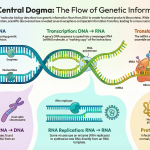
Introduction: Setting the Stage for Protein Synthesis
Every living cell is a bustling metropolis powered by tiny molecular machines called proteins. Proteins are the workhorses of the cell, carrying out a vast array of tasks, from building cellular structures to catalyzing biochemical reactions. To build these essential molecules, the cell relies on a process called translation.
Think of an mRNA molecule as a detailed blueprint containing the specific instructions for building one particular protein. The ribosome, a complex molecular machine, acts as the factory that reads this blueprint and assembles the protein piece by piece.
The Cast of Characters: Meet the Key Players
Before we walk through the process, let’s meet the cast of molecular players involved in translation. Each has a specific and vital role in getting the protein factory up and running.
• The mRNA Blueprint: This is the instruction manual for building a specific protein. It has a special tag at its beginning called the 5′ cap, which acts as a signal for the protein-making machinery to know where to start.
• The Ribosome (The Machine): This is the factory itself, and it comes in two pieces: a small 40S subunit and a large 60S subunit. These two parts must join together to form the complete, functional machine.
• The Initiator tRNA (The First Building Block): This special molecule is responsible for carrying the very first amino acid, Methionine. Its job is to recognize the official “start” signal on the mRNA blueprint.
Eukaryotic translation is a cornerstone of gene expression, representing the final, decisive step in the flow of genetic information from DNA to functional protein. This process, the synthesis of proteins from messenger RNA (mRNA) templates, is fundamental to cellular homeostasis, growth, and the ability of organisms to respond to a vast array of internal and external stimuli. The translational machinery is a dynamic and exquisitely controlled system, ensuring that the right proteins are produced in the correct amounts, at the proper time. This review provides a comprehensive overview of the four primary phases of this process—initiation, elongation, termination, and recycling. We will place a particular focus on the intricate web of regulatory mechanisms that govern this fundamental biological pathway, from the selection of mRNAs for translation to the quality control surveillance systems that maintain proteome integrity. Finally, we will incorporate insights gained from modern high-throughput techniques, such as ribosome profiling, which have revolutionized our understanding of the dynamics of protein synthesis on a genome-wide scale.
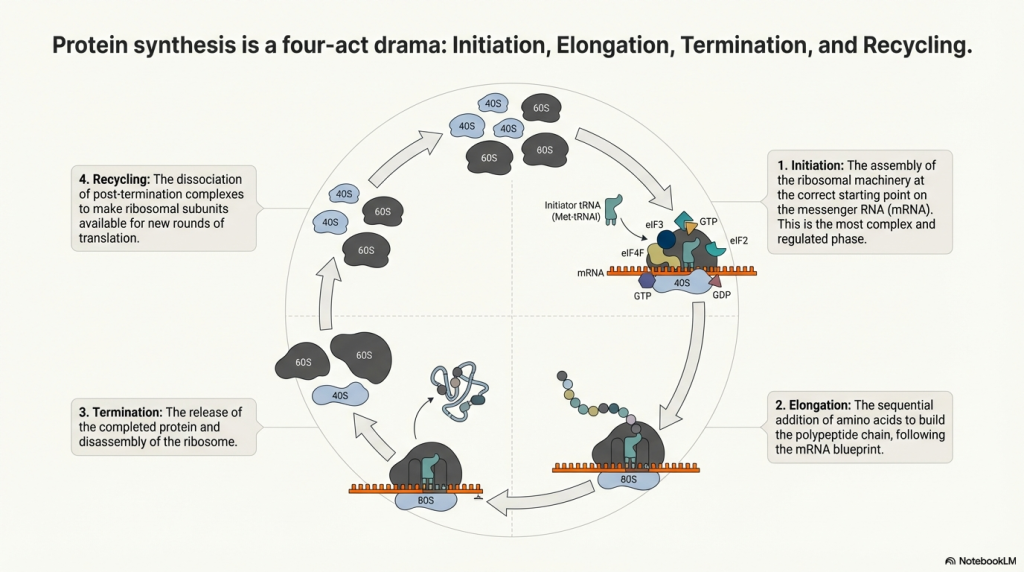
1. The Core Translational Machinery
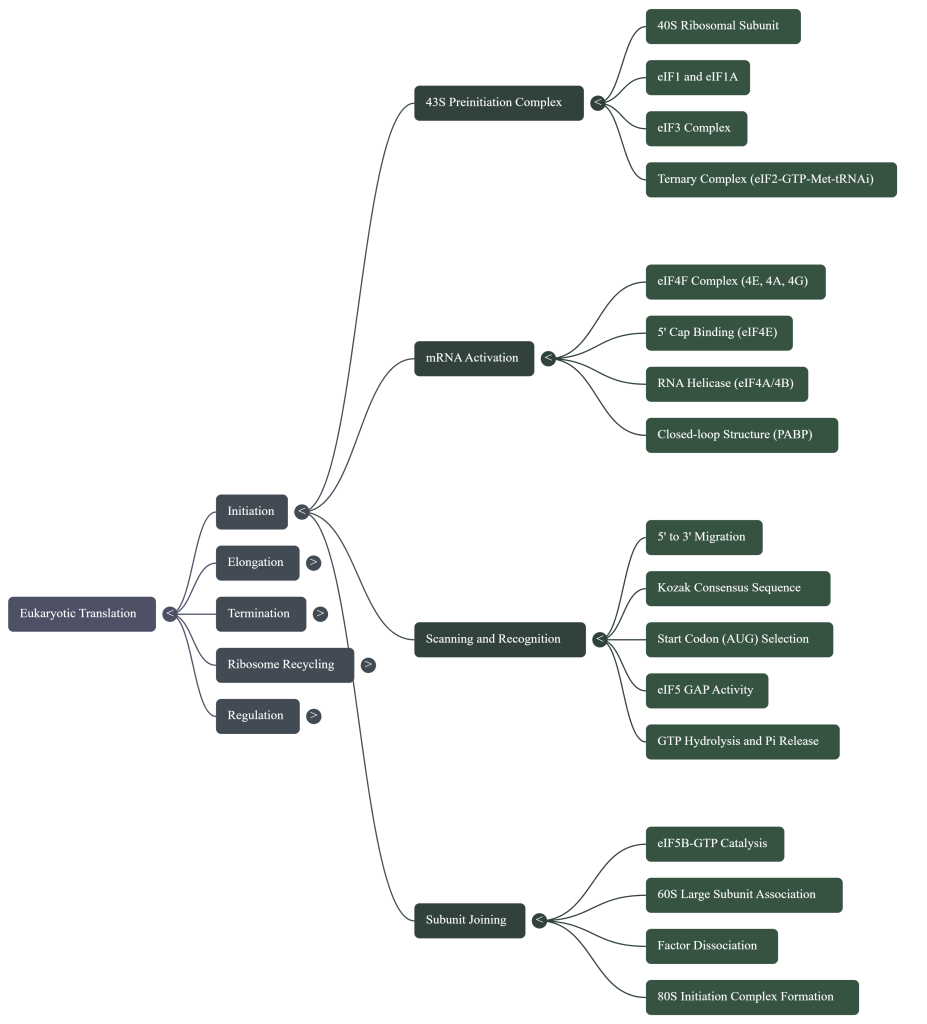
To understand the regulation of protein synthesis, it is essential first to appreciate the core components of the machinery that carry it out. At the heart of the process is the eukaryotic 80S ribosome, a massive ribonucleoprotein complex that serves as the central catalytic machine. The ribosome is composed of two subunits: the small (40S) subunit, which is primarily responsible for decoding the mRNA, and the large (60S) subunit, which catalyzes the formation of peptide bonds. These subunits work in concert with transfer RNAs (tRNAs), which act as adaptors, matching specific mRNA codons to their corresponding amino acids. Guiding the ribosome through the distinct phases of protein synthesis are three major classes of auxiliary protein factors: eukaryotic initiation factors (eIFs), elongation factors (eEFs), and release factors (eRFs).
