1. Introduction to the Global Food Security Challenge

The strategic urgency of modern plant breeding is framed by the imperative to sustain a global population projected to reach 9.6 billion by 2050, a milestone that requires a 70% increase in agricultural productivity. This goal is currently besieged by intensifying biotic and abiotic stresses; pests and diseases alone account for approximately USD 470 billion in annual global losses, while a 1°C rise in mean temperature is estimated to reduce wheat yields by roughly 6%. Consequently, the breeding paradigm is shifting from traditional phenotype-based selection toward data-driven genomic strategies capable of building climate-resilient cultivars. This transition represents a vital move from reacting to environmental pressure to predicting and engineering genetic outcomes at the scale required for global food security.

Agricultural Pressures
The following table illustrates the dual stressors currently eroding our global food foundations:
| Stress Category | Pressures & Impact Data |
|---|---|
| Biotic Stresses | Pests and diseases account for 30–40% of global losses (>$470B USD annually). Fungal infections alone reduce yields by 10–15%. |
| Abiotic Stresses | Each 1°C rise in temperature reduces wheat yield by ~6%. Drought can lower productivity by 40%, while poor soil fertility causes 30–50% of yield gaps in developing regions. |
2. The Modern Breeding Gap: Why We Need New Tools
Despite our successes, traditional breeding is hitting a wall. Historically, breeders relied on phenotype-based selection—choosing the best-performing plants and crossing them. More recently, Genomic Selection (GS) has improved this by using genome-wide markers to predict breeding values. However, both methods are essentially “closed-loop systems,” limited by a species’ own history and unable to precisely navigate complex, polygenic traits like climate resilience.
Critical Constraints of Traditional Breeding:
• Environmental Variability: Selection is often “noisy” because the environment masks a plant’s true genetic potential, making it difficult to predict performance across shifting climates.
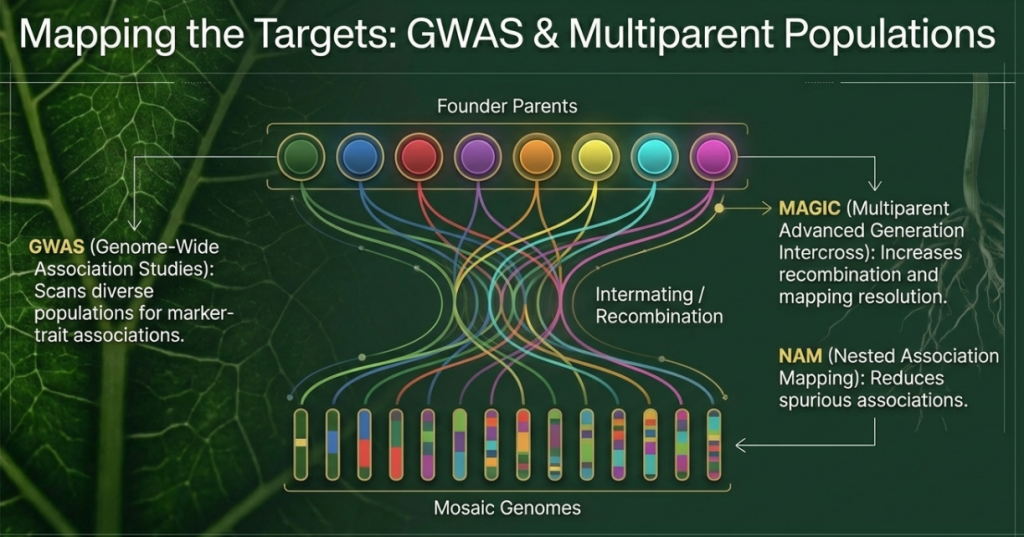
• Missing Heritability: Methods like GWAS (Genome-Wide Association Studies) often fail to identify the many small-effect loci that collectively drive complex traits like heat or drought tolerance.
• Long Breeding Cycles: Developing a new stable cultivar can take years or even decades, a pace that cannot keep up with the accelerating rate of climate change.
Mating Systems and Cultivar Design
| Mating System | Scientific Cultivar Type | Crop Examples |
|---|---|---|
| Self-Pollination | Inbred lines / Pure lines | Wheat, Rice |
| Cross-Pollination | Hybrids / Open-pollinated | Maize, Onion, Sunflower |
| Vegetative/Asexual | Clones | Potato, Sugar Cane |
Traditional methods are confined to the existing gene pool. To secure our 2050 food supply, we must transition to an “open-source architecture” for biology—a programmable framework that can borrow resilience from across the biological kingdoms.
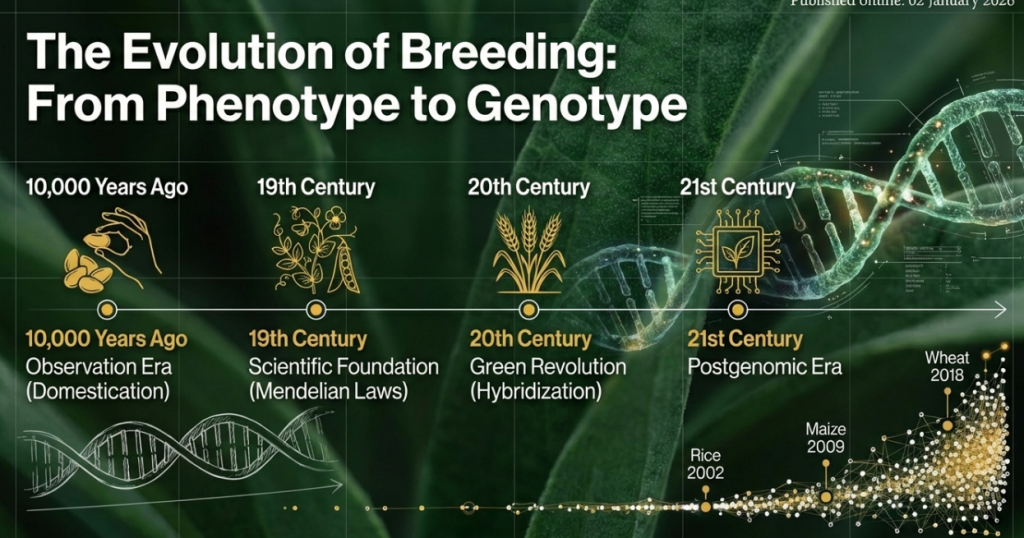
3. The Evolution of Breeding Paradigms: From Landraces to Genomics
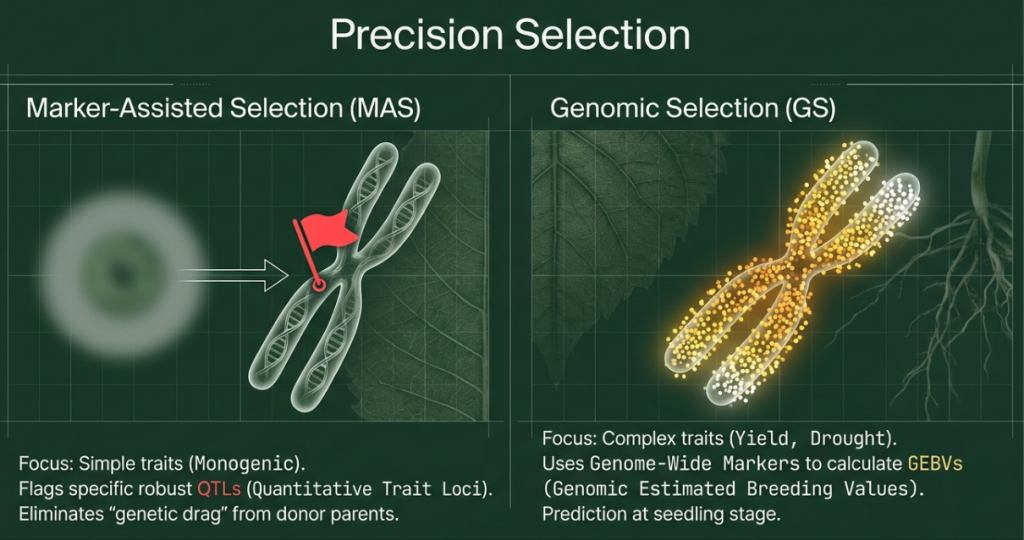
Traditional breeding methodologies, while successful during the Green Revolution, have reached a physiological and statistical plateau for complex, polygenic traits. Early molecular approaches, such as Marker-Assisted Selection (MAS) and Genome-Wide Association Studies (GWAS), initially promised to bridge this gap but ultimately struggled to resolve the genetic architecture of traits like yield and drought tolerance.

Limitations of Pre-Genomic Methodologies:
• The “Missing Heritability” Problem: GWAS often identifies loci that collectively explain only a fraction of total phenotypic variance, failing to account for the numerous small-effect variants that drive complex traits.
• Inefficacy with Polygenic Traits: MAS is optimized for traits governed by major-effect Quantitative Trait Loci (QTLs). It performs poorly for traits with a polygenic architecture where hundreds of loci contribute minute individual effects.
• Context-Dependency and Low Transferability: The effects of identified QTLs are frequently specific to certain populations or environments, limiting their utility in diverse breeding programs.
• Resource Intensity: The requirements for fine mapping and candidate gene validation in GWAS/MAS necessitate extensive recombination and large populations, significantly slowing genetic gain.
The advent of Genomic Selection (GS) represents a fundamental shift. As proposed by Meuwissen et al. (2001), GS utilizes genome-wide marker data to estimate Genomic Estimated Breeding Values (GEBVs). Rather than searching for isolated major-effect loci, GS captures the cumulative contribution of numerous small-effect loci across the entire genome, facilitating an unprecedented acceleration in the breeding cycle.
4. The Conceptual Framework of Integrated Genomic Selection (GS)
The strategic importance of GS lies in its predictive power, which reconfigures the breeding pipeline to prioritize genomic potential, thereby drastically reducing the time required for field-based phenotyping.
Workflow and Workflow Synthesis: The GS framework functions through the coordinated interaction of two distinct populations:
1. Training Population (TRN): This cohort is both genotyped and phenotyped. High-throughput phenotyping (via drones, remote sensing, and imaging) captures multi-environment variation, which is then used to calibrate statistical models.
2. Testing Population (TST): This cohort is genotyped but not phenotyped. The calibrated GS model predicts the GEBVs of these individuals, allowing for the selection of superior progeny before they are ever field-tested.
