR (KASP) technology verifies the genetic purity of rice hybrids by performing quality control (QC) analysis, which identifies errors or “off-types” throughout the breeding pipeline. It is considered a global benchmark for this purpose due to its high precision, speed, and cost-effectiveness.
KASP verifies hybrid purity through the following mechanisms:
1. Codominant Scoring
A primary advantage of KASP is that it is a codominant platform, meaning it can distinguish between homozygous and heterozygous individuals in a single reaction.
• Verifying Hybridity: To confirm a successful hybrid, KASP ensures the plant is heterozygous at specific loci, carrying one unique allele from the female parent and one from the male parent.
• Identifying Off-Types: If a plant expected to be a hybrid is found to be homozygous for a parental allele, it indicates a failure in hybridization (e.g., self-pollination) or seed contamination.
2. Genetic Identity and Fingerprinting
KASP is used for identity control, ensuring that a specific rice line or hybrid matches its expected genetic profile.
• SNP Subsets: For routine purity testing, breeders often use a specific subset of 50–100 SNPs that are uniformly distributed across the genome. This “fingerprint” is sufficient to measure non-homogeneity and confirm genetic identity.
• Parentage Verification: By genotyping the hybrid along with its parents, KASP confirms that the hybrid contains “true-to-type” alleles from its specific lineage. Any hybrid showing more than 10% non-parental alleles is typically discarded.
3. Verification of Donor Lines
In rice breeding programs, KASP is used to maintain the purity of the source material (donor lines).
• Example in BPH Resistance: In studies for brown planthopper (BPH) resistance, KASP gene-specific markers confirmed the purity of original donor lines (such as PTB33 and RathuHeenati) by validating the presence of favorable alleles for resistance genes like Bph17 and Bph32. This ensures that the genetic foundation of the hybrid program remains unpolluted.
4. Operational Advantages for Purity Testing
The technical features of KASP make it ideal for the large-scale screening required in purity verification:
• Speed: KASP is significantly faster than traditional gel-based markers (like SSRs), with some studies noting it is 45 times superior in speed.
• Rapid Turnaround: Genotyping results can be obtained in as little as 14 days, allowing breeders to make decisions about seed purity before the next planting season.
• Cost-Effectiveness: For screening a moderate number of markers (like the 50-100 needed for QC) across thousands of samples, KASP is more affordable than higher-density arrays.
11. What makes a disease-resistance marker ‘perfect’ in breeding?
A “perfect” disease-resistance marker is defined as a gene-specific, diagnostic marker that is derived directly from the DNA sequence of the causal gene responsible for the trait, rather than being merely “linked” to it.
The following characteristics define a perfect marker in a breeding context:
1. Functional Relevance (Causal Markers)
A perfect marker is typically a Functional Marker (FM), meaning it targets mutations—such as Single Nucleotide Polymorphisms (SNPs) or Insertions/Deletions (InDels)—that directly contribute to phenotypic variation.
• Example: The PFT_KASP marker is considered “perfect” because it is based on the pore-forming toxin-like (PFT) gene, which is the actual underlying gene for Fhb1 resistance in wheat.
• Direct Discrimination: These markers accurately discriminate allelic variations among various breeding lines by focusing on the functional region of the gene.
2. Elimination of Recombination Risk
The primary drawback of traditional “linked” markers (such as Simple Sequence Repeats or SSRs) is that recombination can occur between the marker and the underlying gene after multiple selection cycles.
• Permanent Linkage: Because a perfect marker is part of the gene itself, there is zero risk of the marker “separating” from the resistance trait during crossing.
• Reliability: This ensures that if the marker is detected, the resistance gene is guaranteed to be present.
3. High Diagnostic Accuracy (No False Positives)
Linked markers often suffer from “false positives,” where cultivars exhibit the desirable marker genotype but do not actually possess the resistance gene.
• Conclusive Calling: A perfect marker provides complete concordance between the genotype and the known phenotype across diverse germplasm.
• Sensitivity and Specificity: It demonstrates high sensitivity (true positive rate) and specificity (true negative rate), correctly identifying resistant and susceptible lines even when they share very similar haplotypes.
4. Robustness Across Diverse Genetic Backgrounds
Linked markers frequently fail to be polymorphic (show differences) when introgressed into new, diverse genetic backgrounds.
• Universal Utility: Because perfect markers target the causal mutation, they remain diagnostic regardless of the pedigree or region.
• Haplotype Accuracy: They can distinguish between resistant and susceptible haplotypes that might appear identical to less precise markers.
5. Technical Suitability for High-Throughput Platforms
In the context of modern breeding technologies like KASP, a perfect marker must also be technically robust:
• Clear Discrimination: It must produce distinct, tight clusters on a Cartesian graph, allowing for unambiguous allele calling of homozygous resistant, heterozygous, and homozygous susceptible lines.
• Codominant Scoring: It should be a codominant marker, capable of identifying heterozygotes in early-breeding generations, which is critical for efficient selection.
• Cost-Effectiveness: It must be amenable to high-throughput, gel-free platforms to enable real-time decision-making during a single growing season.
12. How does KASP handle dosage in polyploid crops like potato?
In autotetraploid crops like potato (2n = 4x = 48), Kompetitive Allele-Specific PCR (KASP) handles allele dosage by analyzing raw fluorescence intensity data to distinguish between the five possible genotypic states: nulliplex, simplex, duplex, triplex, and quadruplex. While standard KASP is used for bi-allelic scoring in diploids, its application in polyploids requires specialized bioinformatics and visualization techniques to interpret the varying ratios of the two target alleles.
The following are the mechanisms and challenges regarding KASP dosage determination:
1. Cluster-Based Visualization
Dosage levels are identified by plotting the fluorescence signals for the reference allele (X-axis) against the alternate allele (Y-axis) on a Cartesian graph.
• Distinct Groupings: Genotypes with different dosages typically form separate, distinct clusters. For example, in disease resistance screening, the nulliplex lines (zero copies of the resistant allele) cluster at the bottom of the graph, while the simplex lines (one copy) form a group above them, followed by a duplex cluster (two copies).
• Increased Resolution: By including tetraploid allele dosage information, breeders can separate samples that would otherwise appear as a single “heterozygous” group in a diploid model, allowing for more precise genetic characterization.