The fully assembled 80S ribosome contains three distinct tRNA binding sites that are crucial for its function:
• A-site (Aminoacyl): The entry point for incoming aminoacyl-tRNAs.
• P-site (Peptidyl): Holds the tRNA carrying the growing polypeptide chain.
• E-site (Exit): Accommodates the deacylated tRNA before it is released from the ribosome.
The coordinated progression of tRNAs through these sites is orchestrated by the translation factors, as summarized in the table below.
| Factor Class | General Function | Examples |
| Initiation Factors (eIFs) | Mediate the assembly of the 80S ribosome at the correct start codon of an mRNA. This is the most complex phase, involving numerous factors. | eIF1, eIF2, eIF3, eIF4F, eIF5 |
| Elongation Factors (eEFs) | Facilitate the cyclical process of polypeptide chain extension, including the delivery of new aminoacyl-tRNAs and translocation of the ribosome. | eEF1A, eEF2 |
| Release Factors (eRFs) | Recognize stop codons in the A-site, triggering the hydrolysis and release of the completed polypeptide chain. | eRF1, eRF3 |
These components form a highly dynamic and interactive system. The following sections will deconstruct how these players collaborate, beginning with the first and most complex phase, translation initiation.
2. Phase I: Translation Initiation – The Primary Regulatory Hub
Translation initiation is the most intricate and heavily regulated phase of protein synthesis, serving as the primary control point that dictates cellular efficiency and defines the proteome. This stage determines not only which mRNAs are translated from the cellular pool but also where translation begins. The selection of a specific start codon defines the open reading frame (ORF), and by extension, the precise amino acid sequence of the resulting protein. The cell leverages this complexity to enact sweeping changes to the proteome in response to stress, developmental cues, and disease states. This section will deconstruct the canonical scanning mechanism of initiation, from the assembly of preinitiation complexes to start codon recognition, and will also explore key alternative pathways that provide additional layers of regulatory control.
2.1. Formation of the 43S Preinitiation Complex (PIC)
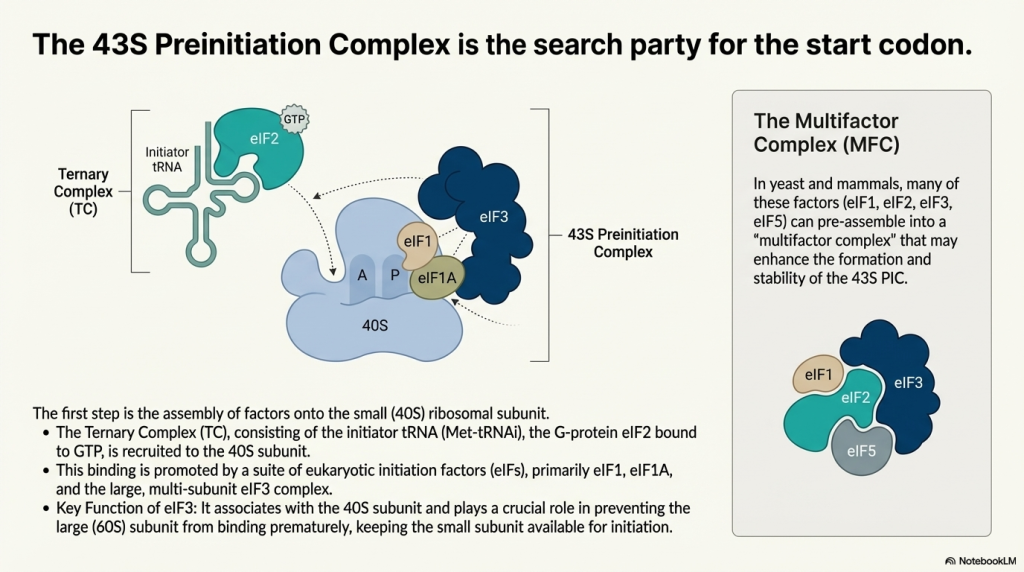
The initiation pathway begins with the assembly of the 43S preinitiation complex (PIC). This process is initiated by the formation of the ternary complex (TC), which consists of the GTP-bound initiation factor eIF2 bound to the initiator methionyl-tRNA (Met-tRNAi). This TC, along with factors eIF1, eIF1A, eIF3, and eIF5, binds to the free 40S ribosomal subunit. The collective action of these factors primes the 40S subunit, inducing a conformational state that is competent for mRNA recruitment and scanning, while also preventing premature association with the 60S subunit.
2.2. mRNA Recruitment and Activation
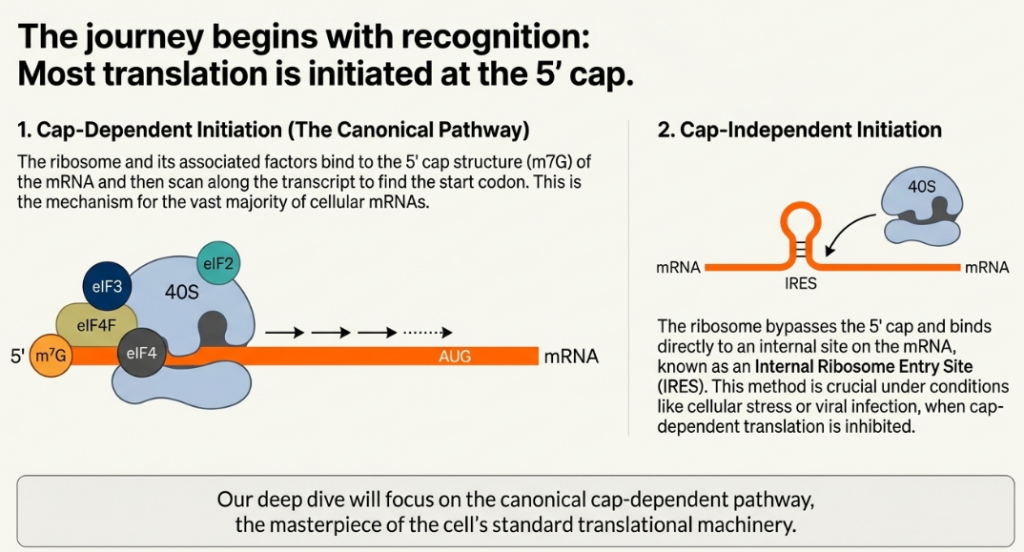
Once formed, the 43S PIC must be recruited to an mRNA template. In eukaryotes, this occurs predominantly through two distinct mechanisms: a cap-dependent pathway used by the vast majority of cellular mRNAs and a cap-independent pathway employed by certain viral and cellular transcripts.
2.2.1. Cap-Dependent Initiation
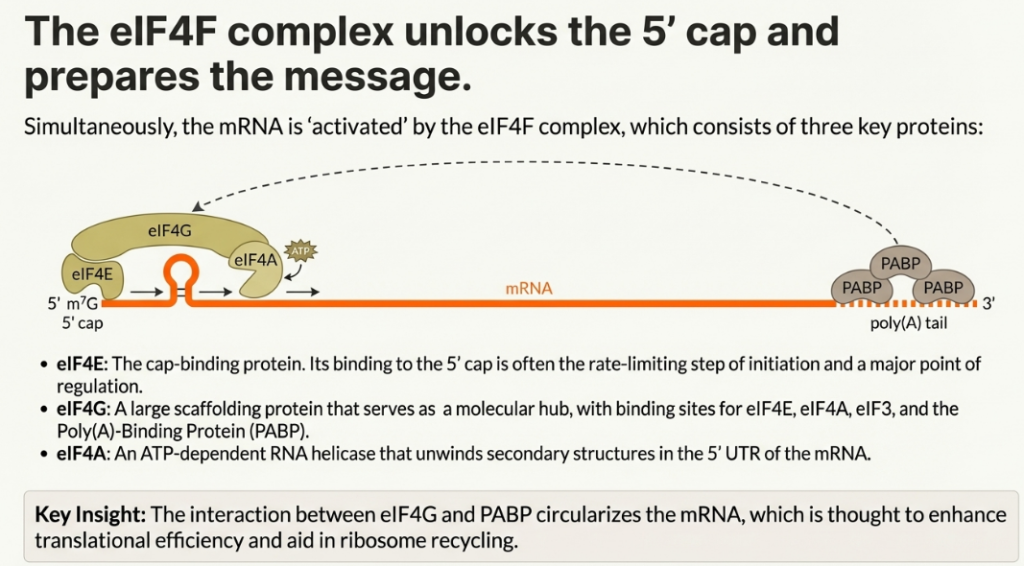
The canonical pathway for mRNA recruitment relies on the recognition of the 7-methylguanosine (m7G) cap structure found at the 5′ end of eukaryotic mRNAs. This recognition is mediated by the eIF4F complex, which is composed of three key proteins:
• eIF4E: The cap-binding protein that directly recognizes and binds the m7G cap.