Comparison of Primary GS Statistical Models
| Model Type | Primary Application | Marker Effect Distribution Assumptions |
|---|---|---|
| GBLUP | Additive effects and highly polygenic traits. | Assumes all markers have a small, equal variance effect. |
| Bayesian Models | Heterogeneous marker effects and sparsity. | (e.g., BayesB) Uses “spike-and-slab” distributions; assumes many markers have zero effect while a few have large effects. |
| Machine/Deep Learning | Non-linear and epistatic interactions. | No rigid assumptions; captures higher-order gene-to-gene interactions and non-additive effects. |
The Statistical Challenge: p>n and Regularization In GS, breeders face the “p>n” problem, where the number of markers (p) exceeds the number of individuals (n). This imbalance poses a severe risk of overfitting, where the model “memorizes” noise rather than identifying generalizable patterns. To mitigate this, models utilize shrinkage and regularization. Shrinkage acts as a mathematical penalty, reducing the effect size of markers with little explanatory power. This ensures the model does not over-react to stochastic variations in the training data, maintaining robustness across the Testing Population.
5. Transitioning to Perennial Cultivation: A Sustainability Mandate
Restructuring crops from annual to perennial systems is a sustainability mandate aimed at reconfiguring agricultural land to provide permanent “ecosystem services.” Perennial crops develop deep, robust root systems that fundamentally alter nutrient and carbon cycles.
Sustainability Differentiators:
• Soil and Resource Retention: Perennials enhance nitrogen (N) retention and promote soil carbon accumulation, significantly reducing nutrient leaching and greenhouse gas emissions.
• Soil Architecture Protection: Permanent soil cover prevents the erosion and eutrophication typical of annual systems, where soil is left bare after harvest.
• Efficiency in Inputs: Established perennials potentially lower requirements for pesticides, labor, and fertilizers, as they do not require annual replanting and tillage.
Case Study Synthesis:
• Perennial Rice (PR23): Developed through interspecific hybridization (O. sativa x O. longistaminata), PR23 can produce eight consecutive harvests over four years from a single planting. Critically, its yield of 6.8 Mg ha⁻¹ matches the 6.7 Mg ha⁻¹ of annual rice, countering the narrative of a perennial yield gap. However, its application is currently restricted to frost-free regions between 40°N and 40°S.
• Intermediate Wheatgrass (Kernza): The first food-grade perennial grain, Kernza offers high-quality grain and minimal lodging, though it currently exhibits a yield reduction of approximately 77% by the third year.
• Perennial Buckwheat (Fagopyrum cymosum): Noted for its waterlogging tolerance and frost resistance, this wild relative is a target for interspecific hybridization to improve common buckwheat, though it still carries unfavorable traits like bitterness and seed shattering.
| Feature | Annual Systems | Perennial Systems (PR23) |
|---|---|---|
| Labor Requirement | High (Replanted annually) | 58.1% reduction per regrowth cycle |
| Input Cost | Standard | 49.2% reduction in input costs |
| Soil Health | High erosion; nutrient loss | Soil carbon accumulation; N retention |
Implementation Barriers: The transition is hampered by “hybrid sterility” resulting from abnormal meiosis in early-generation hybrids. Furthermore, while PR23 shows yield parity, most perennials still face a “yield gap” relative to annual cultivars that have undergone decades of intensive selection.
6. Nature’s Secret Lab: Borrowing Microbial “Shortcuts” for Climate-Resilient Crops
6.1 The Concept of the Evolutionary Shortcut
In the traditional view of biology, plants evolve through the slow, vertical inheritance of genes from parent to offspring. However, nature has a hidden method for rapid innovation: Horizontal Gene Transfer (HGT). Once considered a purely microbial phenomenon, we now recognize HGT as a “nonconventional inheritance” method that has acted as a primary driver of land plant evolution. Rather than waiting millennia for a random mutation to address an environmental crisis, HGT allows plants to bypass slow evolutionary processes by harvesting nature’s pre-packaged solutions—moving genetic material across species boundaries and even between kingdoms of life.
HGT can be viewed as nature’s proprietary speed-dial for adaptation. It allows a plant to “borrow” a complex, pre-refined genetic module from a microbe that has already spent millions of years mastering survival in extreme conditions.
Key Insight: Why does HGT matter? Traditional evolution is incremental and constrained by a species’ own lineage. HGT allows plants to acquire complex, multi-trait adaptive capabilities—such as the ability to survive extreme cold or drought—almost instantly in evolutionary terms. It shifts the paradigm from simple genetic modification to “evolution-guided design,” where we utilize nature’s past successes to secure our future food supply.
While these events were once thought to be rare anomalies, they have actually shaped the very history of land plants, providing the genomic foundation for terrestrial life.
6.2 Ancient Microbial Footprints in Modern Genomes
Our modern crops are walking archives of ancient microbial ingenuity. By mining the genomes of plants and their wild relatives, scientists have discovered that many of the traits we rely on today were originally microbial “shortcuts” integrated into plant DNA during prehistoric environmental shifts. These genes are “preadapted” to extreme environments, providing a proven blueprint for modern resilience.
| Plant Species | Microbial Gene Origin | Survival Benefit (The Shortcut) |
|---|---|---|
| Wheat (Triticum aestivum) | Bacteria | Acquisition of Cold-Shock Protein (CSP) genes that enhance drought tolerance, photosynthesis, and grain yield. |
| Liverworts (Marchantia polymorpha) | Fungi | Fungal genes acquired before the divergence of land plants, providing the essential toolkit for drought adaptation. |
| Red Algae (Pyropia haitanensis) | Bacteria & Saccharothrix spp. | HGT-derived genes (ferrochelatase and reductase) and symbiotic actinobacteria that provide critical heat-stress tolerance in intertidal zones. |
| Green Plants (General) | Bacteria & Fungi | Acquisition of 23 glycosyl hydrolase families, enabling cell wall diversification and defense evolution. |
Because these genes have already proven compatible with plant systems over eons, they offer a perfect starting point for modern synthetic biology.
6.3 CRISPR-Mediated Horizontal Gene Transfer (CRISPR-HGT)
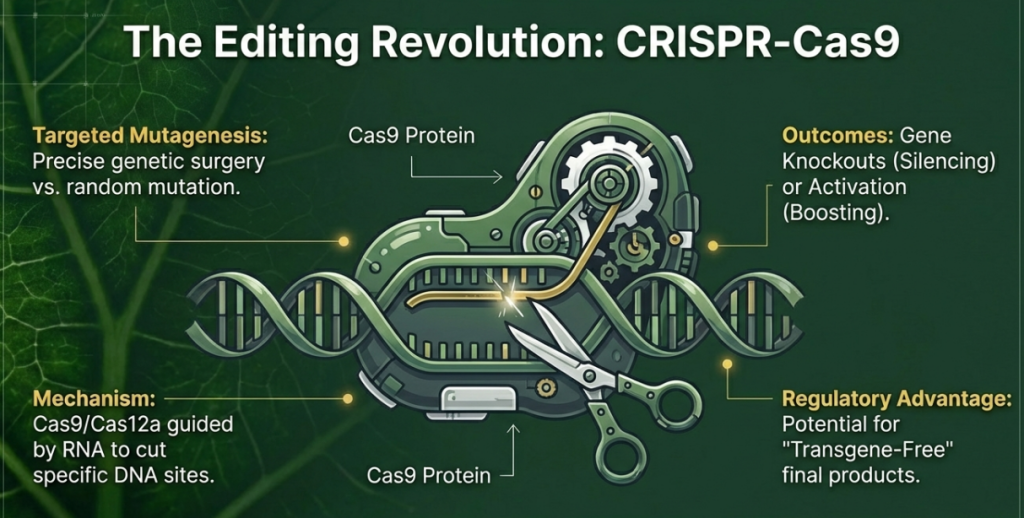
Synthetic biotechnology is moving toward an “evolution-guided design” philosophy. CRISPR-mediated Horizontal Gene Transfer (CRISPR-HGT) recreates the ancient processes through which plants naturally acquired adaptive microbial genes to survive extreme environments.
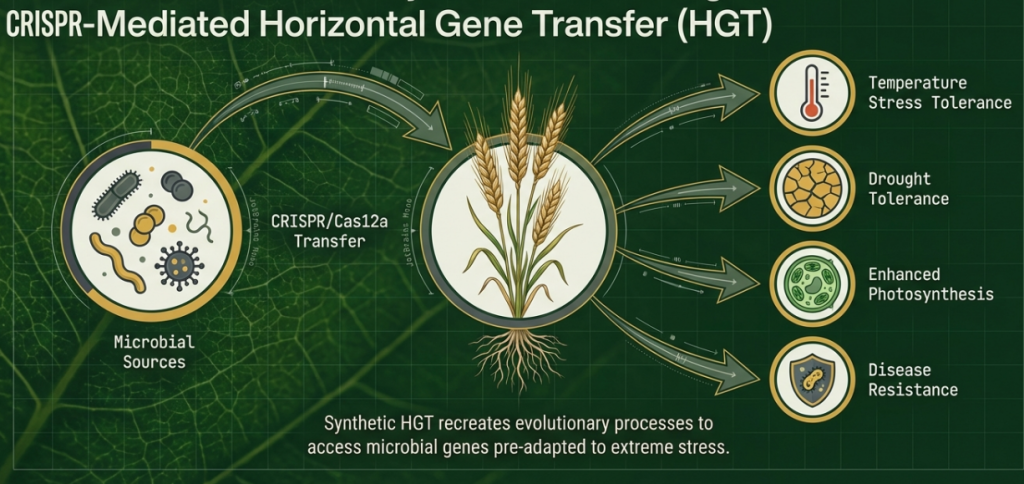
The Evolutionary Precedent: HGT is not merely a theoretical tool; it is a documented evolutionary shortcut. Wheat (Triticum aestivum) already harbors bacterial cold-shock protein (CSP) genes acquired through natural HGT, which contribute to drought tolerance and grain yield. Similarly, red algae (Pyropia haitanensis) possesses heat-stress tolerance genes derived from microbes. These examples provide the “evolutionary proof” that microbial modules are biologically compatible with plant systems.
Mechanisms and Synthetic Advantage:
• Cas12a and CasΦ: These tools enable the multiplexed insertion of large, operon-like gene clusters and multi-gene payloads, allowing for modular stress regulation.
• dCas-based Epigenome Editing: Facilitates reversible gene activation, allowing traits to be “switched on” only under specific environmental cues.
• Functional Superiority: Microbial modules, refined in extremophile environments, offer biological capacities—such as novel detoxification or redox buffering—that are often absent from or developmentally constrained within the endogenous plant gene pool.
The CRISPR-HGT Conceptual Pipeline:
1. Mining: Searching genomic and pan-genomic databases of “extremophile” microbes and wild plant relatives for high-value resilience traits.
2. Phylogenetic Analysis: Utilizing evolutionary history to confirm a gene’s success and functional compatibility as an HGT candidate.
3. Functional Validation: Testing candidate modules (e.g., β-1,3-glucanase or CSPs) in controlled environments to ensure they provide the intended resilience.
4. AI Optimization: Using deep learning tools such as DeepCRISPR, CRISPRon, and CRISPR-PLANT v2 to predict interaction dynamics and ensure stable integration with the plant’s existing metabolic networks.