2. Software for Dosage Calling
Because polyploid calls are more complex than simple bi-allelic scoring, researchers use specialized bioinformatics tools to assign scores. Specifically, the R package fitTetra is used to perform tetraploid allele dosage calling from raw KASP intensity data. This software helps automate the classification of samples into the five tetraploid groups based on their signal intensities.
3. Practical Breeding Applications
Determining dosage is a significant advantage for marker-assisted selection (MAS) in potato:
• Parental Selection: Identifying duplex parents (those with two copies of a resistance gene) is highly valuable because they pass the desired trait to a significantly higher proportion of their progeny than simplex parents.
• Operational Efficiency: Dosage information allows breeders to avoid using susceptible or simplex parents in crossing blocks, thereby maximizing genetic gain.
4. Technical Challenges
Despite its utility, handling dosage in polyploids faces specific hurdles:
• Genome Complexity: The presence of homeologous and paralogous sequences in polyploids requires extensive biological and technical validation to ensure KASP primers are targeting the correct locus.
• Cluster Merging: In some cases, high levels of heterozygosity or genomic complexity can cause different heterozygote clusters to merge into a single group, making it extremely difficult to distinguish between simplex, duplex, and triplex individuals.
• Off-Target Variation: Potato has a high SNP frequency (estimated at 1 per 20 bp), and “off-target” variation near the target SNP can disrupt primer binding, potentially leading to null alleles or inaccurate dosage calls.
13. What are the advantages of using KASP for potato starch content?
Using Kompetitive Allele-Specific PCR (KASP) to analyze potato starch content offers several significant advantages over traditional phenotypic methods, primarily by targeting specific natural variations in the StCWIN1 gene (cell wall invertase) that regulate carbon partitioning,.
The specific advantages include:
1. Superior Efficiency and Speed
Phenotypically measuring starch content (SC) and dry matter (DM) is time-consuming and labor-intensive. KASP provides a high-throughput screening alternative that allows breeders to evaluate large numbers of accessions rapidly, which has historically been a major limiting factor in potato genetic improvement,.
2. Precise Identification of Favorable Alleles
The technology allows breeders to distinguish between specific allelic variations of the StCWIN1 gene, such as the SNP00075 (G/T) locus.
• Performance Differences: Research demonstrated that accessions possessing Allele-T had significantly higher starch content (17.1%) and dry matter (22.9%) compared to those with Allele-G (13.6% SC and 19.5% DM),.
• Haplotype Analysis: KASP markers can identify elite haplotypes, such as Haplotype 113, which is significantly associated with higher dry matter and was likely selected during potato domestication,.
3. Accelerated Marker-Assisted Breeding (MAB)
KASP markers for StCWIN1 act as a “reliable molecular marker toolkit” for the genetic improvement of tuber traits.
• Predictive Power: Because the marker is closely associated with the favorable traits, it allows for the molecular selection of new varieties with improved tuber weight and yield before they reach maturity,.
• Improving Carbon Partitioning: The use of these markers helps identify genotypes that are more efficient at transporting sucrose from source to sink tissues, which is a crucial determinant of final yield,.
4. Technical Robustness and Flexibility
KASP is noted for being a flexible and accurate SNP genotyping method suitable for the complex potato genome.
• High Conformity: The markers developed for starch content show good cluster separation in scatter plots, making it easy to categorize accessions into distinct homozygous or heterozygous groups,.
• Cost-Effectiveness: Compared to other genotyping platforms, KASP is highly cost-effective for screening targeted loci across large populations,.
5. Insight into Plant Homeostasis
Beyond simple yield, the cell wall invertase genes targeted by KASP play a role in regulating the plant’s response to environmental factors, such as pathogen attack and abiotic stress (drought and heat), ensuring a more stable life cycle and yield realization.
14. How is KASP used to identify heat stress resilience in maize?
In maize breeding, Kompetitive Allele-Specific PCR (KASP) technology is used to identify and select for heat stress resilience by converting single nucleotide polymorphisms (SNPs) found in heat stress response (HSR) genes into functional molecular markers.
The identification process typically follows these steps:
1. Comparative Transcriptomics and SNP Discovery
Researchers use RNA-seq (transcriptomics) to compare maize inbred lines with contrasting responses to heat, such as LM11 (extremely heat-sensitive) and CML25 (heat-tolerant).
• Vulnerable Tissues: Sequence libraries are created from tissues most susceptible to heat damage, specifically the top leaf, ovule, and pollen.
• Bioinformatics Pipeline: Accurate SNPs are identified between the sensitive and tolerant lines using mappers like STAR and callers like GATK.
• HSR Gene Targeting: Markers are prioritized based on their location within or near Heat Stress Response (HSR) genes and Differentially Expressed Genes (DEGs). These genes are often associated with pathways like photosynthesis (Rubisco), antioxidation (APX), and heat shock proteins (HSP70, DNAJ).
2. Development of Maize KASPar Assay Markers (MKAMs)
Selected SNPs are converted into high-throughput KASP markers, often designated as MKAMs.
• Genome-Wide Coverage: In one study, 100 genome-wide SNPs (10 per chromosome) were chosen to cover the entire maize genome for genetic dissection.
• Conversion Success: These markers show high design success rates (100% in one study) and reliable “work” success rates (92%), making them more efficient than older markers like SSRs.
3. Validation and Population Genotyping
The developed KASP markers are validated using parental inbred lines and then applied to genotype F2 mapping populations.
• Discrimination of Alleles: KASP provides clear cluster plots that distinguish between alleles from the heat-sensitive parent, the heat-tolerant parent, and heterozygous individuals.
• Polymorphic Utility: In validation tests, a majority of these markers (approximately 71%) were found to be polymorphic and usable for tracking resilience traits.
4. Application in Marker-Assisted Selection (MAS)
The ultimate use of these KASP markers is to accelerate the breeding of heat-resilient maize through marker-assisted selection.
• Speed and Cost: KASP is highly cost-effective and provides a rapid turnaround time, allowing breeders to screen thousands of samples for resilience markers in just a few days.
• Trait Tracking: These markers allow breeders to bypass expensive and labor-intensive phenotypic evaluations—such as measuring tassel blast or pollen sterility in the field—by instead screening for the genetic signatures of tolerance.