• eIF4A: An ATP-dependent RNA helicase that unwinds secondary structures in the 5′ untranslated region (5′ UTR) of the mRNA.
• eIF4G: A large scaffold protein that orchestrates the initiation process by interacting simultaneously with eIF4E, eIF4A, and eIF3.
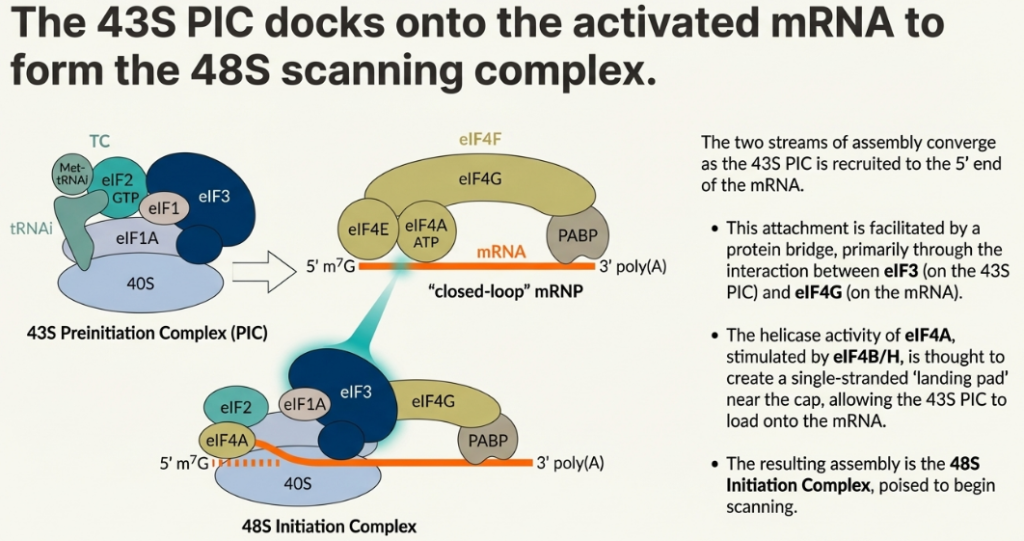
The helicase activity of eIF4A, which is further stimulated by the accessory factor eIF4B, resolves RNA hairpins and other structures near the 5′ end, creating a single-stranded “landing pad” for the 43S PIC. The recruitment is solidified through a chain of interactions: the cap is bound by eIF4E, which is bound by eIF4G, which in turn binds eIF3, a core component of the 43S PIC. This linkage effectively tethers the 40S subunit to the 5′ end of the mRNA, positioning it to begin scanning.
2.2.2. Cap-Independent Initiation via IRESs
Some viral and cellular mRNAs have evolved a mechanism to bypass the requirement for the 5′ cap and the cap-binding factor eIF4E. These transcripts contain highly structured RNA elements known as Internal Ribosome Entry Sites (IRESs). An IRES can directly recruit the 43S PIC to the mRNA, often near the authentic start codon, without the need for 5′ end recognition. This cap-independent mechanism provides a critical advantage under conditions where cap-dependent initiation is globally inhibited, such as during viral infection or cellular stress, allowing for the selective translation of IRES-containing mRNAs.
2.3. Scanning and Start Codon Recognition
Following its recruitment to the mRNA, the 43S PIC begins a kinetic proofreading process, scanning along the 5′ UTR in a 5′ to 3′ direction to locate a suitable start codon. The fidelity of this search is paramount. The 43S PIC is maintained in a scanning-competent “open” conformation (POUT) by the factors eIF1 and eIF1A. eIF1, in particular, antagonizes the conformational changes required for stable codon-anticodon pairing, thereby enabling the complex to discriminate against near-cognate codons (e.g., CUG) or authentic AUG codons situated in a poor sequence context.
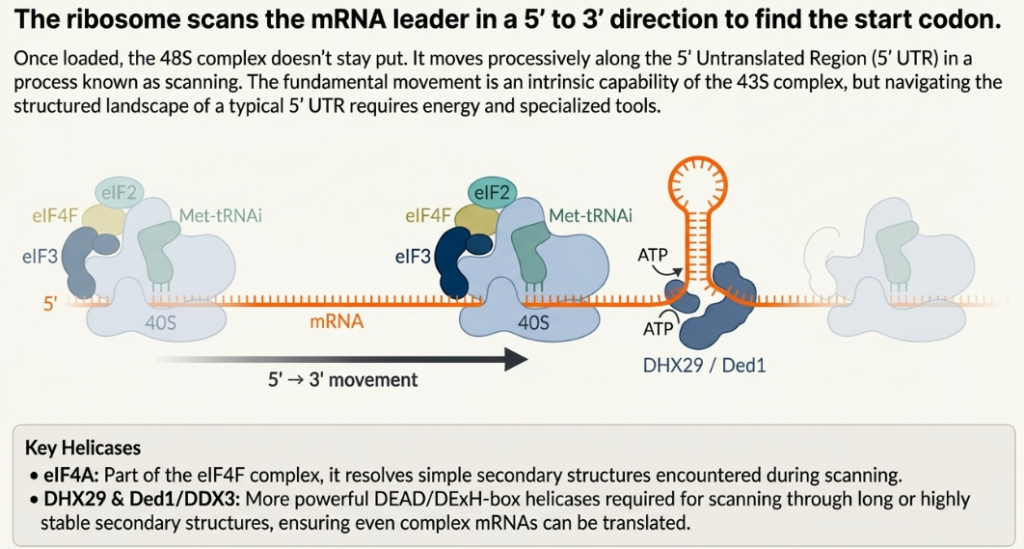
Efficient initiation typically occurs at the first AUG codon encountered, especially when that codon is embedded within an optimal Kozak consensus sequence (GCC(A/G)CCAUGG). The presence of a purine (A or G) at the -3 position and a G at the +4 position (relative to the A of the AUG at +1) strongly promotes recognition.