This modular approach allows us to move away from older, static modification methods toward a dynamic system that anticipates environmental stress.
Comparing Natural Evolution vs. Deliberate Engineering
Understanding where CRISPR-HGT fits into the history of biotechnology requires a rigorous comparison with our existing methodologies.
| Feature | Natural HGT | Traditional Transgenics | CRISPR-HGT (Proposed) |
|---|---|---|---|
| Source of Genetic Material | Accidental cross-kingdom exchange. | Isolated single genes (often constitutive). | Modular microbial clusters (operon-like). |
| Precision/Control | None (Random). | Low (Fixed/Static expression). | High (Inducible, tissue-specific, and reversible). |
| Primary Limitation | Extremely rare and slow. | Static function; cannot respond to environmental variability. | Potential metabolic burden and the need for rigorous biosafety validation of non-plant proteins. |
While traditional transgenics often provide “always-on” traits that can exhaust a plant, CRISPR-HGT offers a multi-layered control framework for “smart” crops.
Breeding Paradigm Evolution
| Criteria | Traditional Phenotypic Selection | Marker-Assisted Selection (MAS) | CRISPR-HGT |
|---|---|---|---|
| Genetic Scope | Limited to variation within sexually compatible species. | Focused on major-effect loci and simple QTLs. | Modular; accesses preadapted microbial gene clusters across kingdoms. |
| Precision | Low; slow and highly environment-dependent. | Medium; precise for simple traits but lacks polygenic scalability. | High; precision insertion of multi-gene payloads with inducible control. |
| Environmental Adaptability | Passive; relies on slow, natural selection cycles. | Limited; QTL effects often lack transferability across environments. | Active; designed to anticipate and adapt to complex, overlapping stressors. |
Horizontal Gene Transfer (HGT) serves as a verified “evolutionary shortcut.” Genomic evidence demonstrates HGT’s role in foundational adaptations: wheat (Triticum aestivum) acquired bacterial cold-shock proteins (CSPs) that enhance drought tolerance and grain yield, while the liverwort Marchantia polymorpha acquired fungal-derived genes for drought adaptation. Most strikingly, the intertidal red alga Pyropia haitanensis acquired 286 HGT-derived genes, including modules for heat-stress tolerance such as sirohydrochlorin ferrochelatase. Synthetic HGT replicates this natural innovation through programmable design.
This roadmap follows two primary directions for CRISPR-HGT:
1. Ancestral Reintroduction: Identifying ancient HGT-derived genes already present in wild relatives and reintroducing them into elite cultivars to restore lost resilience.
2. Synthetic HGT: Testing novel microbial modules (from bacteria, fungi, or archaea) currently absent in plants and tuning them for high-performance stress responses.
3. Phase I: AI-Guided Microbial Gene Mining and Functional Prioritization
To reduce the prohibitive costs of empirical trials, we deploy machine learning to identify high-value genetic modules. This bioinformatic triage is the prerequisite for a system-level evaluation of gene clusters with validated co-expression. We prioritize genes that regulate core homeostasis and redox sensing rather than isolated, transient stress pathways.
The pipeline utilizes specialized tools: HGTector2 for identifying microbial origins, DeepGOPlus for functional prediction, and CRISPR-PLANT v2 for guide RNA optimization. We specifically target the expansion of metabolic and regulatory diversity, such as the acquisition of glycosyl hydrolase families (at least 23 families are known to be HGT-derived) to support cell wall diversification and defense evolution.
Mining Criteria
Before a microbial gene is considered for architectural integration, it must satisfy the following criteria:
• Metabolic Compatibility: High probability of functional integration into plant regulatory networks without pleiotropic risk.
• Functional Targets: Specific focus on cell wall diversification (glycosyl hydrolases), redox sensing, and ribosome protection modules.
• Modular Autonomy: Encapsulation within compact operon-like clusters that operate with minimal regulatory conflict with the host.
• Adaptive Relevance: Documented performance of the cluster in microbial extremophiles exposed to aridity, salinity, or oxidative stress.
4. Phase II: Architectural Integration via Advanced Cas Systems
The integration phase requires a strategic selection of molecular mediators based on the required trade-off between multiplexing capacity and payload size. We utilize diverse CRISPR systems to ensure complex, multi-gene modules are integrated with high fidelity.
For architectural deployment, we utilize:
• Cas12a: Ideal for the multiplexed insertion of operon-like clusters, mirroring natural HGT architectures.
• Compact CasΦ: Specifically selected for its ability to deliver large-scale, multi-gene payloads into complex plant genomes where larger nucleases face delivery constraints.
Inducible Regulation Toolkit
To minimize metabolic burden and avoid growth-defense trade-offs, microbial genes are managed via programmable control:
• RNA-Targeting Systems (Cas13/CasRx): Used for transient modulation of traits under acute stressors, ensuring the trait is active only when necessary.
• dCas-based Epigenome Editors: These allow for reversible activation or silencing. By using stress-inducible promoters, these tools ensure that HGT-derived genes remain silent under optimal growth and activate dynamically only when specific climate triggers (e.g., heat spikes) are detected.
5. Phase III: Multi-Trait Resilience and the Perennialization Strategy
Next-generation resilience requires stacking traits to combat overlapping stressors. A key strategic synergy is the integration of CRISPR-HGT with the development of perennial cropping systems. Perennialization offers deep root systems that enhance nitrogen (N) retention and promote soil carbon accumulation, essential for sustaining long-term productivity on marginal lands.
Next-Generation Crop Candidates
| Candidate Crop | Specific Advantages | Resilience Targets |
|---|---|---|
| Perennial Rice (PR) | Reduces labor by 58.1%; yield potential remains stable for 8 consecutive harvests (4 years). | Resistance to Rice Yellow Mottle Virus (derived from O. longistaminata). |
| Perennial Buckwheat (F. cymosum) | Indigenous growth in damp valleys; high waterlogging resistance. | Frost and heat tolerance in humid subtropical regions; water-tolerant food varieties. |
| Intermediate Wheatgrass (IWG) | Deep roots for carbon sequestration; high grain quality (Kernza®). | Frost and cold tolerance; negligible disease levels in established variety trials. |
6. Phase IV: Accelerating Genetic Gain via Integrated Genomic Selection (GS)
Genomic Selection (GS) is the essential final layer of our framework, providing the predictive power to estimate the breeding value of populations modified by CRISPR-HGT. The GS workflow utilizes a Training Population (TRN)—genotyped and phenotyped via high-throughput platforms—to calibrate models that predict the performance of a Testing Population (TST).
Machine Learning (ML) models, such as Random Forests and Deep Neural Networks, are mandatory for this roadmap. Unlike linear models like GBLUP (Genomic Best Linear Unbiased Prediction) which focus primarily on additive effects, ML approaches excel at capturing the non-additive epistatic interactions—the complex gene-to-gene communications—introduced by novel CRISPR-HGT modules.
Training the GS Model
We utilize the foundational statistical model:
y=Xb+Zu+e
• y: Phenotype vector.
• X/Z: Design matrices for fixed and random effects, respectively.
• b: Fixed effects (e.g., trial, block, or environmental covariates).
• u: Additive genomic breeding values (captured as GEBVs—Genomic Estimated Breeding Values).
• e: Residual error.
By calculating GEBVs, we accurately select individuals based on genomic potential rather than observable phenotypes, drastically shortening breeding cycles.
6. Navigating the Regulatory and Scientific Landscape
The successful deployment of GS and CRISPR-HGT requires a cohesive governance framework and a nuanced understanding of ecological trade-offs.
Comparative Regulatory Analysis
| Jurisdiction | Regulatory Approach | Framework Characteristics |
|---|---|---|
| European Union | Technique-based Oversight | Gene-edited crops are generally subjected to the same stringent and costly oversight as traditional GMOs. |
| United Kingdom | Product-based Framework | The “Precision Bred Plants” model focuses on whether a trait could arise through traditional breeding, prioritizing the final product’s risk profile over the technical method. |
Ecological Trade-offs and AI-Guided Discovery: The integration of microbial modules must account for “growth-defense trade-offs.” For example, Plant Growth-Promoting Bacteria (PGPB) produce auxins and ethylene that are beneficial at moderate levels but can trigger necrosis if overproduced via HGT. Furthermore, HGT-derived traits risk destabilizing native microbial communities. To mitigate this, AI-guided discovery tools like DeepGOPlus and HGTector2 are essential for predicting the compatibility of microbial modules with endogenous plant networks.
Crucially, the Genomic Selection frameworks discussed in Section 3 will serve as the primary tools for predicting the breeding values of these HGT-edited lines, ensuring that synthetic innovations are seamlessly integrated into field-scale breeding populations.
Biosafety and Ethics Checklist
• Immunogenicity Assessment: Ensuring introduced microbial proteins do not elicit allergic reactions.
• Metabolic Burden Analysis: Monitoring for fitness costs where plant energy is diverted from growth to maintaining synthetic modules.
• Eco-Evolutionary Interactions: Evaluating whether HGT traits disrupt beneficial symbioses, such as arbuscular mycorrhizal fungi relationships.
• Gene Flow Containment: Specific monitoring protocols for forestry species to manage risk over extended ecological timescales.
7. Conclusion: The Path Toward 2050
To meet the 2050 food security challenge, the breeding community must synthesize conventional selection, Genomic Selection, and synthetic biotechnology. We are transitioning from “incremental improvement” to a paradigm of “evolution-informed design,” where the genetic ingenuity of the microbial world is harnessed to provide crops with the resilience required for a volatile climate. This future demands transparent biosafety innovation, international regulatory harmonization, and a robust dialogue between scientists and stakeholders to ensure that modern biotechnology remains an ecologically sustainable and socially legitimate endeavor.
Top 3 Considerations for 2050 Global Food Security:
1. Multi-Trait Resilience: Stacking HGT-derived modules to provide “inducible” resistance to drought, heat, and pathogens simultaneously.
2. Perennial Cultivation: Reimagining major grains as multi-year crops to preserve soil health and drastically reduce farmer labor and financial inputs.
3. Predictive AI Design: Moving beyond trial-and-error by using AI to anticipate how cross-kingdom modules will function within the crop holobiont.
Image Summary

Questions/Answers
1. How does genomic selection accelerate breeding compared to traditional methods?
Genomic selection (GS) accelerates crop breeding by transitioning the process from empirical, phenotype-based selection to predictive, data-driven decision-making. Unlike traditional methods that are often slow, imprecise, and dependent on the environment, GS leverages genome-wide markers to predict the performance of individuals before they are even tested in the field.
The following mechanisms detail how GS accelerates the breeding process:
1. Shortening the Breeding Cycle
The most significant advantage of GS is its ability to drastically reduce the length of the breeding cycle, in some cases by up to 50%.