15. Can KASP distinguish different polyploid allele dosages in potatoes?
Yes, KASP technology can distinguish different polyploid allele dosages in potatoes. Because cultivated potato is an autotetraploid (2n = 4x = 48), it possesses four copies of each chromosome, allowing for five possible genotypic states based on the ratio of target alleles.
KASP manages this complexity through the following methods:
1. Discrimination of Five Genotypic Groups
KASP can identify the specific “dosage” or number of copies of a particular allele in a potato genotype. These states are classified as:
• Nulliplex: Zero copies of the target allele.
• Simplex: One copy.
• Duplex: Two copies.
• Triplex: Three copies.
• Quadruplex: Four copies.
2. Analysis of Fluorescence Intensity
The determination of dosage is achieved by analyzing the raw fluorescence intensity data generated during the PCR reaction. On a Cartesian scatter plot (where the X and Y axes represent the two different fluorescent dyes, FAM and HEX), these different dosage levels typically form distinct, separate clusters.
• For example, a cluster at the bottom of a plot (low signal for the alternate allele) identifies nulliplex lines, while subsequent clusters further up the plot identify simplex and duplex individuals.
3. Specialized Data Processing
Because polyploid calls are more complex than standard diploid scoring, researchers often use specialized bioinformatics tools. The sources specifically mention using the R package fitTetra to perform tetraploid allele dosage calling from raw KASP intensity data.
4. Practical Breeding Value
The ability to distinguish dosage levels is highly beneficial for potato breeding programs:
• Identifying Duplex Parents: Breeders specifically look for “duplex” parents (carrying two copies of a resistance gene) because using them in crosses ensures a higher proportion of progeny will inherit the desired trait compared to using simplex parents.
• Enhanced Fingerprinting: Including tetraploid dosage information increases the levels of detectable heterozygosity, allowing for more precise genetic discrimination between closely related clones, such as full-sibs or half-sibs, which might otherwise appear identical at the diploid level.
5. Technical Challenges
While effective, the sources note that dosage determination in polyploids can be challenging. In some instances, different heterozygote levels may merge into a single cluster, making it difficult to accurately separate simplex, duplex, and triplex individuals. Additionally, the high frequency of SNPs in the potato genome (estimated at 1 per 20 bp) requires extensive validation to ensure that “off-target” variation does not interfere with the primer binding and skew the dosage calls.
16. How does KASP detect viruses like Wheat Streak Mosaic Virus?
KASP (Kompetitive Allele-Specific PCR) technology does not typically detect the virus itself; rather, it is used to detect DNA markers (SNPs) associated with virus-resistance genes within the host plant’s genome, such as the Wsm2gene in wheat,.
By identifying and tracking these genetic markers, breeders can use marker-assisted selection (MAS) to rapidly develop cultivars that are naturally resistant to viruses like Wheat Streak Mosaic Virus (WSMV),.
Mechanism of Detection for WSMV Resistance
The process of using KASP to “detect” resistance to WSMV involves the following steps:
• Identifying Genetic Loci: Researchers first identify Single Nucleotide Polymorphisms (SNPs) that are tightly linked to resistance genes. For WSMV, the focus is often on the Wsm2 gene located on chromosome 3BS,.
• SNP-to-KASP Conversion: These linked SNPs (often identified via high-density arrays or sequencing) are converted into breeder-friendly KASP assays,.
• The Reaction Components: A specific KASP assay is designed using three unlabelled primers: two allele-specific forward primers (one for the resistant allele and one for the susceptible allele) and one common reverse primer,.
• Allele Discrimination via Fluorescence: During the PCR reaction, the allele-specific primers compete to bind to the target DNA. If the plant has the resistant allele, a specific fluorescent signal (e.g., FAM) is emitted; if it has the susceptible allele, a different signal (e.g., HEX) is emitted,.
• Clustering and Analysis: The results are plotted on a Cartesian graph where samples group into distinct clusters: homozygous resistant, homozygous susceptible, or heterozygous,.
Key Advantages for Virus Resistance Breeding
• High Throughput and Speed: KASP is an ideal platform for screening thousands of plants for a small number of markers in just a few days, allowing for real-time breeding decisions,.
• Cost-Effectiveness: It is significantly more affordable than traditional gel-based markers or high-density SNP arrays when screening large populations for specific traits,.
• Codominant Scoring: KASP can distinguish between homozygous and heterozygous individuals in a single reaction, which is vital for identifying plants that will pass on the resistance to their offspring,.
• Stability Across Environments: Unlike phenotypic screening, which requires infecting plants and observing symptoms (a process that can be inconsistent due to environmental factors), KASP provides a reliable genetic profile regardless of the growing conditions,.
Versatility Beyond Host Resistance
While the WSMV example focuses on host resistance, the sources also note that KASP technology is versatile enough for direct pathogen identification. For instance, a KASP assay was developed to identify the rice white tip nematode (Aphelenchoides besseyi) by targeting specific SNPs in its ribosomal DNA, allowing researchers to distinguish it from other closely related nematode species,. Similarly, KASP assays have been developed for resistance to other viruses, such as Maize Streak Virus (MSV) and Rice Stripe Virus (RSV),.
17. Can KASP distinguish different disease haplotypes in wheat?
Yes, KASP technology can distinguish between different disease haplotypes in wheat. The sources demonstrate its ability to accurately identify and separate both resistant and susceptible haplotypes for various diseases, even when genetic differences are extremely small.
Specifically, there are following applications:
1. Fusarium Head Blight (Fhb1) Haplotypes
• Discrimination of Similar Haplotypes: KASP can distinguish between the resistant (R) haplotype and four different susceptible haplotypes (S1, S2, S3, and S4).
• Precision Targeting: A specific KASP marker, PFT_KASP, was designed to target a single SNP (G/A polymorphism) that differentiates the resistant haplotype from the S3 susceptible haplotype.
• Clustering Accuracy: The assay accurately groups genotypes into three distinct clusters: the R haplotype, the S3 haplotype, and a third cluster representing all other susceptible haplotypes (S1, S2, and S4) that lack the targeted gene entirely.
• Resolving Errors: KASP is more sensitive than traditional linked markers like UMN10, which often fail to distinguish between the R and S3 haplotypes, leading to false-positive resistance calls.
