Introduction: The Unseen World of Medical Diagnostics

In the landscape of modern medicine, the ability to accurately and swiftly detect the molecular signatures of disease is paramount. Diagnostic tools are the unseen engines that power clinical decisions, from confirming a pregnancy to tracking the spread of a viral outbreak. For decades, one technique has stood out as a foundational and versatile workhorse in laboratories worldwide: the Enzyme-Linked Immunosorbent Assay, or ELISA. This elegant and robust method has become a cornerstone of medical testing, providing critical insights into the invisible battles being waged within our bodies.

This article charts the evolutionary journey of this remarkable technology. We will begin by exploring the fundamental principles that make ELISA so effective and examine the various formats that have been adapted for different diagnostic needs. From there, we will trace its progression from a reliable but limited tool to the new frontier of ultrasensitive, single-molecule detection platforms. This evolution culminates in groundbreaking frameworks like the EV single-molecule array (eSimoa), which promises to redefine early disease detection by decoding biological information with unprecedented precision. Our exploration begins with the core principles that have made ELISA an indispensable tool for nearly half a century.
1. The Bedrock of Detection: Understanding the ELISA Technique
To fully appreciate the quantum leaps made in diagnostic technology, it is essential to first understand the core principles of immunoassays. This foundational knowledge provides the strategic context for the subsequent advancements, highlighting how each new generation of tests builds upon the simple yet powerful interaction between two key biological molecules.
ELISA, an acronym for Enzyme-Linked Immunosorbent Assay, is a laboratory technique designed to detect and quantify substances such as antibodies, antigens, proteins, and hormones in bodily fluids like blood, urine, or saliva. The technique is a type of immunoassay, which relies on the highly specific binding interaction between an antigen and an antibody. Antigens are molecular markers—usually proteins or sugars—found on the surfaces of pathogens and cells, including viruses, bacteria, parasites, allergens, and tumor cells. Antibodies are specialized proteins produced by the immune system that recognize and bind to these specific antigens to help eliminate them from the body.

The ELISA method was developed in the early 1970s as a safer and more practical alternative to radioimmunoassay (RIA), which used radioactive materials. Two independent research teams, led by Engvall and Perlman, and Van Weemen and Schuurs, are credited with its simultaneous invention. Since then, its versatility has made it a routine method in both research and clinical diagnostics across the globe.
The clinical utility of ELISA is vast, enabling a wide range of critical medical tests:
• Diagnosing Infections: Detecting antibodies produced in response to bacteria (e.g., Lyme disease), viruses (e.g., HIV, Hepatitis A/B/C), and fungi (e.g., Candida).
• Identifying Autoimmune Conditions: Measuring autoantibodies that mistakenly attack the body’s own cells, as seen in conditions like Type 1 diabetes.
• Detecting Hormones: Confirming pregnancy by measuring levels of human chorionic gonadotropin (hCG).
• Screening and Monitoring: Screening donated blood for viral contaminants, tracking disease outbreaks in populations, and detecting drug abuse.
At its core, every ELISA variation harnesses this antigen-antibody partnership. The “enzyme-linked” component refers to a specific enzyme, such as horseradish peroxidase (HRP) or alkaline phosphatase (AP), chemically attached to an antibody. When the corresponding substrate is added, the enzyme catalyzes a reaction that produces a measurable signal—often a color change, like the blue product from HRP’s reaction with its substrate—indicating the presence and quantity of the target substance. From this elegant foundation, several distinct methodologies have emerged to answer different clinical questions.
2. The ELISA Toolkit: A Comparative Analysis of the Four Major Formats
The broad utility of ELISA is enabled by its adaptability into several distinct formats. Different diagnostic questions—such as detecting an antibody versus an antigen, or prioritizing speed versus sensitivity—require different methodological approaches. Over the years, four primary formats have become standard in the laboratory, each with a unique architecture of antibodies and antigens and a corresponding set of strengths and weaknesses.