• Early Selection: In traditional breeding, breeders must wait for plants to reach maturity or undergo multi-environment trials to observe traits like yield or stress tolerance. GS allows for early selection based on genomic potential (Genomic Estimated Breeding Values or GEBVs) rather than waiting for observable phenotypes.
• Time Savings: Empirical evidence shows that GS can shorten breeding cycles by 2 to 4 years for staple crops such as wheat, maize, rice, and chickpea.
• Reduced Field Testing: Because the “testing population” is genotyped but not necessarily phenotyped, the time, cost, and environmental dependency of traditional selection cycles are significantly reduced.
2. Capturing Complex Polygenic Traits
Traditional methods, including earlier molecular tools like Marker-Assisted Selection (MAS), are primarily effective for traits governed by one or a few major genes.
• Cumulative Effects: GS captures the cumulative contribution of numerous small-effect loci across the entire genome. This makes it uniquely suited for improving complex traits such as yield, disease resistance, and drought tolerance, which are governed by many genes.
• Increased Genetic Gain: By accurately predicting these complex traits early, GS enhances annual genetic gain by 20–45% compared to conventional approaches. Some studies suggest potential increases in genetic gain of 30–50%.
3. Decoupling Selection from Phenotyping
Traditional breeding requires extensive and resource-intensive phenotyping in every generation.
• Data-Driven Advancement: GS uses a “training population” (which is both genotyped and phenotyped) to build predictive models. Once these models are calibrated, they can predict the performance of a much larger “testing population” using only genotypic data.
• Rapid-Cycle Breeding: This allows for rapid-cycle and recurrent selection, where multiple generations of crossing and selection can occur in a single year, especially when GS is integrated with technologies like speed breeding and doubled haploids.
4. Integration with Advanced Technologies
• AI and Machine Learning: Integrating GS with artificial intelligence and machine learning further enhances predictive accuracy for complex traits by 10–20% over conventional statistical methods.
• Decision Support Systems: AI-assisted digital decision support systems can synthesize GS outputs with climate and soil models to optimize field trials, potentially reducing breeding cycle times by an additional 25–35%.
2. Explain the role of CRISPR in creating climate-resilient crops.
CRISPR–Cas9 has emerged as a foundational technology in plant science, offering a precise, efficient, and scalable approach to creating climate-resilient crops. By enabling targeted modifications to the plant genome, CRISPR addresses the limitations of conventional breeding—such as slow breeding cycles and the “linkage drag” of unfavorable traits—to help crops withstand intensifying abiotic and biotic stresses driven by climate change.
Targeted Enhancement of Stress Tolerance
CRISPR allows for the direct manipulation of genes responsible for how plants perceive and respond to environmental fluctuations.
• Abiotic Stress Pathways: The technology is used to modify genes involved in stress perception, signal transduction, and transcriptional regulation. For example, it can fine-tune pathways for osmoprotectant biosynthesis, allowing plants to maintain productivity under drought, salinity, and extreme temperatures without the negative trade-offs often seen with broad gene overexpression.
• Precision and “Climate-Smart” Breeding: Researchers use CRISPR to eliminate unfavorable alleles in elite cultivars or fine-tune endogenous gene expression to enhance yield stability under stress. Integrating these traits into breeding programs supports “climate-smart” agriculture, intended to sustain food systems despite increasing environmental volatility.
Advanced CRISPR Applications for Resilience
The versatility of the CRISPR toolkit provides several specialized methods for engineering resilience:
• Multiplex Editing: CRISPR enables the simultaneous targeting of multiple loci. This is particularly valuable for developing durable resistance against evolving pathogens and for “stacking” multiple resilience traits (e.g., combined drought and salinity tolerance).
• Transcriptional Regulation: Tools like CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi) allow breeders to reversibly upregulate or repress endogenous stress-response genes without altering the underlying DNA sequence.
• Base and Prime Editing: These innovations allow for precise single-nucleotide substitutions or specific insertions/deletions, which reduce the risk of unintended mutations and genomic instability compared to traditional double-strand break methods.
Innovative Strategies: CRISPR-HGT and AI
Emerging frameworks are pushing CRISPR beyond simple allelic modification toward evolution-guided design:
• CRISPR-Mediated Horizontal Gene Transfer (CRISPR-HGT): This proposed framework synthetically reconstructs the evolutionary process by which plants acquired adaptive genes from microbes. Microbial genes, refined in extreme environments, offer a “naturally preadapted” resource for traits like detoxification, protein stabilization, and immune modulation.
• AI Integration: Artificial intelligence is being integrated with CRISPR to predict the best guide RNA (sgRNA) designs and to model how complex microbial modules will integrate into plant metabolic networks to maximize resilience.
Advantages Over Traditional Methods
• Speed: CRISPR significantly shortens the experimental timelines required to develop stable, homozygous phenotypes, potentially reducing time-to-market for improved cultivars by up to 40%.
• Preservation of Elite Backgrounds: The ability to edit endogenous genes directly in elite varieties avoids the need for years of backcrossing to remove undesirable traits introduced during conventional hybridization.
• Regulatory Potential: CRISPR-edited crops can be developed without the introduction of foreign DNA (using DNA-free ribonucleoproteins), which may simplify regulatory acceptance in some jurisdictions compared to transgenic GMOs.
Despite this potential, challenges remain regarding off-target effects, the efficiency of delivery in certain crop species, and inconsistent global regulatory frameworks. Nevertheless, CRISPR technologies are positioned as a cornerstone of modern precision breeding to ensure future global food security.
3. What are the benefits and challenges of developing perennial crops?
Developing perennial crops—which are planted once and harvested repeatedly over multiple years—offers a sustainable alternative to the annual cropping systems that currently provide 80% of the global food supply but require significant annual inputs.
Benefits of Perennial Crops
Perennial cultivation systems provide several environmental and economic advantages:
• Environmental Sustainability: Unlike annual crops, perennials provide permanent living cover for the soil. Their deep root systems enhance nitrogen retention, promote soil carbon accumulation, and prevent soil erosion and nutrient loss caused by heavy rainfall on bare soil.
• Resource Efficiency: Perennial crops typically have an extended photosynthetic period, which improves annual light capture and boosts overall productivity. They can significantly reduce the need for fertilizers, pesticides, energy, and labor inputs.
• Economic Gains for Farmers: Perennial rice, for example, has been shown to reduce labor by 58.1% and input costs by 49.2% in each regrowth cycle compared to annual varieties.
• Resilience and Stability: Perennial crops can provide more stable grain production in the face of rapid weather fluctuations that often hinder first-year grains. For instance, certain perennial buckwheat species exhibit high waterlogging tolerance and can survive in humid subtropical regions where other perennials cannot overwinter.
Challenges in Development
Despite their potential, the practical application of perennial crops faces significant hurdles:
• Biological Barriers: A primary challenge is hybrid sterility in interspecific hybrids (crosses between annual and perennial relatives), often caused by abnormal meiosis.
• Yield and Quality Gaps: Developed perennial varieties often lag behind their annual counterparts in terms of grain yield, seed size, and other agronomic traits. For example, intermediate wheatgrass yields significantly less than annual wheat.
• Unfavorable Traits: Wild perennial relatives often possess undesirable characteristics such as scrawny seeds, seed shattering, and strong bitterness that must be bred out.
• Weed Potential: There is a risk that perennial crops could become weeds themselves due to their ability to sprout from wintering roots or shattering seeds.
• Climate Adaptation: For perennial crops to winter successfully in high-altitude or high-latitude regions, they must possess traits for frost or soil freezing tolerance.
• Genetic Complexity: Perenniality is often a polygenic trait, meaning its successful introduction requires targeting multiple genetic loci rather than a single gene.
Overcoming Challenges with Technology
Advanced breeding strategies are being employed to address these bottlenecks:
• Genomic Selection (GS): This technique uses genome-wide markers to predict breeding values, capturing the cumulative contribution of many small-effect loci to improve complex traits like perenniality faster than traditional methods.
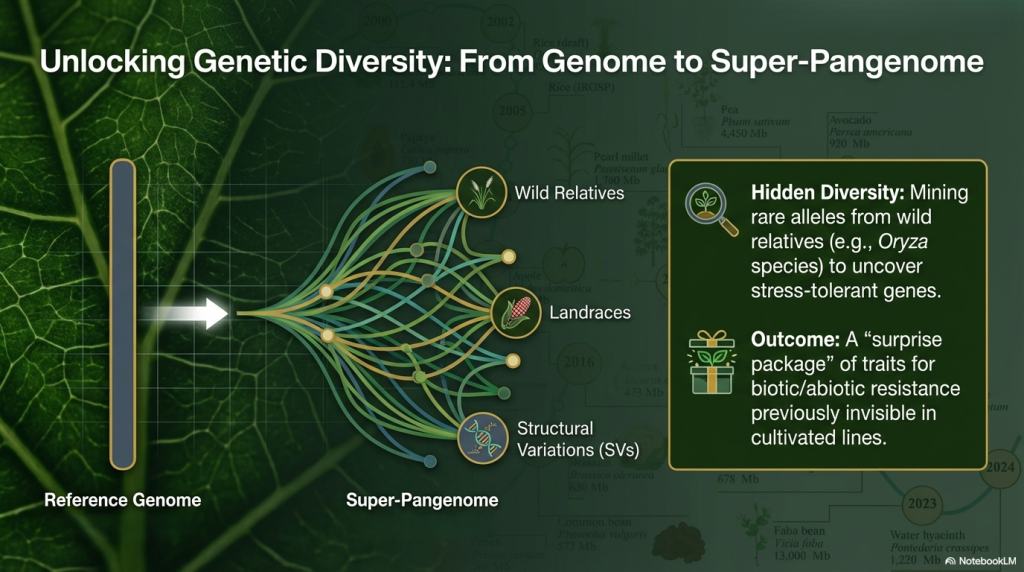
• Super-pangenomes: By integrating genomic data from both cultivated crops and their wild perennial relatives, super-pangenomes help identify structural variations and rare alleles lost during domestication, providing a “toolbox” for precision breeding.
• Embryo Rescue: This tissue culture technique is used to regenerate plants from interspecific hybrids that would otherwise degenerate due to incompatibility.
4. How is speed breeding integrated with genomic selection?
The integration of speed breeding (SB) with genomic selection (GS) represents a powerful synergy that transitions crop improvement into a predictive, rapid-cycle process. By combining the data-driven predictive power of GS with the rapid generation turnover of SB, breeders can significantly increase annual genetic gain and reduce the time required to develop improved cultivars.
The following mechanisms illustrate how these two technologies are integrated:
1. Rapid-Cycle and Recurrent Selection
The primary integration point is the transition from traditional selection to rapid-cycle genomic prediction pipelines.
• Early Selection: GS predicts the performance of individuals (Genomic Estimated Breeding Values or GEBVs) using genome-wide markers before they are tested in the field.