2. Wheat Streak Mosaic Virus (Wsm2) Haplotypes
• Multiple Haplotype Identification: Research using three closely linked KASP markers identified six distinct haplotypes associated with Wsm2.
• Resistance vs. Susceptibility: The platform successfully categorized these into three haplotypes associated with resistance (R1, R2, and R3) and three associated with susceptibility (S1, S2, and S3).
• Effectiveness in Diverse Backgrounds: These KASP-based haplotypes were validated across multiple advanced breeding populations, including RIL (recombinant inbred line) and DH (doubled haploid) populations.
3. Pre-Harvest Sprouting (PHS) Haplotypes
• Haplotype-Specific Selection: A major QTN on chromosome 3D was used to develop the QSS.TAF9-3D KASP marker, which distinguishes between two primary haplotypes: TT and CC.
• Predictive Value: The TT haplotype was found to be highly correlated with PHS resistance, particularly in white-grained wheat varieties, allowing breeders to identify and select resistant materials that might have been overlooked by other markers.
4. Leaf Rust (Lr21) Haplotypes
• Indel-Based Haplotypes: KASP assays were developed to target specific insertion/deletion (InDel) polymorphisms within the Lr21 gene.
• Sorting Discrepancies: In cases where earlier SNP-based markers gave ambiguous results in soft winter wheat, a second KASP assay was able to correctly categorize genotypes by identifying a unique deletion present only in resistant germplasm.
Summary of Haplotype Diagnostic Capabilities
| Disease Trait | Target Haplotypes | KASP Diagnostic Advantage |
|---|---|---|
| Fusarium Head Blight | R, S1, S2, S3, S4 | Can distinguish R from S3 (2-SNP difference). |
| Wheat Streak Mosaic Virus | R1-R3, S1-S3 | Identifies resistant vs. susceptible profiles across populations. |
| Pre-Harvest Sprouting | TT (Resistant), CC (Susceptible) | Effectively identifies resistance in white-grained varieties. |
| Leaf Rust | Resistant vs. Susceptible InDels | Corrects errors caused by ambiguous SNP signals in specific backgrounds. |
18. How is KASP used to identify submergence tolerance in rice?
In rice breeding, Kompetitive Allele-Specific PCR (KASP) technology is used to identify submergence tolerance by targeting the SUBMERGENCE 1 (SUB1) locus on chromosome 9. This locus is the primary factor conferring tolerance to complete inundation, which is a major constraint for rice production in rainfed lowlands.
The following details on how KASP is applied to this trait:
1. Targeting Natural Allelic Variants
The Sub1A gene at the SUB1 locus has two main natural variants that determine the plant’s response to flooding:
• Sub1A-1: The tolerant allele type found in varieties like FR13A.
• Sub1A-2: The susceptible allele type; natural variants of this type fail to show phenotypic tolerance even if the gene is present.
2. Development of Specific KASP Markers
KASP markers were developed to replace traditional gel-based markers (like AEX1 and GnS2) by targeting the specific Single Nucleotide Polymorphisms (SNPs) that distinguish these two alleles. Two primary KASP markers are used:
• Sub1A_SNP1 (T/C): Targets the SNP at the 556th position.
• Sub1A_SNP2 (A/G): Targets the SNP at the 678th position.
3. Validation and Accuracy
These KASP markers have been validated using segregating F2 populations (such as a cross between the susceptible variety Komboka and the tolerant IR49830). The results from KASP assays have shown 100% consistency with traditional gel-based marker analysis, proving their reliability for marker-assisted breeding (MAB).
4. Operational Advantages Over Traditional Methods
The sources identify several reasons why KASP is the preferred method for identifying submergence tolerance:
• Co-dominant Scoring: Unlike the traditional dominant marker AEX1, which only identifies the presence of the tolerant allele, the Sub1A_SNP1 KASP marker can distinguish between homozygous tolerant, homozygous susceptible, and heterozygous individuals in a single reaction.
• Elimination of Enzyme Digestion: The Sub1A_SNP2 KASP marker is more efficient than the traditional GnS2 marker because it eliminates the need for a time-consuming additional enzyme reaction step.
• High Throughput and Cost: KASP enables the screening of a large number of seeds quickly and at a lower cost than gel-based methods, which are often labor-intensive and expensive.
• Accurate Clustering: In KASP assays, tolerant and susceptible varieties form distinct, easily identifiable clusters on scatter plots based on their fluorescence signals, allowing for rapid automated genotype calling.
19. How do breeders use KASP to pyramid multiple resistance genes?
Breeders use Kompetitive Allele-Specific PCR (KASP) technology to pyramid (or “stack”) multiple resistance genes by utilizing its high-throughput, cost-effective, and highly precise SNP genotyping capabilities. Pyramiding is a strategy to combine several genes into a single variety to achieve broad-spectrum and more durable resistance that is harder for pathogens or pests to overcome.
The process of using KASP for pyramiding involves several key stages:
1. Identifying and Validating Donor Lines
Breeders use KASP to screen germplasm collections and “MAGIC” (multi-parent advanced generation inter-cross) lines to find donor genotypes that already possess one or more desired resistance genes.
• Gene-Specific Accuracy: Using “perfect” or gene-specific KASP markers—such as the PFT_KASP marker for Fhb1 in wheat—ensures that the breeder is selecting the actual functional mutation rather than a marker that might recombine and separate from the resistance trait.
• Example in Rice: Breeders used KASP to identify “digenic” MAGIC lines (e.g., M201, M272, and M344) that carry favorable alleles for both Bph17 and Bph32 resistance genes against brown planthopper.
2. High-Throughput Selection in Segregating Generations
Once crosses are made, breeders must screen thousands of plants in segregating generations (like F2 or BC populations) to find individuals that have inherited all the targeted genes.
• Operational Speed: KASP is significantly faster than traditional gel-based methods, allowing results in a few days so breeders can make selection decisions before the next crossing or planting cycle.
• Low Cost: Pyramiding often requires testing a few SNPs across tens of thousands of samples; KASP is more cost-effective for this specific “low-density, high-volume” application compared to high-density SNP arrays.
3. Codominant Scoring for Efficient Breeding
Because KASP is a codominant platform, it can distinguish between homozygous and heterozygous individuals in a single assay.
• Homozygote Identification: For a pyramid to be stable, the resulting variety needs to be homozygous for all the favorable alleles. KASP allows breeders to identify these “true-to-type” multigenic plants early in the breeding cycle, saving time and field space.