Direct ELISA
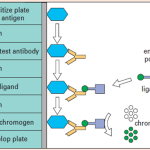
A direct ELISA involves an enzyme-conjugated primary antibody that binds directly to an antigen that has been coated onto the assay plate.
| Advantages | Disadvantages |
| Rapid due to fewer steps. | High cost of reaction. |
| Eliminates secondary antibody cross-reactivity. | Low sensitivity compared to other formats. |
| No signal amplification. | |
| Inflexible; requires a specific conjugated antibody for each target. |
Indirect ELISA
In an indirect ELISA, an unlabeled primary antibody first binds to the antigen, which is then detected by an enzyme-conjugated secondary antibody that specifically binds to the primary antibody.
| Advantages | Disadvantages |
| Higher sensitivity than Direct ELISA due to signal amplification. | Risk of cross-reactivity from the secondary antibody. |
| More flexible; one labeled secondary antibody can be used with many primary antibodies. | Longer procedure due to an extra incubation step. |
| Less expensive. |
Sandwich ELISA
The target antigen is “sandwiched” between two distinct antibodies—a capture antibody coated on the plate and an enzyme-conjugated detection antibody.

| Advantages | Disadvantages |
| Highest sensitivity among all ELISA types. | Time-consuming and expensive. |
| High specificity due to using two antibodies for the antigen. | Requires the use of “matched pair” antibodies. |
| Suitable for complex/impure samples as the antigen is purified by the capture antibody. | Antibody optimization can be difficult. |
Competitive ELISA
The concentration of a target substance in a sample is determined by its ability to compete with a labeled reference molecule for a limited number of binding sites; thus, a lower signal indicates a higher concentration of the target in the sample.
| Advantages | Disadvantages |
| Less sample purification needed. | Low specificity. |
| Can be used for small antigens. | Cannot be used in dilute samples. |
| Low variability. | |
| Can measure a large range of antigens. |
While these four formats have served as the bedrock of diagnostics for decades, the relentless scientific drive for earlier and more precise disease detection created a demand for technologies that could push beyond their inherent limits.
3. Pushing the Boundaries: The Leap Toward Ultrasensitive Detection
The next evolutionary leap in diagnostics was driven by a critical clinical need: the ability to detect disease biomarkers at extremely low concentrations. Identifying these molecular whispers is essential for diagnosing diseases like cancer at their earliest, most treatable stages. However, traditional ELISA formats, despite their utility, often lack the sensitivity required to detect rare target proteins circulating in complex samples like blood plasma. This limitation spurred a wave of innovation aimed at dramatically amplifying the diagnostic signal.

Initial incremental advancements bridged the gap between conventional colorimetric detection and the next generation of assays. Researchers developed techniques that used fluorogenic, quantitative PCR, and electrochemiluminescent reporters to generate stronger and more quantifiable signals. While these methods improved sensitivity, a truly revolutionary breakthrough came with the development of Single Molecule Array (Simoa) technology.
Simoa represents a paradigm shift in protein detection. This represents a fundamental shift from traditional analog methods, which measure the average signal from all molecules in a sample, to a digital approach that counts individual molecular events. By isolating and detecting single enzyme molecules in thousands of femtoliter-sized wells, Simoa effectively digitizes protein detection, providing its core value proposition: unrivaled sensitivity. Simoa is capable of quantifying proteins at attomolar (aM; 10⁻¹⁸ M) concentrations, representing an astounding million-fold improvement over existing methods. This leap in sensitivity opened the door to analyzing biomarkers that were previously undetectable, particularly those contained within one of the most promising sources of diagnostic information: extracellular vesicles.
4. The Next Generation: Spatially Decoding Biomarkers with eSimoa
The latest frontier in diagnostics involves a strategic shift from analyzing bulk fluid samples to decoding spatially compartmentalized information within nanoscale biological structures. This approach recognizes that the location of a biomarker—whether on the surface or inside a particle—can provide critical insights into its function and origin.
A key focus of this new approach is Extracellular Vesicles (EVs). These are nanosized particles, enclosed by a lipid bilayer, that are released by all cells in the body. They act as “biomarker reservoirs,” carrying a cargo of proteins and nucleic acids that directly reflects the biological state of their parent cells. Because tumor cells actively release EVs, these particles hold immense promise for the non-invasive diagnosis and monitoring of cancer.

To unlock this potential, a groundbreaking framework known as the EV single-molecule array (eSimoa) was developed. This innovative method combines the targeted isolation of EVs with the high-throughput power of Simoa technology. Its core purpose is to enable the spatial decoding of EV protein biomarkers, distinguishing between those on the surface and those inside, with exceptional sensitivity and resolution.