• Generation Turnover: While GS removes the need for extensive field testing in every cycle, SB provides the means to physically grow and cross these predicted high-performing individuals at an accelerated rate.
• Annual Gains: Together, they enable multiple generations of crossing and selection to occur within a single year—a process that normally takes years in traditional breeding.
2. Shortening the Breeding Cycle
The integration of these tools can shorten breeding cycles for staple crops (like wheat, rice, and maize) by 2 to 4 years.
• Predictive Advancement: In an integrated pipeline, a “testing population” is genotyped and their performance is predicted using models trained on a “training population”.
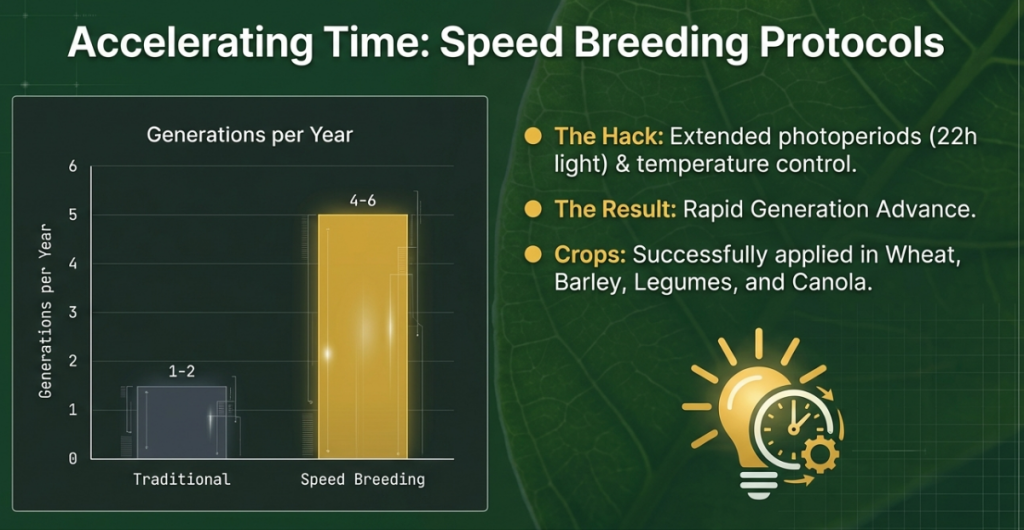
• Time Savings: Once superior candidates are identified via GS, SB protocols—which use regulated environmental conditions and prolonged photoperiods to achieve 4 to 6 generations per year—are used to rapidly advance these candidates toward homozygosity or for use as parents in the next cycle.
3. Rapid Fixation of Desirable Alleles
SB and GS are used together to rapidly fix desirable polygenic traits, such as yield or stress tolerance.
• Controlled Environments: SB uses optimized light intensity, temperature, and day length (e.g., 22-hour light regimes) to stimulate early flowering and seed set.
• Homozygous Phenotypes: This method allows breeders to obtain stable, homozygous phenotypes from predicted elite lines in less than a year.
4. Synergy with Other Technologies
The integrated framework often incorporates other advanced methods to further optimize outcomes:
• Doubled Haploids (DH): Integrating GS and SB with DH technology can reduce the time to develop homozygous lines from five generations to just two.
• AI and Machine Learning: Artificial intelligence is used to synthesize GS outputs with climate and soil models, further refining the selection process within the speed breeding pipeline.
• Genome Editing: CRISPR/Cas9 can be used to directly modify traits (e.g., knocking out late-flowering genes), and these edited plants are then grown under SB conditions to produce transgenic seeds and stable lines as quickly as possible.
5. Can CRISPRi and CRISPRa regulate stress responses without DNA changes?
Yes, CRISPRi (CRISPR interference) and CRISPRa (CRISPR activation) can regulate stress responses without making any permanent changes to the plant’s DNA sequence. These advanced CRISPR technologies focus on transcriptional regulation, allowing for the precise control of gene expression levels rather than altering the underlying genetic code.
Mechanism: Using Catalytically Inactive Cas9 (dCas9)
Unlike the standard CRISPR–Cas9 system that induces double-strand breaks to edit DNA, CRISPRi and CRISPRa utilize a catalytically inactive version of the Cas9 protein, often called “dead Cas9” or dCas9.
• dCas9 binding: This protein lacks nuclease activity (meaning it cannot cut DNA) but still retains its ability to bind to specific genomic loci when directed by a guide RNA.
• Reversible control: Because these systems do not break the DNA, the modifications they produce are reversible, enabling breeders to temporarily silence or activate genes in response to environmental cues.
Roles in Stress Response Regulation
These tools are particularly valuable for creating climate-resilient crops by managing how plants react to environmental volatility:
• Inducible Stress Regulation: dCas-based systems can be engineered for conditional regulation, allowing specific genes—such as those acquired from adaptive microbes—to remain silent under normal growth conditions and activate dynamically only when the plant encounters specific stress combinations.
• Fine-tuning Pathways: Instead of the broad, permanent overexpression of stress genes (which can often lead to negative trade-offs in yield or growth), CRISPRi and CRISPRa allow for the fine-tuning of stress-responsive pathways. This precision helps plants maintain productivity under fluctuating conditions like drought, salinity, or extreme temperatures.
• Studying Essential Genes: These systems are essential for research into regulatory networks and essential genes, where traditional permanent knockouts might be lethal to the plant.
• Epigenome Editing: CRISPRi/a can be integrated with epigenome editors to facilitate reversible gene activation or silencing through chemical modifications (like methylation) that affect how the DNA is read without changing the sequence itself.
By leveraging these “non-editing” CRISPR tools, scientists can move beyond simple allelic modification toward evolution-guided design, engineering crops that can anticipate and adapt to stress in real-time while sustaining yield.
6. How do AI and CRISPR work together in precision breeding?
The convergence of artificial intelligence (AI) and CRISPR–Cas9 is transforming precision breeding into a highly automated, predictive ecosystem that addresses the limitations of traditional, empirical methods. While CRISPR provides the molecular “scissors” to make precise genomic changes, AI serves as the “architect” that identifies optimal targets and predicts the outcomes of these modifications.
The synergy between these technologies manifests in the following key areas:
1. Optimized Design and Precision
AI and machine learning (ML) are critical for improving the technical efficiency of the CRISPR toolkit:
• sgRNA Design: AI models are used to design highly specific single-guide RNAs (sgRNAs), ensuring they bind to the correct target site with maximum efficiency.
• Off-Target Prediction: Tools such as DeepCRISPR and CRISPRon leverage deep learning to predict and minimize unintended “off-target” mutations, which is a significant biosafety concern in genome editing.
• Transcriptional Regulation: AI helps optimize the design of inducible promoters and CRISPR regulators (like CRISPRa or CRISPRi), allowing for reversible and context-specific gene control.
2. AI-Guided Gene Discovery
AI helps researchers “mine” massive genomic datasets to find the most effective traits to edit:
• Mining Super-Pangenomes: AI-driven analysis of super-pangenomes allows breeders to identify structural variations (SVs) and rare alleles in wild relatives that were lost during domestication. These rare variants serve as high-value targets for CRISPR-based reintroduction into elite cultivars.
• CRISPR-Mediated Horizontal Gene Transfer (CRISPR-HGT): In this proposed framework, AI mining tools identify naturally preadapted microbial genes (from bacteria or fungi) that confer resilience to extreme environments. CRISPR is then used to synthetically integrate these modules into crop genomes.
3. Predicting Functional Compatibility
Editing a single gene can have complex ripple effects throughout a plant’s biology. AI models address this complexity by:
• Metabolic Modeling: AI predicts how new genetic modules will integrate with the plant’s existing regulatory and metabolic networks.
• Anticipating Trade-offs: Machine learning models trained on multi-omics data can anticipate biological trade-offs, such as a gain in stress resistance that might inadvertently reduce grain yield.
• Phenotype Modeling: AI integrates genomic data with environmental metadata to simulate how an edited plant will perform under future climatic scenarios.
4. Accelerating the Breeding Pipeline
The integration of AI and CRISPR significantly reduces the experimental timelines required to develop field-ready crops:
• Time-to-Market: AI-assisted digital decision support systems can reduce the time required to develop improved cultivars by up to 40%.
• Stable Homozygosity: CRISPR can be used to knock out late-flowering genes, and when these edited plants are grown under speed breeding conditions, researchers can obtain stable, homozygous phenotypes in less than a year.
7. How does CRISPR-mediated HGT use microbial genes for resilience?
CRISPR-mediated horizontal gene transfer (CRISPR-HGT) is a programmable framework that synthetically recreates the evolutionary process by which plants historically acquired adaptive genes from microbes. This approach shifts genome editing from simple allelic modification to evolution-guided design, using microbial ingenuity as a modular toolkit to engineer crops that can anticipate and withstand stress.
The role of microbial genes in this framework includes the following:
1. Utilizing “Naturally Preadapted” Genetic Modules
Microbes evolving in extreme environments (such as those exposed to high radiation, desiccation, or salinity) have developed compact operons or enzyme clusters for multi-trait resilience. CRISPR-HGT treats these as preadapted genetic modules that can be transferred into plants to provide biochemical capacities—such as detoxification, protein stabilization, and immune modulation—that are otherwise absent from plant genomes.
2. AI-Guided Discovery and Integration
Artificial intelligence is used to “mine” genomic and pan-genomic databases to identify these beneficial microbial elements.
• Bioinformatic Mining: AI prioritizes microbial-like genes within stress-responsive pathways, such as those encoding antioxidant defenses or ion transport.
• Predicting Compatibility: AI models predict how these complex microbial modules will integrate with a plant’s existing regulatory and metabolic networks to ensure functional stability and minimize energetic burdens.
3. Precision Delivery of Complex Payloads
Advanced CRISPR tools are used to physically integrate these microbial modules into the crop genome:
• Cas12a: This nuclease enables the multiplexed insertion of operon-like clusters, mirroring how these genes are naturally organized in microbes.
• CasΦ: Due to its compact size, this nuclease supports the delivery of large multi-gene payloads even in complex genomes.
4. Modular and Inducible Stress Regulation
Instead of permanent, broad overexpression—which can lead to yield trade-offs—CRISPR-HGT allows for conditional regulation.
• dCas-based Tools: Tools like CRISPRa and CRISPRi (using catalytically inactive or “dead” Cas9/Cas12) can be used as epigenome editors to facilitate reversible gene activation.
• Dynamic Response: This allows microbial-derived resilience genes to remain silent under normal growth conditions and activate only when the plant encounters specific stress combinations, thereby preserving energy for yield under benign conditions.
5. Leveraging Existing HGT Events
A near-term strategy within this framework involves identifying ancient HGT-derived genes already present in wild relatives of crops. For example, bacterial cold-shock protein (CSP) genes found in wheat relatives have been shown to contribute to drought tolerance and improved photosynthesis. Researchers can use CRISPR to reintroduce these “natural” microbial footprints into elite cultivars to enhance their resilience.