• Example in Wheat: KASP has been used to verify wheat germplasm containing pyramids for Fusarium head blight resistance, specifically stacking Fhb1 and Fhb5 (or Qfhb-5AS) alleles derived from the cultivar Sumai-3.
4. Dosage Determination in Polyploids
In polyploid crops like potato, KASP provides allele dosage information (e.g., nulliplex, simplex, duplex), which is vital for pyramiding.
• Targeted Parental Selection: Breeders can use KASP to identify duplex parents (carrying two copies of a resistance gene), ensuring that a higher proportion of their offspring will inherit the desired pyramid.
• Stacking in Potato: KASP markers are used to stack resistance genes for different biotic stresses, such as late blight (e.g., R2 and R8-QTL) and viruses like Potato Virus Y (e.g., Ryadg and Rysto) into elite breeding backgrounds.
5. Quality Control and Purity Verification
To ensure the pyramid remains intact during maintenance and multiplication, KASP is used for quality control (QC).
• Identity Control: Breeders use KASP “fingerprints” (50–100 SNPs) to verify that a multigenic line has not been mislabeled or contaminated with self-pollinated seeds during the breeding process.
• Hybrid Purity: It confirms that hybrid progeny have inherited the correct set of alleles from both parents to maintain the desired combination of genes.
20. How is KASP used to identify submergence tolerance in rice?
In rice breeding, Kompetitive Allele-Specific PCR (KASP) technology is used to identify submergence tolerance by targeting the SUBMERGENCE 1 (SUB1) locus on chromosome 9. This locus is the primary factor conferring tolerance to complete inundation, which is a major production constraint in the rainfed lowlands of Asia and Africa.
The application of KASP for this trait involves the following specific methods and advantages:
1. Targeting Natural Allelic Variants
The Sub1A gene at the SUB1 locus has two main natural variants that determine the plant’s response to flooding:
• Sub1A-1: The tolerant allele found in varieties like FR13A and IR49830.
• Sub1A-2: The susceptible allele; natural variants of this type fail to show phenotypic tolerance even if the gene is present.
2. Development of Specific KASP Markers
Two primary KASP markers were developed to replace traditional gel-based markers by targeting the specific Single Nucleotide Polymorphisms (SNPs) that distinguish the tolerant and susceptible alleles:
• Sub1A_SNP1 (T/C): Targets the SNP at the 556th position of the gene sequence.
• Sub1A_SNP2 (A/G): Targets the SNP at the 678th position.
3. Operational Advantages Over Traditional Methods
KASP offers several critical improvements over the older gel-based markers AEX1 and GnS2:
• Co-dominant Scoring: Unlike the traditional dominant marker AEX1, which only identifies the presence of the tolerant allele, the Sub1A_SNP1 KASP marker can distinguish between homozygous tolerant, homozygous susceptible, and heterozygous individuals in a single reaction.
• Elimination of Enzyme Digestion: The Sub1A_SNP2 KASP marker is more efficient than the traditional GnS2 (a CAPS marker) because it eliminates the need for a time-consuming additional restriction enzyme reaction step.
• High Throughput and Low Cost: KASP enables the screening of a large number of seeds quickly and at a lower cost than gel-based methods, which are labor-intensive and expensive.
4. Validation and Reliability
The KASP markers have been validated using segregating F2 populations, such as a cross between the susceptible variety Komboka and the tolerant IR49830. The results from KASP assays have shown 100% consistency with traditional gel-based marker analysis, proving their high accuracy for marker-assisted breeding (MAB). In these assays, tolerant and susceptible varieties form distinct, easily identifiable clusters on scatter plots based on their fluorescence signals, allowing for rapid automated genotype calling.
21. How does KASP support marker-assisted selection for drought tolerance?
Kompetitive Allele-Specific PCR (KASP) supports marker-assisted selection (MAS) for drought tolerance by enabling high-throughput, cost-effective screening of complex polygenic traits across large breeding populations. Drought tolerance is often controlled by many minor-effect quantitative trait loci (QTLs), making traditional phenotypic selection difficult and expensive.
KASP technology facilitates drought tolerance breeding through the following specific applications:
1. Marker-Assisted Recurrent Selection (MARS)
KASP is an ideal platform for MARS, a strategy used to accumulate favorable alleles for highly polygenic traits like drought tolerance.
• Maize Projects: KASP has been central to large-scale initiatives such as the Drought-Tolerant Maize for Africa (DTMA) and Water-Efficient Maize for Africa (WEMA) projects.
• Operational Efficiency: The technology allows breeders to genotype any number of samples with polymorphic SNPs without designing specific multiplex sets, providing the rapid turnaround time (as little as 14 days) necessary for pre-flowering selection and rapid cycle recombination.
2. Validation of Specific Drought-Resilience Genes in Wheat
In wheat breeding, KASP markers have been developed and validated for specific functional genes that underpin drought tolerance:
• TaDreb-B1 and 1-feh w3: These specific KASP markers are used to identify genotypes with superior alleles for drought tolerance.
• Enzyme Activity: The 1-FEH w3 variant is associated with enzyme activity for remobilizing stem water-soluble carbohydrates (WSC) to the grain under drought stress.
• Marker Toolkit: These assays provide a robust molecular toolkit for tracking favorable yield-related alleles under water-limited conditions.
3. High-Throughput Screening in Rice and Sorghum
• Rice: Specific KASP markers (e.g., HZ_Os03273 and HZ_Os03275) have been developed to target drought tolerance QTLs such as qDT3.9 and qDT6.3. These are used for pyramiding drought tolerance with other traits like low nitrogen tolerance.
• Sorghum: KASP technology has been used for the introgression of drought tolerance QTLs into farmers’ preferred varieties.
4. Implementation in Maize Breeding Pipelines
CIMMYT has successfully used KASP panels to identify drought-tolerant varieties globally:
• In one study, a set of 96 KASP markers (derived from 39,846 initial SNPs) was used to genotype 450 F2 families.
• This screening identified 57 KASP markers that were highly effective in assessing drought tolerance in these specific populations.
5. Technical and Economic Advantages
• Low Cost: For applications requiring a small to moderate number of markers across many samples—typical of drought MAS—KASP can be up to 46% cheaper than chip-based platforms.