The eSimoa framework is built on three complementary pipelines, each designed to answer a specific diagnostic question about the spatial arrangement of proteins:
• i. Surface eSimoa: This pipeline captures and detects intact EVs based on the simultaneous presence, or co-localization, of two distinct proteins on their surface. This pipeline answers the question: ‘Which distinct EV subpopulations, defined by their surface markers, exist in a sample?’
• ii. Luminal eSimoa: This pipeline is designed to quantify the absolute concentration of proteins located inside the EV (luminal proteins) by first lysing the EVs to release their internal contents for measurement. This pipeline answers: ‘What is the total concentration of a specific internal protein across the entire EV population?’
• iii. Pulldown (Surface-Luminal) eSimoa: This integrated pipeline first captures a specific subpopulation of EVs from a sample based on a single surface protein, then analyzes the luminal proteins contained only within that captured group. This powerful integrated approach answers the most clinically relevant question: ‘What is the internal cargo of a specific, targeted EV subpopulation?’

The clinical power of this framework has been demonstrated by its ability to detect the KRASG12D protein—a highly specific marker for pancreatic ductal adenocarcinoma—within EVs derived from pancreatic tumor cells. This is a crucial step toward developing a minimally invasive blood test for a notoriously hard-to-detect cancer.
The key advancements of the eSimoa framework can be summarized as follows:
• Unmatched Sensitivity: It can detect specific EV populations in plasma at concentrations as low as 10⁵ EVs per mL and quantify protein concentrations down to the femtomolar (fM) level. This sensitivity allows for the detection of tumor-derived EVs that may constitute a fraction as low as five orders of magnitude below the total EV population in blood.

• Absolute Quantification: Unlike many other methods that provide relative measurements, eSimoa provides the absolute concentration of luminal proteins like the general tumor marker RAS and the cancer-specific KRASG12D mutant.
• Clinical Feasibility: The framework can be applied directly to clinical samples like plasma with minimal pre-purification, making it automated, scalable, and suitable for high-throughput laboratory settings.
This ability to precisely quantify spatially defined biomarkers represents a profound step forward, transitioning diagnostics from a broad overview to a high-resolution molecular snapshot.
5. Conclusion: The Future of Diagnostics is Precise and Personalized
The journey from the foundational color-changing reactions of the 1970s to the single-molecule precision of today marks a dramatic evolution in diagnostic science. We have traveled from the broad utility of the four classic ELISA formats—each a powerful tool in its own right—to the highly specific, ultrasensitive eSimoa framework that can decode the molecular cargo of individual nanoparticles. This progression is not merely an academic achievement; it represents a fundamental shift in our ability to understand and combat disease.

This leap from bulk analysis to spatially-defined, single-molecule quantification is the engine of personalized medicine. The ability to isolate a specific subpopulation of EVs from a patient’s blood and precisely measure its cancer-specific cargo, such as the KRASG12D mutant protein, enables therapies to be tailored not just to a disease, but to the unique molecular activity of that disease in that individual at that moment. As these ultrasensitive diagnostic frameworks continue to be refined and applied, they hold the transformative potential to significantly improve patient outcomes in oncology and a wide array of other diseases, heralding a future where medicine is more predictive, precise, and powerful than ever before.


Image Summary


References
Rudolf M Lequin, Enzyme Immunoassay (EIA)/Enzyme-Linked Immunosorbent Assay (ELISA), Clinical Chemistry, Volume 51, Issue 12, 1 December 2005, Pages 2415–2418, https://doi.org/10.1373/clinchem.2005.051532
Crowther, J.R. (1995). Basic Principles of ELISA. In: Crowther, J.R. (eds) ELISA. Methods in Molecular Biology™, vol 42. Humana Press. https://doi.org/10.1385/0-89603-279-5:35
https://www.thermofisher.com/np/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-elisa.html
https://my.clevelandclinic.org/health/articles/24990-elisa
https://www.ncbi.nlm.nih.gov/books/NBK555922/
https://en.wikipedia.org/wiki/ELISA
Wang X, Ogata AF, Walt DR. Ultrasensitive Detection of Enzymatic Activity Using Single Molecule Arrays. J Am Chem Soc. 2020 Sep 2;142(35):15098-15106. doi: 10.1021/jacs.0c06599. Epub 2020 Aug 20. PMID: 32797755; PMCID: PMC7472518.
Cheng, C. A., Hou, K. C., Hsu, C. W., & Chiang, L. C. (2024). Ultrasensitive and high‐resolution protein spatially decoding framework for tumor extracellular vesicles. Advanced Science, 11(3), 2304926. https://doi.org/10.1002/advs.202304926