8. How do AI and CRISPR work together in precision breeding?
Artificial intelligence (AI) and CRISPR work together in precision breeding to create a highly automated, predictive ecosystem that overcomes the limitations of traditional, empirical selection. While CRISPR provides the molecular tools to make precise genomic changes, AI acts as the “architect” that identifies optimal genetic targets and predicts the outcomes of those modifications.
The synergy between these technologies functions through the following mechanisms:
1. AI-Guided Target Discovery
AI is used to “mine” massive genomic datasets to identify the most effective traits for CRISPR-based editing.
• Super-Pangenome Mining: AI-driven analysis of super-pangenomes allows breeders to identify structural variations (SVs) and rare alleles in wild relatives that were lost during domestication, providing a catalog of functional targets for reintroduction via CRISPR.
• Microbial Gene Discovery: In frameworks like CRISPR-mediated Horizontal Gene Transfer (CRISPR-HGT), AI tools (such as DeepGOPlus and HGTector2) identify “naturally preadapted” microbial genes that confer resilience to extreme environments.
• Regulatory Elements: AI identifies conserved noncoding sequences (CNSs) and cis-regulatory elements (CREs) that can be targeted to fine-tune gene expression networks without altering the underlying coding sequence.
2. Optimized Design and Precision
AI tools significantly improve the technical efficiency and safety of the CRISPR toolkit.
• sgRNA Design: Deep learning models, such as DeepCRISPR and CRISPRon, are used to design highly specific single-guide RNAs (sgRNAs) with maximum on-target efficiency.
• Off-Target Prediction: AI algorithms predict and minimize unintended “off-target” mutations, which is a critical requirement for biosafety and regulatory acceptance.
• Modeling Epigenetics: AI is used to optimize the design of programmable epigenome editors (like dCas9-based systems) for reversible and environmentally responsive gene regulation.
3. Predicting Functional Compatibility
Editing a single gene often has complex ripple effects throughout a plant’s biology, which AI helps to manage.
• Network Integration: AI models predict how new genetic modules will integrate with a plant’s existing regulatory and metabolic networks, reducing the risk of biological conflict.
• Anticipating Trade-offs: Machine learning models trained on multi-omics data can anticipate biological trade-offs, such as ensuring that increased stress resistance does not inadvertently reduce grain yield.
• Phenotype Modeling: AI integrates genomic data with environmental metadata (satellite, soil, and weather data) to simulate how an edited plant will perform under future climatic scenarios.
4. Accelerating the Breeding Pipeline
The integration of AI and CRISPR drastically reduces the experimental timelines required to develop field-ready crops.
• Reduced Time-to-Market: AI-assisted digital decision support systems can reduce the time required to develop improved cultivars by up to 40%.
• Synergy with Speed Breeding: CRISPR can be used to knock out late-flowering genes, and when these edited plants are grown under speed breeding conditions, researchers can obtain stable, homozygous phenotypes in less than a year.
• Automated Decision-Making: AI-augmented genomic selection improves the prediction accuracy for complex traits by 10–20% over conventional methods, particularly in stress environments.
9. Can CRISPRi and CRISPRa regulate stress responses without DNA changes?
Yes, CRISPRi (CRISPR interference) and CRISPRa (CRISPR activation) can regulate stress responses without making permanent changes to a plant’s DNA sequence. These advanced tools focus on transcriptional regulation, allowing breeders to precisely control how much a gene is expressed rather than altering its underlying genetic code.
The following details the mechanisms and advantages of this approach:
The Mechanism: Dead Cas9 (dCas9)
Unlike standard CRISPR–Cas9, which creates double-strand breaks to edit DNA, CRISPRi and CRISPRa use a catalytically inactive version of the Cas9 protein, often called “dead Cas9” or dCas9.
• DNA Binding without Cutting: dCas9 lacks the nuclease activity required to cut DNA but retains its ability to bind to a specific genomic locus directed by a guide RNA.
• Reversibility: Because the DNA is not broken, the modifications are reversible, allowing for temporary silencing or activation of genes in response to environmental conditions.
Roles in Stress Response and Climate Resilience
These tools provide a highly programmable, multi-layered framework for managing environmental volatility:
• Inducible Regulation: dCas-based systems can be engineered for conditional regulation, where specific stress-response genes remain silent under normal conditions and activate dynamically only when the plant encounters specific combinations of stress, such as drought and heat.
• Fine-Tuning Pathways: Instead of permanent, broad overexpression of stress genes (which often leads to negative trade-offs in yield or growth), CRISPRi and CRISPRa allow for the fine-tuning of endogenous gene networks.
• Epigenome Editing: These systems can function as programmable epigenome editors, facilitating reversible gene regulation through modifications that affect how the DNA is read without changing the sequence itself.
• Studying Essential Genes: CRISPRi and CRISPRa are particularly valuable for researching regulatory networks and essential genes where permanent “knockouts” might be lethal to the plant.
By using these “non-editing” CRISPR tools, researchers can move toward evolution-guided design, creating crops that can anticipate and adapt to shifting climates in real-time while maintaining productivity.
10. How do plants decide which genes to activate during stress?
Plants decide which genes to activate during stress through a complex series of biological events that transition from environmental sensing to genetic execution. This process is governed by multi-layered regulatory networks that integrate various internal and external signals.
Perception and Signaling
The decision-making process begins with stress perception, where the plant detects environmental fluctuations such as drought, salinity, or extreme temperatures. Once a stressor is perceived, the plant initiates signal transduction pathways that carry this information from the cell surface to the nucleus. These signaling pathways are essential for coordinating the plant’s overall physiological response to environmental volatility.
Transcriptional Regulation
The primary “switches” in this process are transcription factors, which are specialized proteins that bind to specific DNA sequences to turn genes on or off.
• Cis-regulatory elements (CREs): These are non-coding DNA sequences, often containing stress-responsive motifs, located near the start of genes that serve as landing sites for transcription factors.
• Gene Expression Networks: By binding to these motifs, transcription factors can activate entire networks of genes involved in osmoprotectant biosynthesis, detoxification, and protein stabilization.
Epigenetic and Functional Layers
The plant’s decision is also influenced by its epigenetic state, which includes chemical modifications like DNA methylation that affect how easily a gene can be read without changing the DNA sequence itself. Conserved noncoding sequences (CNSs) across different species act as universal or species-specific regulatory elements to maintain core functions under pressure. Furthermore, research into multi-omics—including transcriptomics and metabolomics—shows that these decisions occur at multiple functional levels to ensure the plant maintains core homeostasis and immune modulation.
Evolutionary Influence
Interestingly, the “logic” behind some of these stress responses was shaped by horizontal gene transfer (HGT), where plants historically acquired adaptive genetic modules from microbes. These acquired microbial genes often regulate signaling and detoxification rather than just isolated stress pathways, providing plants with a “preadapted” toolkit for multi-trait resilience.
11. What is the difference between CRISPRi and traditional gene silencing?
The primary difference between CRISPR interference (CRISPRi) and traditional gene silencing methods, such as RNA interference (RNAi), lies in the molecular level at which they operate and the precision of the control they offer.
1. Level of Regulation: Transcriptional vs. Post-transcriptional
• CRISPRi: Operates at the transcriptional level. It uses a catalytically inactive or “dead Cas9” (dCas9) protein that lacks nuclease activity and cannot cut DNA. Instead, the dCas9 is directed by a guide RNA to bind to a specific genomic locus, physically blocking the transcription machinery (like RNA polymerase) from reading the gene.
• Traditional Silencing (RNAi): Operates at the post-transcriptional level. It targets and degrades messenger RNA (mRNA) after it has already been transcribed from the DNA, preventing it from being translated into a protein.
2. Reversibility and Dynamics
• CRISPRi: Provides reversible repression. Because it does not alter the underlying DNA sequence or degrade the transcripts permanently, it allows for temporary or conditional regulation. This is particularly useful for studying essential genes that, if permanently knocked out, would be lethal to the plant.
• Traditional Silencing: While also capable of varying levels of suppression, traditional methods are often described as having a lower level of control compared to CRISPR-based tools. CRISPRi can be engineered to respond dynamically to environmental cues, such as activating only when a plant encounters specific stresses like drought or heat.
3. Precision and Scalability
• CRISPRi: Is highly programmable and scalable. It is part of an advanced “CRISPR toolkit” that allows for multiplexing, where multiple genes can be silenced or regulated simultaneously with high specificity.
• Traditional Silencing: Methods like RNAi have historically been more prone to “off-target effects,” where genes with similar sequences are unintentionally silenced. CRISPRi, when integrated with AI-guided design, can significantly reduce these risks by predicting and avoiding unintended genomic interactions.
4. Integration with Epigenetics
• CRISPRi: Can function as a programmable epigenome editor. It can be fused with enzymes that chemically modify the DNA (such as through methylation) to silence genes without changing their sequence. This allows for a more complex layer of regulation that traditional gene silencing methods cannot easily achieve.
12. Could CRISPRa be used to make crops more nutritious?
Yes, CRISPR activation (CRISPRa) can be used to make crops more nutritious. This technology allows for the precise, controlled upregulation of endogenous genes without altering the underlying DNA sequence.
The sources highlight several ways CRISPRa and related genome-editing tools contribute to nutritional enhancement:
• Upregulation of Beneficial Pathways: CRISPRa enables gain-of-function studies by activating specific gene networks responsible for producing essential nutrients. This precision is particularly valuable for improving traits like micronutrient accumulation, protein composition, and starch biosynthesis.
• Specific Nutritional Targets: Researchers use CRISPR-based technologies to target noncoding regions to control gene expression. A notable example is the engineering of noncoding regions to enhance vitamin C biosynthesis in crops like kiwifruit.
• Preserving Elite Backgrounds: Because CRISPRa can modify the expression of a plant’s own genes, it allows for the direct improvement of elite cultivars’ nutritional value without introducing foreign DNA or requiring years of backcrossing to remove “linkage drag” from wild relatives.
• Broad Applications in Biofortification: The technology is part of a broader shift in precision breeding aimed at creating nutritionally enhanced, high-yield, and resilient crops to address global food security and “hidden hunger”.
By fine-tuning metabolic pathways rather than using broad gene overexpression, CRISPRa offers a sophisticated method for developing crops with improved flavor, nutrient profiles, and shelf life.
13. Can CRISPRi target multiple genes simultaneously for stress resistance?
Yes, CRISPRi (CRISPR interference) can target multiple genes simultaneously to enhance stress resistance. This capability, known as multiplex editing, is one of the primary advantages of the CRISPR toolkit in plant science.