• Flexibility: Breeders can select only the polymorphic SNPs relevant to a specific population, which is crucial for drought traits where genetic significance varies between crosses.
• Sample Quality: KASP works effectively with DNA bulks (pooling 10–15 plants per sample), which reduces costs and time when screening large germplasm collections for favorable drought alleles.
22. How do KASP markers assist in breeding for wheat quality?
KASP (Kompetitive Allele-Specific PCR) markers assist in breeding for wheat quality by enabling high-throughput, cost-effective selection of genes that control end-use quality traits, which are often difficult, expensive, or destructive to measure phenotypically.
KASP markers assist in the following specific areas of wheat quality breeding:
1. Gluten and Bread-Making Quality
Gluten quality is a primary determinant of bread-making characteristics and is optimized by selecting for specific high-molecular-weight glutenin subunits (HMW-GS).
• Targeted Loci: KASP assays have been developed for the Glu-A1, Glu-B1, and Glu-D1 loci.
• Predicting Performance: These assays can distinguish between subunits that significantly affect mixing properties. For example, assays at the Glu-D1 locus differentiate between the Dx2+Dy12 and Dx5+Dy10 subunits, which showed average mixing time (MTxW) values of 5.8 mm and 11.4 mm, respectively.
• Selection Ranking: Markers at the Glu-A1 locus allow breeders to rank allelic effects on mixing time as AxNull < Ax1 < Ax2*.
2. Grain Hardness
Grain hardness is essential for determining the milling and baking utility of wheat.
• Gene Selection: KASP markers for the Pina-D1, Pinb-D1, and Pinb-B2 genes have been validated using Single Kernel Characterization System (SKCS) values.
• Haplotype Identification: These markers allow breeders to categorize and rank haplotypes based on hardness levels (e.g., ranking Pina-D1a + Pinb-D1a as the softest and Pina-D1b + Pinb-D1b as the hardest).
3. Flour Color and Appearance
The sources identify several enzymes and pigments that determine flour color and the aesthetic quality of wheat products:
• Yellow Pigment Content (YPC): KASP assays target the Phytoene Synthase (PSY) and Zeta-Carotene Desaturase (ZDS) genes, which are essential components of the pathways encoding flour color.
• Enzymatic Discoloration: KASP markers for polyphenol oxidase (PPO) activity (Ppo-A1 and Ppo-D1) and peroxidase (POD) activity (TaPod-A1) identify alleles associated with lower activity levels, which is desirable for preventing the darkening of noodles and other products.
• Black Point Resistance: Black point causes undesirable discoloration at the embryonic end of grains, lowering their processing and nutrient quality. A KASP marker was developed for a candidate gene (TraesCS3B02G404700) to facilitate selection for black point resistance.
4. Starch Quality
KASP markers assist in selecting for starch granule size distribution, which affects the quality of steamed bread and noodles.
• Waxy Genes: A dominant KASP assay for the Wx-B1 gene identifies the “null-type” allele, which results in significantly fewer A/B-type starch granules compared to the wild-type allele.
5. Pre-Harvest Sprouting (PHS) Resistance
PHS is a major constraint that leads to a significant reduction in grain and end-use quality due to increased enzyme activity.
• Multi-Gene Toolkits: KASP markers for genes such as TaPHS1, TaSdr-B1, TaVp-1B, and TaMFT-A1 allow breeders to combine multiple beneficial alleles to improve sprouting tolerance.
• White-Grained Wheat: While red wheat is naturally more resistant, KASP markers like QSS.TAF9-3D and those on chromosome 3AL have been identified as valuable tools for maintaining quality in popular white-grained varieties.
6. Operational Advantages for Quality Breeding
• Non-Destructive and Fast: Traditional quality testing often requires milling and seed destruction. KASP allows for genetic profiling using only a small amount of leaf tissue, saving seeds for the next generation.
• Speed: KASP assays are reported to be 45 times faster than traditional gel-based PCR markers, allowing for real-time selection decisions.
• Efficiency in Segregating Populations: KASP is more cost-effective than high-density arrays for screening large numbers of plants for a few specific quality-related SNPs.
23. How does KASP compare to TaqMan for SNP genotyping?
Based on the sources, Kompetitive Allele-Specific PCR (KASP) and TaqMan are both recognized as advanced, uniplex, single-tube SNP genotyping platforms that utilize terminal fluorescence reading for high-throughput analysis. While they share similar principles, they differ significantly in their chemistry, cost-effectiveness, and conversion success rates.
1. Reagent Cost and Probe Requirements
A major difference between the two platforms lies in their probe systems:
• TaqMan: This technology requires the synthesis of a site-specific probe for every individual SNP being tested.
• KASP: Instead of site-specific probes, KASP uses universal FRET cassettes (probes) that can be used across a wide variety of different gene-specific primers. Because researchers do not need to synthesize a new probe for each specific site, KASP greatly reduces the reagent cost of experiments compared to TaqMan.
2. Success and Conversion Rates
KASP generally demonstrates higher technical reliability in moving from SNP identification to a functional assay:
• Assay Design Success Rate: KASP achieves a design success rate of 98–100%, compared to 72% for TaqMan.
• Working Assay Rate: KASP successfully converts into working assays at a rate of 93–94%, whereas TaqMan averages 61%.
3. High-Throughput Capability and Error Rates
Both platforms are considered benchmarks for high-throughput genotyping because they reduce the time and labor required for large-scale screening.
• Performance: Both systems are noted for lowering error rates compared to older gel-based markers.
• Amenability to Automation: KASP is specifically highlighted for its flexibility and ease of automation, making it a “benchmark technology” for industrial-scale genotyping.
4. Application Suitability
Both TaqMan and KASP are ideal for applications that requires genotyping a small to moderate number of SNPs across a large number of samples. This makes them preferred choices for:
• Quality Control (QC): Verifying genetic identity and hybrid purity.
• Marker-Assisted Selection (MAS): Screening breeding populations for specific disease or stress-resistance traits where only a few loci are relevant.
• Pyramiding: Tracking the inheritance of multiple specific genes in segregating generations.
While KASP and TaqMan are the primary competitors in this space, the sources note that other platforms like rhAmp have recently joined them as highly competitive technologies for these purposes.