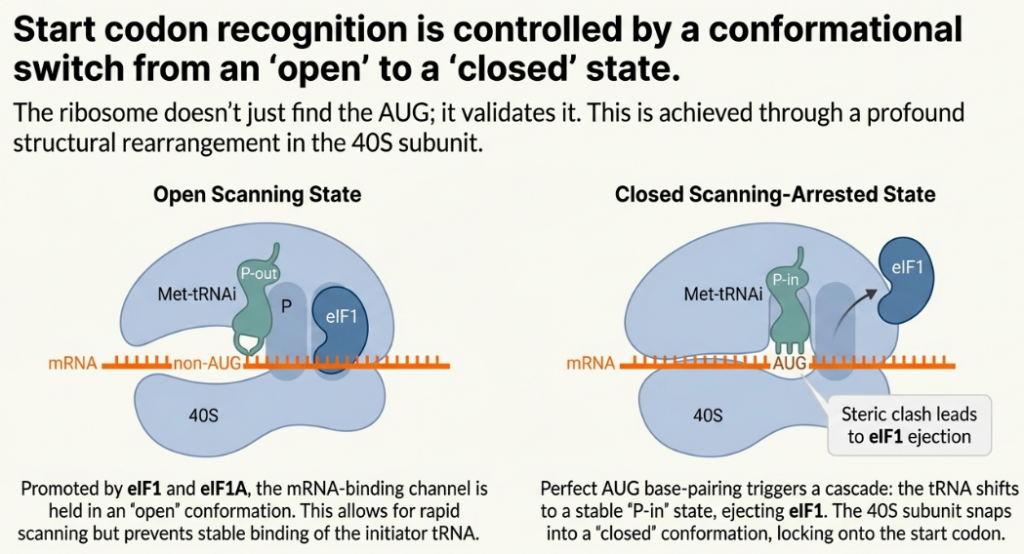
Upon successful and stable base-pairing between an AUG start codon and the anticodon of the Met-tRNAi, the 43S PIC undergoes a major conformational rearrangement. This switch from the “open” state to a scanning-arrested, initiation-committed “closed” state (PIN) involves the displacement of eIF1. This irreversible transition locks the complex onto the start codon and signals the successful completion of the search phase.
2.4. 80S Ribosome Assembly
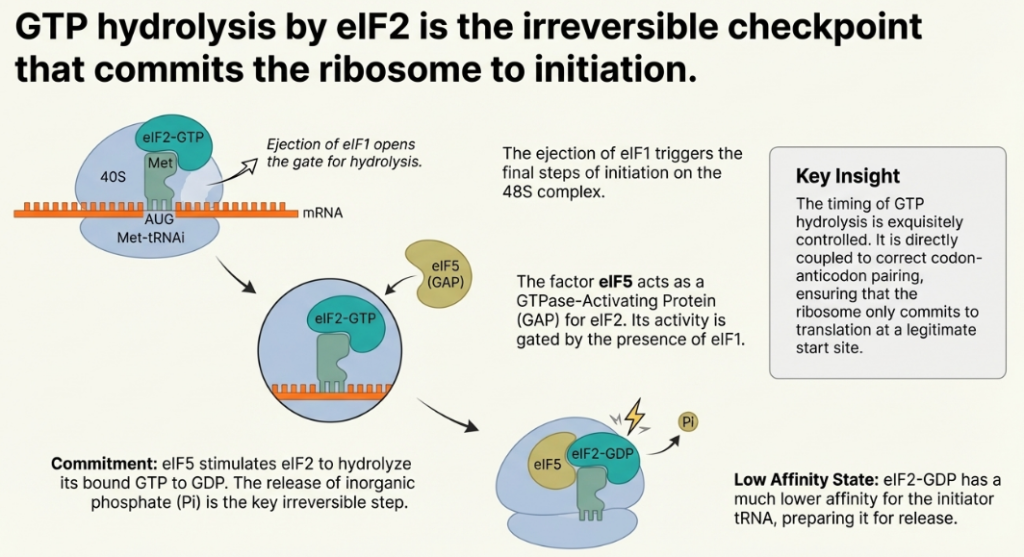
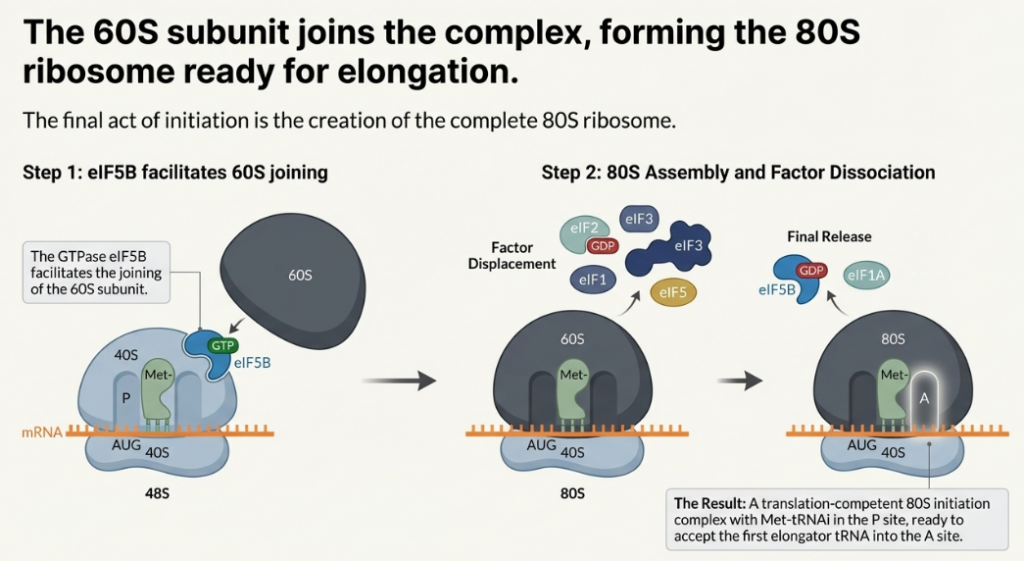
Start codon recognition is the trigger for the final steps of initiation. The factor eIF5, acting as a GTPase-activating protein (GAP), promotes the hydrolysis of the GTP bound to eIF2. This event leads to the release of eIF2-GDP and other initiation factors from the 40S subunit. Subsequently, the large 60S ribosomal subunit is recruited in a process mediated by another GTPase, eIF5B. The joining of the 60S subunit forms the complete, elongation-competent 80S ribosome, with the Met-tRNAi securely positioned in the P-site, ready to accept the first elongator tRNA. With the 80S ribosome fully assembled at the start of the coding sequence, the processive phase of polypeptide synthesis can begin.
3. Phase II: Elongation – Synthesizing the Polypeptide Chain
Once the 80S ribosome is properly assembled at the start codon, the elongation phase commences. This phase is not merely a passive assembly line but a kinetic battleground where the speed of polypeptide synthesis is constantly balanced against translational accuracy. In this highly processive and cyclical process, the ribosome moves along the mRNA transcript, decoding the genetic message one codon at a time to covalently link amino acids into a growing polypeptide chain. This section examines the core steps of the elongation cycle, the factors that modulate its speed, and the cellular responses to events that cause ribosomes to stall.