The following details explain how CRISPRi and multiplexing work together to improve crop resilience:
Multiplexing for Trait Stacking
CRISPR-based strategies allow for the simultaneous modification of multiple genetic loci. In the context of climate resilience, this allows breeders to “stack” various traits—such as combined drought and salinity tolerance—within a single plant. This approach is more effective than traditional breeding, which often addresses stressors in isolation.
Reversible and Inducible Regulation
Because CRISPRi uses a catalytically inactive or “dead” Cas9 (dCas9) to repress gene expression without changing the underlying DNA sequence, its multiplexed applications offer high levels of control.
• Inducible Responses: Multiplexed CRISPRi can be engineered to remain silent under normal conditions and activate or repress multiple target genes dynamically only when the plant encounters specific combinations of stressors.
• Coordinated Repression: Researchers can use this technology to precisely repress multiple negative regulators of stress responses simultaneously, thereby enhancing the plant’s overall defense capacity.
Parallel Discovery in Advanced Frameworks
Newer breeding frameworks, such as CRISPR-mediated Horizontal Gene Transfer (CRISPR-HGT), rely on multiplexing to integrate complex microbial genetic modules into plants.
• Evaluating Modules: CRISPR-HGT integrates multiplex CRISPR delivery to evaluate multiple microbial-derived resilience genes in parallel.
• Multi-Gene Payloads: Advanced nucleases like Cas12a and CasΦ are specifically highlighted for their ability to support the delivery and integration of large multi-gene payloads or operon-like clusters, mirroring how genes are organized in resilient microbes.
By leveraging multiplexed CRISPRi, scientists can create “climate-ready” crops that possess durable, multi-stress resilience against evolving pathogens and volatile environmental conditions.
14. Can CRISPRa be used to biofortify staple crops with vitamins?
Yes, CRISPRa (CRISPR activation) can be used to biofortify staple crops with vitamins and other essential nutrients by enabling the precise, controlled upregulation of endogenous genes without altering the underlying DNA sequence.
The sources provide the following details on how this technology supports biofortification:
• Mechanism of Action: CRISPRa uses a catalytically inactive version of the Cas9 protein (dCas9) that binds to specific genomic loci but does not cut the DNA. This allows breeders to activate or increase the expression of a plant’s own genes that are responsible for nutrient production.
• Targeting Nutritional Pathways: These “gain-of-function” studies allow researchers to turn on or enhance specific metabolic networks. This precision is used to improve traits such as micronutrient accumulation, starch biosynthesis, and protein composition in crops.
• Specific Examples:
◦ Researchers are using CRISPR-based tools to target noncoding regions to control gene expression, with a notable example being the engineering of noncoding regions to enhance vitamin C biosynthesis in crops like kiwifruit.
◦ While some biofortified staples like “Golden Rice” (enriched with pro-vitamin A) were originally created using traditional transgenic methods, CRISPRa offers a way to achieve similar results by modifying the expression of a plant’s existing genes rather than introducing foreign DNA.
• Advantages for Staple Crops: CRISPRa allows for the direct improvement of elite cultivars, preserving their high yield and other desirable traits while enhancing their nutritional value. This avoids the lengthy process of backcrossing typically required to remove “linkage drag” introduced when breeding with wild relatives.
By fine-tuning metabolic pathways rather than using broad gene overexpression, CRISPRa serves as a cornerstone of modern precision breeding aimed at addressing global food and nutritional security, specifically to combat “hidden hunger” in regions that rely heavily on staple crops.
15. Can CRISPRi target disease resistance and drought tolerance simultaneously?
Yes, CRISPRi can target disease resistance and drought tolerance simultaneously through a process known as multiplex editing, which allows for the simultaneous modification of multiple genetic loci within a single plant. This capability enables breeders to “stack” various resilience traits, effectively addressing both biotic (disease) and abiotic (drought) stresses at the same time.
The integration of these traits is achieved through several advanced CRISPR frameworks:
1. Multiplex Delivery and Trait Stacking
The CRISPR toolkit is specifically designed to handle multiple guide RNAs (sgRNAs), which can direct the catalytically inactive Cas9 (dCas9) to different genomic locations. This approach is significantly more efficient than traditional breeding, which often struggles with “linkage drag” or the difficulty of addressing complex, overlapping stressors in isolation.
2. CRISPR-Mediated Horizontal Gene Transfer (CRISPR-HGT)
A cutting-edge framework called CRISPR-HGT is being developed specifically to evaluation and engineer multi-trait resilience.
• Parallel Evaluation: CRISPR-HGT uses multiplex CRISPR delivery to evaluate various microbial-derived genes in parallel, seeking to reintroduce “naturally preadapted” modules for stacked resistance to both abiotic and biotic stressors.
• Complex Payloads: Advanced nucleases like Cas12a and CasΦ support the delivery of large, multi-gene payloads, mirroring how resilient microbes organize their genes into clusters to manage diverse environmental pressures.
3. Dynamic and Inducible Regulation
Unlike permanent gene knockouts, CRISPRi allows for reversible and modular regulation.
• Conditional Responses: Multiplexed CRISPRi can be engineered so that the genes for drought tolerance and disease resistance remain silent under normal conditions, activating dynamically only when the plant encounters those specific stressors.
• Preserving Yield: This precision helps minimize the “energetic or regulatory burdens” (yield trade-offs) often associated with the permanent overexpression of stress genes.
4. Integration with Predictive Tools
To ensure that stacking these traits does not cause biological conflict, researchers use artificial intelligence (AI) to model how these diverse modules will integrate into the plant’s existing regulatory and metabolic networks. This predictive approach helps identify the specific combination of edits most likely to provide stable, multi-stress resilience across different environments.
16. Can CRISPRi be used to reduce toxins in bitter crops?
Yes, CRISPRi (CRISPR interference) can be used to reduce toxins in bitter crops by enabling the reversible and precise repression of genes responsible for toxin biosynthesis. Unlike traditional knockout methods that permanently delete genes, CRISPRi allows breeders to silence specific pathways without making permanent changes to the plant’s DNA sequence.
The following points detail how CRISPRi and related technologies address toxins in bitter crops:
1. Mechanism of Toxin Reduction
CRISPRi utilizes a catalytically inactive “dead” Cas9 (dCas9) protein that binds to a specific genomic locus but does not cut the DNA. By binding to the promoter or coding region of a target gene, it physically blocks the transcription machinery, effectively “turning off” or reducing the production of specific compounds. This is particularly useful for targeting the biosynthetic pathways of:
• Cucurbitacins: These are the compounds primarily responsible for bitterness in most cucurbits, such as wild bottle gourds.
• Quinolizidine Alkaloids: Found in “bitter lupins,” these alkaloids are toxic to humans and affect the nervous, circulatory, and digestive systems.
• Steroidal Glycoalkaloids: Toxic compounds like α-solanine and α-chaconine are found in crops like tomatoes and potatoes.
2. Advantages of Reversible Repression
Many toxic or bitter compounds also serve as natural defense mechanisms for the plant against pests and pathogens. The reversible nature of CRISPRi is advantageous because:
• Fine-Tuning: It allows for the fine-tuning of gene expression rather than a total, permanent loss of function, which can sometimes reduce a plant’s overall fitness.
• Inducible Control: CRISPRi can be engineered to be inducible, meaning the toxin production could potentially be silenced in the edible parts of the plant (like the fruit or tuber) while remaining active in other tissues for defense.
3. Precedents in Genome Editing
While specific large-scale commercial applications of CRISPRi for de-bittering are still emerging, other genome-editing tools have already demonstrated success in this area. For example, Zinc Finger Nucleases (ZFNs) have been used in tomatoes to reduce cholesterol and toxic steroidal glycoalkaloids successfully. Additionally, researchers have identified specific genetic markers and structural variations, such as deletions in the ClBt gene, that explain the loss of bitterness during the domestication of crops like watermelon, providing a clear roadmap for CRISPR-based interventions.
4. Precision and Speed
The integration of CRISPR-based tools into breeding programs significantly accelerates the process of removing unfavorable traits compared to traditional backcrossing. This allows for the direct improvement of elite cultivars, preserving their high yield and quality while specifically addressing the toxin or bitterness issue.
17. How do doubled haploids further accelerate speed breeding?
Doubled haploids (DH) further accelerate speed breeding by providing a rapid method to achieve complete genetic homozygosity, effectively bypassing the multiple generations of selfing typically required in traditional breeding.
The integration of DH technology contributes to the breeding process in the following ways:
• Rapid Generation of Homozygous Lines: DH production allows for the creation of 100% homozygous inbred lines in just one generation. In contrast, traditional methods can take five or more generations to reach a similar level of genetic uniformity.
• Drastic Reduction in Cycle Time: By producing homogeneous lines so quickly, DH technology can reduce the breeding cycle from five generations down to just two. This speed is particularly valuable for creating new varieties and homozygous parental lines for F1 hybrid production.
• Predictable Trait Expression: Because DH plants are genetically homogeneous, they allow for more predictable and stable trait expression early in the breeding process. This precision helps breeders evaluate candidates more effectively for complex traits like yield and disease resistance.
• Enhanced Genetic Gain: When integrated into advanced frameworks like Nested Association Mapping (DH NAM), DH technology enables the development of homozygous lines in a short span of time, significantly increasing annual genetic gain.
• Complementary Synergy: DH production is an “accelerated breeding strategy” that works in tandem with speed breeding and genomic selection to reduce generation time and rapidly fix desirable alleles within a population.
While DH technology has biological limitations and can be genotype-dependent, its ability to save years of time makes it a foundational tool in modern precision breeding ecosystems.
18. How do doubled haploids simplify F1 hybrid production?
Doubled haploids (DH) simplify the production of F1 hybrids primarily by drastically accelerating and streamlining the creation of the 100% homozygous inbred lines required as parent material.
The following mechanisms explain how DH technology facilitates this process:
1. Rapid Achievement of Absolute Homozygosity
Traditional F1 hybrid production requires parental lines to be completely homozygous to ensure that the resulting offspring (the F1 generation) are genetically uniform.
• Time Savings: Conventional methods for developing these inbred lines typically require five to eight generations of selfing and selection. DH technology achieves the same result in just one generation by doubling the chromosomes of haploid plants.
• Cycle Reduction: This efficiency can reduce the overall breeding cycle from five generations down to just two, allowing breeders to move to large-scale hybrid production much faster.
2. Enhanced Predictability and Uniformity
Because DH lines are 100% homozygous, they produce highly predictable and stable trait expression.
• Genetic Uniformity: Crossing two genetically distinct, completely homozygous DH lines results in an F1 hybrid population that is 100% uniform (heterozygous at all loci but genetically identical across all individuals).
• Trait Selection: In crops like cabbage, DH is used to develop inbred lines that are homozygous for essential horticultural traits, disease resistance, and specific self-incompatibility genes, which are necessary for controlled hybrid crossing.
3. Streamlined Commercial Scalability
DH technology provides an efficient pathway for maintaining and preserving elite parental lines for commercial use.