24. How is KASP used to identify submergence tolerance in rice?
In rice breeding, Kompetitive Allele-Specific PCR (KASP) is used to identify submergence tolerance by targeting the SUBMERGENCE 1 (SUB1) locus located on chromosome 9. This locus is the primary genetic factor conferring tolerance to complete inundation, a major production constraint in the rainfed lowlands of Asia and Africa.
The application of KASP for identifying this trait involves the following key components:
1. Targeting Natural Allelic Variants
The Sub1A gene at this locus has two main natural variants that determine how a rice plant responds to flooding:
• Sub1A-1: The tolerant allele found in varieties such as FR13A and IR49830.
• Sub1A-2: The susceptible allele; natural variants of this type do not exhibit phenotypic tolerance even when the gene is present.
2. Specific KASP Marker Design
Researchers developed KASP markers to replace older, less efficient gel-based markers (AEX1 and GnS2) by targeting the specific Single Nucleotide Polymorphisms (SNPs) that distinguish these two alleles. There are two primary KASP markers used:
• Sub1A_SNP1 (T/C): Targets the SNP at the 556th position of the gene sequence.
• Sub1A_SNP2 (A/G): Targets the SNP at the 678th position.
3. Operational Advantages Over Traditional Methods
KASP provides several critical improvements for identifying submergence tolerance compared to the older gel-based markers:
• Co-dominant Scoring: Unlike the traditional dominant marker AEX1—which can only detect the presence of the tolerant allele—the Sub1A_SNP1 KASP marker can distinguish between homozygous tolerant, homozygous susceptible, and heterozygous individuals in a single reaction.
• Elimination of Enzyme Digestion: The Sub1A_SNP2 marker is more efficient than the traditional GnS2 (a CAPS marker) because it eliminates the need for a time-consuming restriction enzyme reaction step.
• High-Throughput and Efficiency: KASP allows for the screening of a large number of seeds quickly and at a lower cost, which is essential for large-scale marker-assisted breeding (MAB).
4. Validation and Reliability
These KASP markers have been validated using segregating F2 populations (such as a cross between the susceptible variety Komboka and the tolerant IR49830). Results from KASP assays have shown 100% consistency with traditional gel-based marker analysis, proving their high accuracy and reliability for selecting submergence-tolerant rice lines.
25. How does KASP support marker-assisted selection for drought tolerance?
Kompetitive Allele-Specific PCR (KASP) technology supports marker-assisted selection (MAS) for drought tolerance by providing a high-throughput, flexible, and cost-effective platform to track the complex genetic components of this trait across various crops. Drought tolerance is typically a polygenic trait controlled by many minor-effect loci, making traditional phenotypic evaluation expensive and difficult.
KASP facilitates drought-tolerance breeding through the following mechanisms:
1. Marker-Assisted Recurrent Selection (MARS)
KASP is an ideal platform for MARS, a strategy used to accumulate favorable alleles for highly polygenic traits.
• Maize Projects: KASP has been central to major initiatives such as the Drought-Tolerant Maize for Africa (DTMA) and Water-Efficient Maize for Africa (WEMA) projects.
• Operational Speed: The platform allows for the rapid turnaround time necessary for pre-flowering selection, which enables rapid cycle recombination in the same growing season.
• Flexibility: Breeders can genotype any number of samples with specific polymorphic SNPs without the need to design fixed multiplex sets for every unique population.
2. Targeted Selection of Drought-Resilience Genes
Researchers have developed KASP markers for specific functional genes known to underpin drought tolerance:
• Wheat: KASP assays for TaDreb-B1 and 1-feh w3 are used to identify genotypes with superior drought-resilient alleles. The 1-FEH w3 variant is specifically associated with enzyme activity that remobilizes stem water-soluble carbohydrates to the grain when the plant is under drought stress.
• Rice: Specific KASP markers (e.g., HZ_Os03273 and HZ_Os03275) target quantitative trait loci (QTLs) like qDT3.9 and qDT6.3 to facilitate the simultaneous improvement of drought and low nitrogen tolerance through designed QTL pyramiding.
• Sorghum: KASP technology has been successfully applied to the introgression of drought tolerance QTLs into farmers’ preferred varieties.
3. Efficiency in Large Breeding Pipelines
• Industrial Scale: KASP facilitates shared, industrial-scale genotyping for various breeding organizations, allowing them to screen thousands of samples for drought-related markers in just a few days.
• Vietnam Case Study: In one application, a set of 96 KASP markers was used to genotype 450 F2 families, identifying 57 markers that were highly effective for assessing drought tolerance in those specific maize populations.
4. Technical and Cost Advantages
• Reduced Costs: For applications requiring a small to moderate number of markers across many samples—typical of drought MAS—KASP can be between 7.9% and 46.1% cheaper than chip-based platforms like GoldenGate.
• DNA Bulking: KASP is robust enough to work with DNA bulks (pooling 10–15 plants per sample), which significantly reduces the cost and time required to screen large germplasm collections for favorable drought alleles.