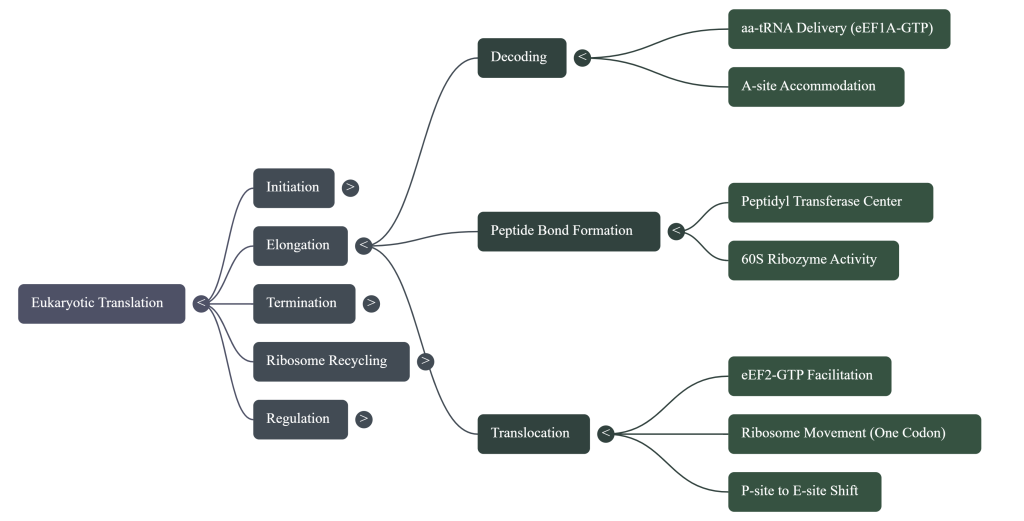
3.1. The Elongation Cycle
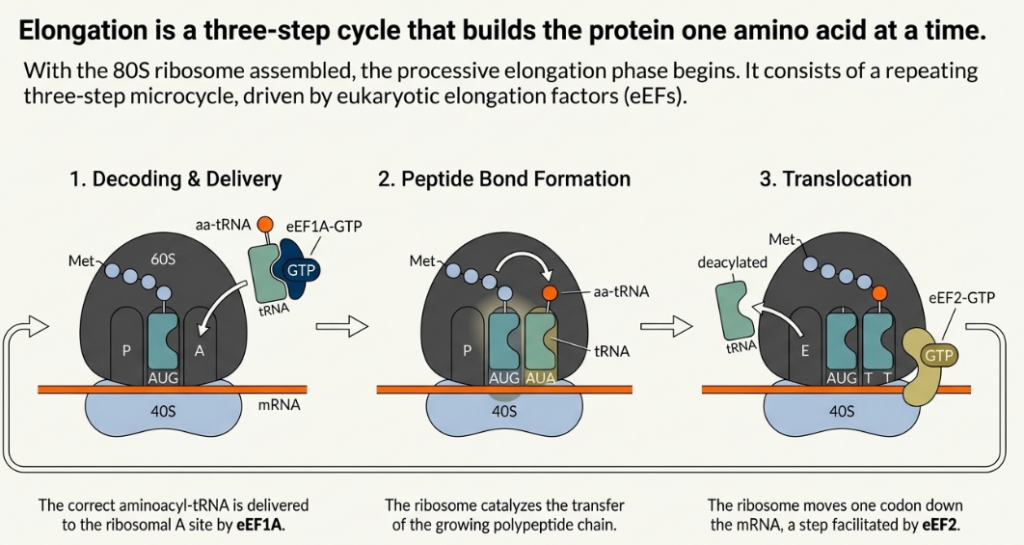
The elongation cycle can be broken down into three fundamental, repeated steps:
1. Aminoacyl-tRNA Accommodation: A cognate aminoacyl-tRNA, corresponding to the codon present in the ribosomal A-site, is delivered to the ribosome. This delivery is mediated by the eukaryotic elongation factor eEF1A, which binds the aminoacyl-tRNA in a GTP-dependent manner.
2. Peptide Bond Formation: Following the successful accommodation of the correct tRNA in the A-site, the peptidyl transferase center (PTC), a catalytic hub composed primarily of ribosomal RNA (rRNA) within the 60S subunit, catalyzes the formation of a peptide bond. This reaction transfers the growing polypeptide chain from the tRNA in the P-site to the amino group of the amino acid attached to the tRNA in the A-site.
3. Translocation: The elongation factor eEF2 promotes the movement of the ribosome one codon further along the mRNA in the 3′ direction. This translocation event shifts the tRNAs from the A and P sites into the P and E sites, respectively. The deacylated tRNA in the E-site is then released, leaving the A-site vacant and ready to accept the next aminoacyl-tRNA, thereby completing the cycle.
3.2. Determinants of Elongation Rate
The rate of translation elongation is not uniform and is influenced by several key factors. One of the primary determinants is codon optimality, which refers to the correlation between the frequency of a codon’s usage in a transcript and the cellular abundance of its corresponding tRNA. Codons that are recognized by abundant tRNA species (“optimal” codons) are decoded rapidly, while codons corresponding to low-abundance tRNAs (“rare” or “non-optimal” codons) are decoded more slowly. This can lead to ribosome pausing, which in turn can influence protein folding and mRNA stability. Furthermore, post-transcriptional modifications of tRNAs, particularly in the anticodon loop, can significantly affect decoding efficiency and the accuracy of translation.
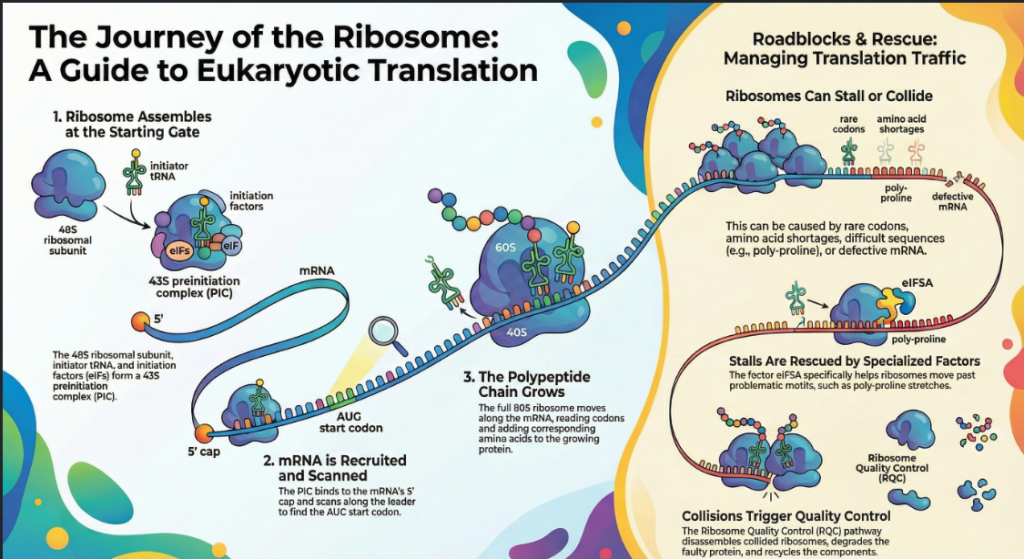
3.3. Ribosome Stalling and Resolution
While codon optimality causes transient pauses, certain mRNA sequences can induce more prolonged ribosome stalling. Stretches of proline codons (poly-proline motifs), for instance, are notoriously difficult for the ribosome to translate. To overcome such roadblocks, cells employ specialized factors. The translation factor eIF5A is critical for resolving stalls at specific problematic motifs, including not only poly-proline but also sequences containing codons for glycine, aspartate, and others. Recent evidence suggests that eIF5A’s role is more central than previously appreciated; it may function more broadly to enhance the rate of peptide bond formation in every elongation cycle and even to stimulate the rate of peptide release during termination. When stalling is not resolved, it can trigger downstream quality control pathways. The end of the coding sequence presents a final, unique challenge that requires a switch from elongation to termination machinery.
4. Phase III & IV: Termination, Recycling, and Reinitiation
The process of translation concludes when the ribosome reaches the end of the open reading frame, signaled by one of three stop codons. This event triggers a highly coordinated series of events: the release of the newly synthesized polypeptide, the dissociation of the ribosomal subunits, and their recycling for subsequent rounds of translation. This canonical pathway, however, is not absolute. Cells can employ regulated exceptions, such as stop codon readthrough and reinitiation, to generate protein diversity and exert additional layers of control.