References
Devran, Z., & Göknur, A. (2020). Development and validation of a SNP-based KASP assay for rapid identification of Aphelenchoides besseyi Christie, 1942. Crop Protection, 136. https://doi.org/10.1016/j.cropro.2020.105235
Devran, Z., Göknur, A., & Mesci, L. (2016). Development of molecular markers for the Mi-1 gene in tomato using the KASP genotyping assay. Horticulture Environment and Biotechnology, 57(2), 156–160. https://doi.org/10.1007/s13580-016-0028-6
Dhakal, S., Tan, C. T., Anderson, V., Yu, H., Fuentealba, M. P., Rudd, J. C., Haley, S. D., Xue, Q., Ibrahim, A. M. H., Garza, L., Devkota, R. N., & Liu, S. (2018). Mapping and KASP marker development for wheat curl mite resistance in “TAM 112” wheat using linkage and association analysis. Molecular Breeding, 38(10). https://doi.org/10.1007/s11032-018-0879-x
Dipta, B., Sood, S., Mangal, V., Bhardwaj, V., Thakur, A. K., Kumar, V., & Singh, B. (2024). KASP: a high-throughput genotyping system and its applications in major crop plants for biotic and abiotic stress tolerance. In Molecular Biology Reports (Vol. 51, Number 1). Springer Science and Business Media B.V. https://doi.org/10.1007/s11033-024-09455-z
Gong, L., Guo, X., Nie, F., Fang, M., Zhang, L., Liu, X., Yang, W., Shi, L., Li, W., Zhang, G., Guo, Z., Wang, A., & Zhang, H. (2023). Identification of single nucleotide polymorphism in StCWIN1 and development of Kompetitive Allele-Specific PCR (KASP) marker associated with tuber traits in potato. Plant Growth Regulation, 101(2), 453–462. https://doi.org/10.1007/s10725-023-01032-x
Ishwarya Lakshmi, V. G., Sreedhar, M., JhansiLakshmi, V., Gireesh, C., Rathod, S., Bohar, R., Deshpande, S., Laavanya, R., Kiranmayee, K. N. S. U., Siddi, S., & Vanisri, S. (2022). Development and Validation of Diagnostic KASP Markers for Brown Planthopper Resistance in Rice. Frontiers in Genetics, 13. https://doi.org/10.3389/fgene.2022.914131
Jagtap, A. B., Vikal, Y., & Johal, G. S. (2020). Genome-wide development and validation of cost-effective KASP marker assays for genetic dissection of heat stress tolerance in maize. International Journal of Molecular Sciences, 21(19), 1–21. https://doi.org/10.3390/ijms21197386
Kante, M., Lindqvist-Kreuze, H., Portal, L., David, M., & Gastelo, M. (2021). Kompetitive allele specific pcr (Kasp) markers for potato: An effective tool for increased genetic gains. Agronomy, 11(11). https://doi.org/10.3390/agronomy11112315
Kou, C., Peng, C. J., Dong, H. Bin, Hu, L., & Xu, W. G. (2023). Mapping quantitative trait loci and developing their KASP markers for pre-harvest sprouting resistance of Henan wheat varieties in China. Frontiers in Plant Science, 14. https://doi.org/10.3389/fpls.2023.1118777
Liu, G., Mullan, D., Zhang, A., Liu, H., Liu, D., & Yan, G. (2023). Identification of KASP markers and putative genes for pre-harvest sprouting resistance in common wheat (Triticum aestivum L.). Crop Journal, 11(2), 549–557. https://doi.org/10.1016/j.cj.2022.09.002
Meade, F., Byrne, S., Griffin, D., Kennedy, C., Mesiti, F., & Milbourne, D. (2020). Rapid Development of KASP Markers for Disease Resistance Genes Using Pooled Whole-Genome Resequencing. Potato Research, 63(1), 57–73. https://doi.org/10.1007/s11540-019-09428-x
Moon, J. H., Son, D., Lee, J. W., & Yoo, S. C. (2019). Development of Kompetitive Allele Specific PCR markers for submergence tolerant gene Sub1 in rice. Plant Breeding and Biotechnology, 7(1), 62–66. https://doi.org/10.9787/PBB.2019.7.1.62
Neelam, K., Brown-Guedira, G., & Huang, L. (2013). Development and validation of a breeder-friendly KASPar marker for wheat leaf rust resistance locus Lr21. Molecular Breeding, 31(1), 233–237. https://doi.org/10.1007/s11032-012-9773-0
Pandurangan, S., Nilsen, K. T., & Kumar, S. (2021). Validation of a snp-kasp marker for the fusarium head blight resistance quantitative trait loci on chromosome 5as. Canadian Journal of Plant Science, 101(1), 135–139. https://doi.org/10.1139/cjps-2020-0099
Rasheed, A., Wen, W., Gao, F., Zhai, S., Jin, H., Liu, J., Guo, Q., Zhang, Y., Dreisigacker, S., Xia, X., & He, Z. (2016). Development and validation of KASP assays for genes underpinning key economic traits in bread wheat. Theoretical and Applied Genetics, 129(10), 1843–1860. https://doi.org/10.1007/s00122-016-2743-x
Semagn, K., Babu, R., Hearne, S., & Olsen, M. (2014). Single nucleotide polymorphism genotyping using Kompetitive Allele Specific PCR (KASP): Overview of the technology and its application in crop improvement. In Molecular Breeding (Vol. 33, Number 1, pp. 1–14). Kluwer Academic Publishers. https://doi.org/10.1007/s11032-013-9917-x
Singh, L., Anderson, J. A., Chen, J., Gill, B. S., Tiwari, V. K., & Rawat, N. (2019). Development and validation of a perfect KASP marker for fusarium head blight resistance gene Fhb1 in wheat. Plant Pathology Journal, 35(3), 200–207. https://doi.org/10.5423/PPJ.OA.01.2019.0018
Smith, S. M., & Maughan, P. J. (2015). SNP genotyping using KASPar assays. Methods in Molecular Biology, 1245, 243–256. https://doi.org/10.1007/978-1-4939-1966-6_18
Sood, S., Bhardwaj, V., Chourasia, K. N., Kaur, R. P., Kumar, V., Kumar, R., Sundaresha, S., Bohar, R., Garcia-Oliveira, A. L., Singh, R. K., & Kumar, M. (2022). KASP markers validation for late blight, PCN and PVY resistance in a large germplasm collection of tetraploid potato (Solanum tuberosum L.). Scientia Horticulturae, 295. https://doi.org/10.1016/j.scienta.2021.110859
Tan, C. T., Assanga, S., Zhang, G., Rudd, J. C., Haley, S. D., Xue, Q., Ibrahim, A., Bai, G., Zhang, X., Byrne, P., Fuentealba, M. P., & Liu, S. (2017). Development and validation of kasp markers for wheat streak mosaic virus resistance gene wsm2. Crop Science, 57(1), 340–349. https://doi.org/10.2135/cropsci2016.04.0234
Tan, C. T., Yu, H., Yang, Y., Xu, X., Chen, M., Rudd, J. C., Xue, Q., Ibrahim, A. M. H., Garza, L., Wang, S., Sorrells, M. E., & Liu, S. (2017). Development and validation of KASP markers for the greenbug resistance gene Gb7 and the Hessian fly resistance gene H32 in wheat. Theoretical and Applied Genetics, 130(9), 1867–1884. https://doi.org/10.1007/s00122-017-2930-4






Pingback: Molecular Markers in Genetic Analysis and Marker-Assisted Selection: A Comprehensive Professional Resource - Aneknowledge.com