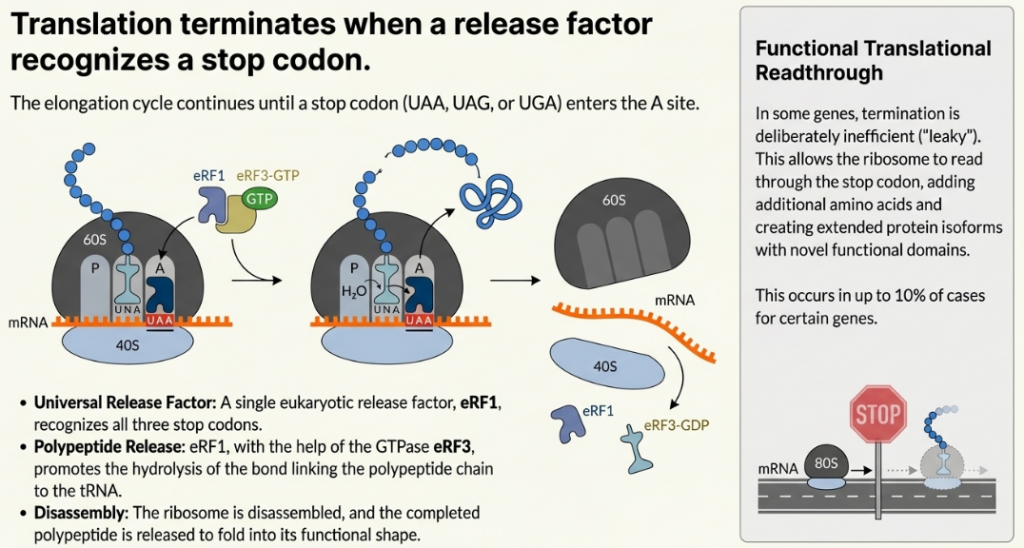
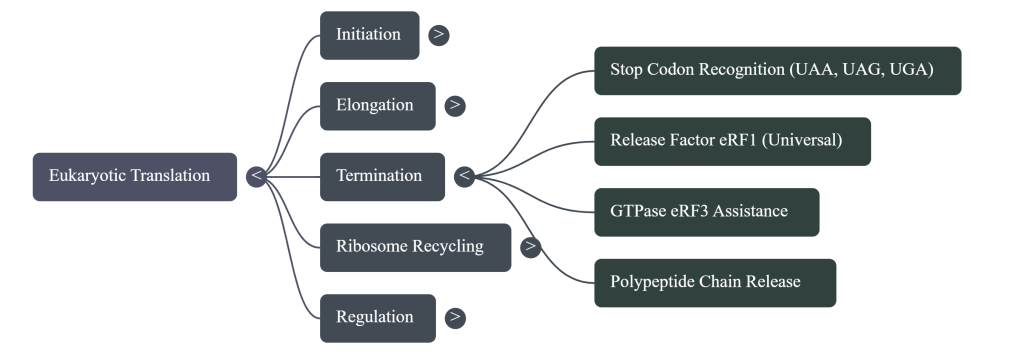
4.1. Canonical Termination and Polypeptide Release
When a stop codon (UAA, UAG, or UGA) enters the ribosomal A-site, it is not recognized by a tRNA. Instead, it is bound by a complex of eukaryotic release factors, eRF1 and the GTPase eRF3. eRF1 is the factor that recognizes all three stop codons. The binding of this complex, coupled with GTP hydrolysis by eRF3, induces a conformational change that positions a conserved GGQ (Gly-Gly-Gln) motif of eRF1 into the peptidyl transferase center (PTC). Rather than catalyzing peptide bond formation, this motif promotes the hydrolysis of the ester bond linking the completed polypeptide chain to the P-site tRNA, leading to the release of the nascent protein.
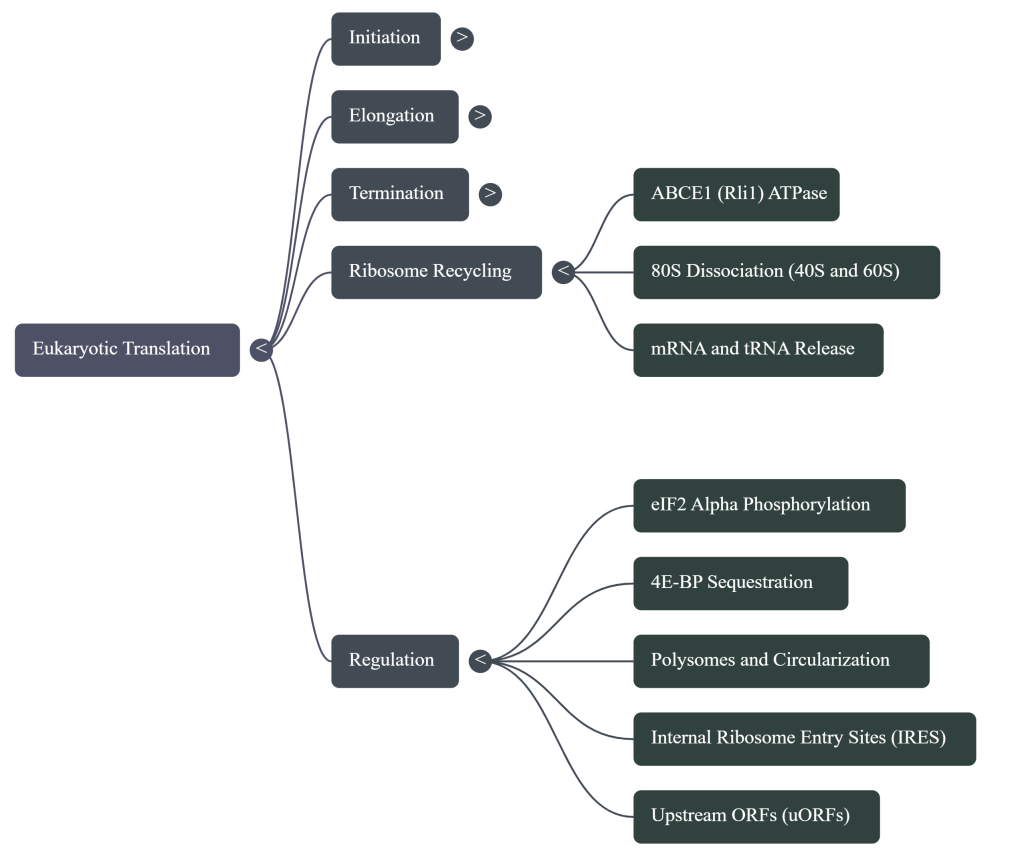
4.2. Ribosome Recycling
Following polypeptide release, the ribosome remains on the mRNA as a post-termination complex with a deacylated tRNA. The dissociation of this complex is an active, energy-dependent process. The key player in this stage is the ATPase ABCE1, which binds to the post-termination ribosome and utilizes the energy from ATP hydrolysis to split it into its constituent 40S and 60S subunits. This event also facilitates the release of the deacylated tRNA and the mRNA, freeing all components to participate in a new round of translation.
4.3. Regulated Deviations: Readthrough and Reinitiation
Cells can subvert the standard termination and recycling processes for specific regulatory purposes.
• Stop Codon Readthrough: At a stop codon, a kinetic competition occurs between the binding of the eRF1/eRF3 complex and the binding of a near-cognate aminoacyl-tRNA that can weakly base-pair with the codon. Normally, termination is highly favored. However, certain conditions can increase the likelihood of readthrough, where an amino acid is inserted at the stop codon and elongation continues. Factors that promote readthrough include the specific stop codon sequence (UGA is often “leakiest”), the surrounding mRNA sequence context, and the presence of downstream secondary structures. This phenomenon, while rare, is a regulated mechanism for producing C-terminally extended protein isoforms with novel functions.
• Reinitiation: After translating a short upstream open reading frame (uORF), the 40S ribosomal subunit does not always fully dissociate from the mRNA. Instead, it can remain associated but is initially initiation-incompetent because it lacks a ternary complex (TC). It resumes scanning in a 3′ direction, and the probability of reinitiating at a downstream ORF depends on the efficiency and distance required to recruit a new TC. This process is a sophisticated regulatory mechanism, as the efficiency of reacquiring the TC, and thus reinitiating, can be modulated by cellular conditions.
The entire translation cycle, from initiation to recycling, is not a static pathway but a dynamic one, subject to multiple, overlapping layers of regulation that allow the cell to fine-tune protein synthesis in response to its needs.
5. Landscapes of Translational Control
Cells must dynamically modulate protein synthesis to adapt to developmental programs and respond to environmental stress. This section explores the cell’s sophisticated toolbox for such regulation, revealing translation not as a constitutive process but as a major nexus for cellular signaling. Control exerted at virtually every stage allows for rapid changes in the proteome without the need for new transcription and RNA processing. Key paradigms of translational control include regulation by protein phosphorylation, the role of cis-acting mRNA elements like upstream ORFs, and the influence of trans-acting factors such as RNA-binding proteins and microRNAs.
5.1. Regulation by Phosphorylation
Reversible protein phosphorylation is a primary mechanism for transmitting cellular signals to the translational machinery, enabling rapid and global adjustments to protein synthesis rates.
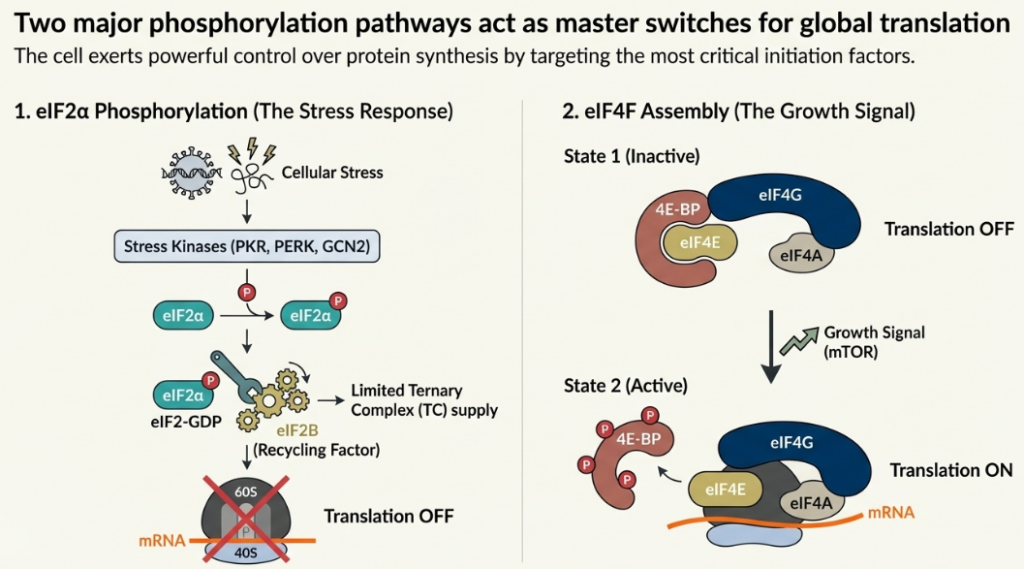
• eIF2α Phosphorylation and the Integrated Stress Response (ISR): A variety of cellular stresses, including amino acid starvation, viral infection, and endoplasmic reticulum (ER) stress, converge on a common pathway known as the Integrated Stress Response (ISR). Central to the ISR is the phosphorylation of the α-subunit of the initiation factor eIF2. Phosphorylated eIF2-GDP acts as a potent inhibitor of its own guanine nucleotide exchange factor (GEF), eIF2B. By tightly binding to and sequestering eIF2B, phosphorylated eIF2 prevents the recycling of eIF2-GDP to its active GTP-bound state. This leads to a dramatic drop in the cellular levels of the eIF2-TC, resulting in a global reduction in translation initiation.
• The mTOR Pathway and 4E-BPs: In contrast to the repressive ISR, the mTOR signaling pathway is a key activator of translation in response to growth factors and nutrient availability. A primary target of mTOR is a family of translational repressors known as eIF4E-binding proteins (4E-BPs). In their hypophosphorylated state, 4E-BPs bind tightly to the cap-binding protein eIF4E, preventing it from assembling into the eIF4F complex. Upon activation, mTOR hyperphosphorylates the 4E-BPs, causing them to dissociate from eIF4E. The liberated eIF4E is then free to form the eIF4F complex, promoting cap-dependent translation and driving cell growth.
5.2. Regulation by cis-Acting mRNA Elements: The Role of uORFs
Upstream Open Reading Frames (uORFs), short coding sequences located in the 5′ UTR of many eukaryotic mRNAs, are powerful cis-acting regulatory elements. While generally repressive, uORFs can function as sophisticated molecular sensors that modulate the translation of the main downstream ORF in response to cellular conditions.
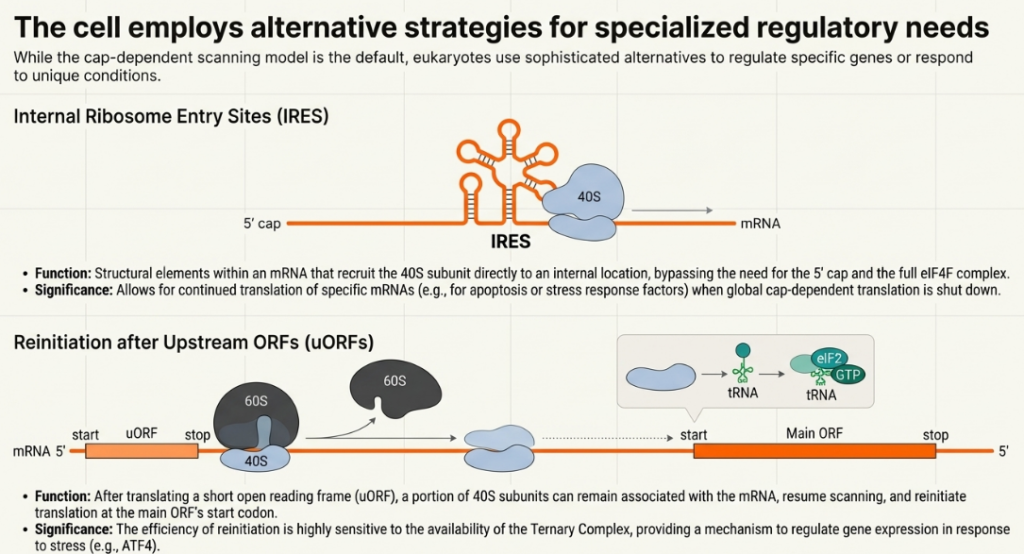
The mRNA encoding the transcription factor ATF4 provides a classic example of uORF-mediated control. Under normal conditions, where eIF2-TC levels are high, ribosomes that translate a short initial uORF (uORF1) efficiently reacquire a TC and reinitiate at a longer, repressive uORF (uORF2). Translation of this second uORF prevents initiation at the main ATF4 coding sequence. However, under stress conditions that trigger eIF2α phosphorylation and lower TC levels, the reinitiation process is delayed. Ribosomes scanning downstream of uORF1 take longer to reacquire a TC, causing them to bypass the start codon of the repressive uORF2 and instead successfully reinitiate at the main ATF4 ORF. This mechanism results in the specific translational upregulation of ATF4 during stress, precisely when its transcriptional activity is needed.
5.3. Regulation by trans-Acting Factors
Translation can also be controlled by trans-acting factors that bind directly to mRNA transcripts, typically within the untranslated regions.
• RNA-Binding Proteins: A diverse array of sequence-specific RNA-binding proteins can bind to motifs in the 3′ UTR to repress translation. A common mechanism involves the protein acting as a bridge to create an inhibitory “closed loop.” By simultaneously interacting with the 3′ UTR and a component of the initiation machinery at the 5′ end (such as eIF4E), these proteins can physically block the assembly of a productive eIF4F complex, thereby preventing initiation.
• MicroRNAs (miRNAs): MicroRNAs are small (~21 nucleotide) non-coding RNAs that act as potent post-transcriptional repressors. Guided by an Argonaute (AGO) protein, a miRNA binds to partially complementary target sites, usually located in the 3′ UTR of its target mRNA. The miRNA-AGO complex serves as a platform to recruit effector proteins, most notably members of the GW182 family. GW182 then actively dismantles the initiation-competent state of the messenger ribonucleoprotein (mRNP) particle. It can achieve this by disrupting the productive “closed loop” interaction between the poly(A)-binding protein (PABP) and eIF4G, and by recruiting enzymes that promote the deadenylation and subsequent decay of the mRNA.
The complexity of these regulatory networks necessitates robust quality control mechanisms to ensure that the process of translation remains accurate and that aberrant products are swiftly eliminated.
6. Translational Quality Control
The synthesis of a functional proteome requires not only precise regulation but also high fidelity. Errors during translation, such as ribosome stalling or the translation of faulty mRNAs, can produce truncated or misfolded proteins that are toxic to the cell. To mitigate these risks, eukaryotes have evolved a sophisticated suite of quality control mechanisms that act as a surveillance system, identifying and resolving aberrant translational events to maintain proteome integrity.
6.1. Ribosome-Associated Quality Control (RQC)
Ribosomes can stall on an mRNA for various reasons, including encountering damaged RNA or translating difficult sequences. If a stalled ribosome is not resolved, trailing ribosomes can collide with it, creating a “traffic jam.” Such collisions are a signal for the Ribosome-Associated Quality Control (RQC) pathway. The RQC complex is recruited to the stalled ribosomal subunit, where it initiates a multi-pronged response. It targets the partially synthesized nascent polypeptide chain for ubiquitination and subsequent degradation by the proteasome, and it facilitates the splitting and recycling of the stalled ribosomal subunits, freeing them for future use.
6.2. Nonsense-Mediated mRNA Decay (NMD)
Nonsense-Mediated mRNA Decay (NMD) is a crucial surveillance pathway that protects the cell from the potentially harmful effects of truncated proteins. NMD recognizes and selectively degrades mRNAs that contain a premature termination codon (PTC). PTCs can arise from nonsense mutations in the DNA or errors during transcription or splicing. By eliminating these faulty transcripts before they can be fully translated, NMD ensures that cellular resources are not wasted producing non-functional or dominant-negative protein fragments. The discovery of these intricate quality control and regulatory mechanisms has been greatly accelerated by modern experimental techniques that provide a global view of translation.
7. Modern Approaches to Interrogating Translation
In recent decades, technological advancements have fundamentally transformed the study of translation, moving from the analysis of single genes to genome-wide landscapes. Among these, Ribosome Profiling (Ribo-Seq) has emerged as a particularly transformative technique. By providing a “snapshot” of all ribosome positions across the entire transcriptome at nucleotide resolution, Ribo-Seq has generated unprecedented insights into the dynamics, regulation, and complexity of protein synthesis in vivo.
7.1. Principles and Insights from Ribosome Profiling (Ribo-Seq)
The core principle of Ribo-Seq is to capture and sequence the small fragments of mRNA that are physically protected by translating ribosomes. The experimental workflow involves several key steps: (1) translation is arrested, often using inhibitors or flash-freezing; (2) cellular lysates are treated with ribonucleases to digest all mRNA not enclosed within a ribosome; and (3) the 80S ribosomes containing these protected mRNA fragments, or “footprints,” are isolated. These footprints, typically 18-34 nucleotides in length, are then converted to a cDNA library and subjected to deep sequencing.
By mapping these footprint reads back to a reference transcriptome and comparing their density to parallel measurements of mRNA abundance (RNA-Seq), researchers can derive a wealth of information, including:
• Identification of translated ORFs: The characteristic three-nucleotide periodicity of ribosome footprints allows for the precise identification of actively translated coding sequences, including novel uORFs and N-terminally extended proteoforms.
• Measurement of ribosome density: The number of footprints mapping to a gene provides a direct measure of its translation level.
• Calculation of Translation Efficiency (TE): The ratio of ribosome footprint density to mRNA abundance for a given gene provides a relative measure of its translation efficiency.
• Analysis of translation dynamics: Ribo-Seq can pinpoint specific locations of ribosome pausing, such as at rare codons (site-specific pause) or at stop codons (queuing), and reveal translation occurring in unexpected regions like 5′ and 3′ UTRs.
7.2. Methodological Artifacts and Considerations
While Ribo-Seq offers unprecedented resolution, its power is contingent upon a rigorous understanding of its inherent methodological biases. Failure to account for these potential artifacts, which can arise at multiple stages of the protocol, can lead to significant misinterpretation of translational dynamics. Key considerations include:
• Use of Translation Inhibitors: Pre-treatment of cells with translation inhibitors can significantly distort ribosome density profiles. For example, cycloheximide, a commonly used elongation inhibitor, can cause an artificial accumulation of ribosome footprints near the start codon. This occurs because the drug efficiently arrests elongating ribosomes but is less effective against initiating ribosomes, which can still assemble on the mRNA and join the queue, leading to a pile-up.
• Nuclease and Ligation Biases: The enzymes used during library preparation, such as the nucleases for digestion and the ligases for adapter attachment, can exhibit sequence specificity. This can lead to the over- or under-representation of certain footprint sequences, introducing bias that is unrelated to the actual ribosome occupancy.
• Fragment Size Selection: Standard Ribo-Seq protocols often involve selecting a narrow range of footprint lengths (e.g., ~30 nt). This may inadvertently exclude footprints from alternative ribosome conformations. For instance, collided or queued ribosomes can protect longer mRNA fragments, and selecting only for monosome-sized footprints may deplete the data of these important stalling events.
Despite these limitations, careful experimental design and bioinformatic analysis allow these new methodologies to continue to shape and refine our understanding of the complex landscape of translation dynamics.
8. Conclusion and Future Perspectives
Eukaryotic translation is not a static, assembly-line process but a highly dynamic and exquisitely regulated pathway that lies at the core of cellular life. As this review has detailed, translational control is a story of fundamental trade-offs. The cell must balance the speed of elongation against the fidelity of decoding, and global shifts in protein synthesis via signaling pathways like mTOR and the ISR must be reconciled with gene-specific regulation conferred by cis-acting uORFs and trans-acting factors. Likewise, canonical mechanisms for initiation and termination are punctuated by alternative pathways like IRES entry and stop codon readthrough, which provide crucial regulatory flexibility. This intricate regulatory network is surveyed by sophisticated quality control mechanisms that ensure the fidelity of protein synthesis and the integrity of the proteome.

The advent of powerful techniques like ribosome profiling has provided a systems-level view of these processes, but has also underscored the complexity and remaining challenges in the field. Looking ahead, a primary goal will be the continued optimization of experimental methods to minimize artifacts and provide an even clearer picture of translation in vivo. The future of the field lies in the integration of multiple omics approaches. Combining ribosome profiling with techniques like RNA-protein crosslinking and coupling these global analyses with genome editing will enable the precise dissection of the roles of known and putative translation factors. These powerful combinations promise to deepen our understanding of the molecular mechanisms governing gene expression and will be invaluable for exploring the genome-wide effects of drugs that target the translation machinery, opening new avenues for therapeutic intervention.
Image Summary
References
Blanchet, S., Ranjan, N. (2022). Translation Phases in Eukaryotes. In: Entian, KD. (eds) Ribosome Biogenesis. Methods in Molecular Biology, vol 2533. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2501-9_13
Dominique Chu, Tobias von der Haar, The architecture of eukaryotic translation, Nucleic Acids Research, Volume 40, Issue 20, 1 November 2012, Pages 10098–10106, https://doi.org/10.1093/nar/gks825
Hinnebusch AG, Lorsch JR. The mechanism of eukaryotic translation initiation: new insights and challenges. Cold Spring Harb Perspect Biol. 2012 Oct 1;4(10):a011544. doi: 10.1101/cshperspect.a011544. PMID: 22815232; PMCID: PMC3475172.
Jackson, R., Hellen, C. & Pestova, T. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat Rev Mol Cell Biol 11, 113–127 (2010). https://doi.org/10.1038/nrm2838
Sokabe M, Fraser CS. Toward a Kinetic Understanding of Eukaryotic Translation. Cold Spring Harb Perspect Biol. 2019 Feb 1;11(2):a032706. doi: 10.1101/cshperspect.a032706. PMID: 29959192; PMCID: PMC6360857.