• Parental Line Maintenance: DH lines serve as an excellent method for preserving the exact genetic identity of parental components, which is critical for the long-term consistent production of a specific commercial hybrid.
• Overcoming Inbreeding Depression: While many cross-pollinated species experience a decline in vigor during the inbreeding process, DH technology allows breeders to skip the intermediate generations of declining vigor and move directly to the homozygous state for immediate hybrid testing and restoration of vigor (heterosis).
4. Synergy with Other Advanced Tools
The use of DH is often integrated with other modern techniques to further simplify hybrid development:
• CMS Systems: DH parental lines are frequently combined with cytoplasmic male sterility (CMS) systems to enable large-scale, cost-effective hybrid seed production by eliminating the need for manual emasculation.
• Genetic Research: DH lines are valuable resources for QTL mapping and basic genetic research, helping breeders identify the best possible parental combinations to maximize hybrid vigor.
19. Which crops currently benefit most from DH technology?
Based on the sources, maize is identified as having one of the most common and well-established doubled haploid (DH) systems. While efficient and reproducible DH protocols are currently available for only about 30 crop species, the following crops are noted as benefiting significantly from the technology:
Major Field Crops and Cereals
DH technology is widely utilized in major staple crops to reduce breeding cycles from five generations down to just two. Key beneficiaries include:
• Maize: The maize DH system is particularly prominent and frequently uses markers like the R1-NJ color marker for identification.
• Inbreeding Cereals: This group includes wheat, rice, and barley.
• Other Large-Scale Crops: Reproducible protocols are established for potato, rapeseed (canola), and sugar beet.
Cruciferous (Brassica) Vegetables
DH technology, often achieved through anther or microspore culture, is a “cornerstone” for creating 100% homozygous inbred lines for these crops in a single generation.
• Cauliflower: DH lines are used to preserve parental lines and enhance traits like disease resistance.
• Cabbage: The technology is used to develop homozygous lines more quickly, allowing for more predictable trait expression.
• Broccoli: It facilitates the quick production of genetically homogeneous lines.
• Radish: DH produced by microspores is used as an appealing tool for saving time and creating new hybrids.
Strategic Applications
DH technology is particularly valuable when integrated into advanced breeding frameworks:
• Hybrid Seed Production: It is highly advantageous for the commercial production of F1 hybrid seeds by providing stable, homozygous parental lines.
• Nested Association Mapping (DH NAM): In crops with well-established protocols, DH is used to advance half-sib populations rapidly, contributing to enhanced annual genetic gain.
20. Are there limits to using DH in certain plant species?
Yes, there are significant limits to using doubled haploid (DH) technology, primarily due to its genotype dependency and the limited number of species for which efficient protocols currently exist.
The following points detail the specific limitations identified in the sources:
1. Limited Species Applicability
While DH technology is a powerful tool for achieving complete homozygosity in a single generation, it is not universally applicable.
• Narrow Range: Currently, efficient and reproducible DH protocols are available for only about 30 crop species.
• Established Examples: The technology is most commonly used in inbreeding cereals like wheat, rice, and barley, as well as in maize, potato, rapeseed, and sugar beet.
2. Genotype Dependency
Even within a species where DH protocols exist, the success of the process is often highly dependent on the specific genetic background of the plant being used.
• Induction Rates: Different crop species and varieties show varying rates of haploid induction, meaning some genotypes respond well to the process while others do not. For example, in tropical maize, the in vivo haploid induction rate is influenced by both the source germplasm and the season [138 (ref)].
• Tissue Culture Requirements: The process often requires specialized tissue culture techniques, such as anther or microspore culture, which can be difficult to optimize for every variety within a species.
3. Biological and Technical Barriers
The DH process involves complex biological manipulations that present several hurdles:
• Meiotic Abnormalities: In some cases, such as interspecific hybrids (crosses between different species), abnormal meiosis can lead to hybrid sterility, making it difficult to produce viable haploids for doubling.
• Chromosome Doubling: For many species, the doubling of the haploid genome requires chemical agents like colchicine, which can be toxic and may cause unintended genetic changes or poor plant survival [226, 280 (ref)].
• Regeneration Challenges: For many vegetable crops, such as cauliflower or broccoli, regenerating whole plants from tissue culture (the necessary step after induction) remains a major technical challenge.
4. Impact on Genetic Diversity
There are also strategic limitations concerning the long-term health of a breeding program:
• Loss of Variation: Breeders using DH systems may unintentionally practice selection for certain loci during the induction process, which can limit genetic variation in the resulting breeding populations.
• Reduced Opportunities for Recombination: Because DH production bypasses several generations of traditional selfing, it provides fewer opportunities for recombination to break unfavorable genetic linkages compared to methods like Single Seed Descent (SSD).
In summary, while DH technology can reduce breeding cycles from five generations down to just two, its practical use is constrained by biological complexity, a requirement for high-tech infrastructure, and a strong dependence on the specific genotype of the starting material.
21. How do DH technology and CMS systems work together?
Doubled haploid (DH) technology and cytoplasmic male sterility (CMS) systems work together as complementary tools to accelerate and automate the large-scale production of F1 hybrid seeds. While DH technology rapidly creates the homozygous parental material needed for hybrids, CMS provides the biological mechanism to cross those parents efficiently without manual intervention.
The synergy between these two technologies functions through the following mechanisms:
1. Rapid Development of the “Three-Line” System
Large-scale hybrid seed production often relies on a “three-line system,” which requires three distinct parental types. Integrating DH technology drastically shortens the development of these lines:
• A-Line (Male Sterile): The female parent that carries sterile cytoplasm and recessive nuclear genes.
• B-Line (Maintainer): Genetically identical to the A-line but carries fertile cytoplasm to produce pollen for maintaining the A-line.
• R-Line (Restorer): The male parent that carries dominant restorer genes to ensure the final F1 hybrid is fertile for the farmer.
DH technology reduces the time required to develop these lines from five to eight generations of traditional selfing down to just one or two generations.
2. Ensuring Absolute Homozygosity and Uniformity
For a hybrid to be commercially viable, the parental lines must be genetically uniform.
• 100% Homozygosity: DH production ensures that the A, B, and R lines are completely homozygous in a single step.
• Predictable Performance: Crossing two completely homozygous DH lines results in an F1 hybrid population that is 100% uniform, possessing the exact same genetic identity across every individual.
3. Commercial Scalability and Cost Efficiency
The combination of DH and CMS addresses the primary economic and technical bottlenecks in hybrid breeding:
• Elimination of Manual Emasculation: CMS makes the female parent sterile, meaning breeders do not have to manually remove anthers to prevent self-pollination.
• Exclusivity and Maintenance: DH lines serve as an excellent method for preserving the exact genetic identity of elite parental components, which is critical for consistent long-term production and protecting a breeder’s investment.
• Improved Seeding Performance: The use of sturdy CMS lines and DH-derived parents improves the overall seeding performance and reliability of the hybrid production process.
4. Crop-Specific Beneficiaries
Several major crops currently utilize the combined power of CMS and DH technology to advance their breeding programs:
• Brassica Vegetables: This includes cauliflower, cabbage, broccoli, and radish, where CMS systems (like Ogura) are used alongside DH to produce uniform, high-yielding hybrids.
• Maize: Maize hybrid systems are well-established and frequently integrate DH technology to fix target genes and fix desirable alleles quickly.
• Onions and Sugar Beet: CMS is essential for creating hybrids in these crops, and DH technology is used to develop the required inbred lines in a fraction of the traditional time.
22. Can doubled haploids help in breeding self-incompatible vegetables?
Yes, doubled haploids (DH) are a foundational tool for breeding self-incompatible (SI) vegetables, particularly those in the Brassicaceae family such as cauliflower, cabbage, broccoli, and radish.
DH technology helps overcome the biological and logistical challenges of breeding these crops in several key ways:
1. Rapid Achievement of Absolute Homozygosity
Traditional inbreeding in self-incompatible crops is difficult because plants naturally reject their own pollen. Breeders often have to use labor-intensive methods like bud pollination over 6–8 generations to produce inbred lines.
• Time Savings: DH technology bypasses this process, creating 100% homozygous inbred lines in just a single generation.
• Fixing Traits: It allows for the rapid and permanent fixation of desirable alleles, such as those for disease resistance or specific horticultural qualities.
2. Streamlining F1 Hybrid Production
The commercial success of many vegetables relies on F1 hybrids, which require two genetically distinct, completely homozygous parental lines.
• Fixing S-Alleles: For a self-incompatibility system to work effectively in hybrid seed production, parental lines must be homozygous for a single SI gene (S-allele). DH technology is used to develop these stable, homozygous parental components more quickly than traditional methods.
• Genetic Uniformity: Because DH-derived parents are genetically homogeneous, the resulting F1 hybrids exhibit high levels of uniformity and predictable trait expression, which are critical for commercial harvesting and marketing.
3. Overcoming Inbreeding Depression
Many cross-pollinated and self-incompatible species experience a significant decline in vigor (inbreeding depression) during the many cycles of selfing required by traditional breeding. DH technology allows breeders to move immediately to a homozygous state, where they can evaluate the line’s potential and restore vigor through hybridization (heterosis) without spending years on intermediate, low-vigor generations.
4. Application in Specific SI Vegetables
• Cauliflower: DH parental lines are increasingly used to address challenges in commercial seed production, helping to reduce the presence of “sibs” (unintended selfed offspring) in hybrid batches.
• Cabbage: It is a “cornerstone” for creating the inbred lines homozygous for both horticultural traits and the SI genes necessary for controlled crossing.
• Radish: DH produced by microspore culture is considered an “appealing tool” for saving time and developing new, genetically uniform hybrids.
Challenges and Limitations
While powerful, the use of DH in these vegetables is often genotype-dependent, meaning the process (such as anther or microspore culture) may work well for some varieties but fail for others. Additionally, regenerating whole plants from the tissue culture stage remains a technical hurdle for some species.
23. What are the biological limits of using DH technology?
Doubled haploid (DH) technology is a powerful tool for achieving complete genetic homozygosity in a single generation, but its application is constrained by several genotypic and biological limitations. These hurdles range from the technical complexity of the induction process to the broader impact on the genetic health of breeding populations.
The primary biological limits of DH technology include:
1. Genotype Dependency
The success of DH production is highly dependent on the specific genetic background of the plant species or variety being used. Different genotypes exhibit varying rates of haploid induction, meaning some respond efficiently to induction protocols while others remain recalcitrant. For example, in tropical maize, induction rates are significantly influenced by both the source germplasm and the season.
2. Limited Species Applicability
Despite its advantages, efficient and reproducible DH protocols have been established for only about 30 crop species. While common in inbreeding cereals like wheat, rice, and barley, as well as in maize, potato, rapeseed, and sugar beet, many other crops lack the biological responsiveness required for routine DH applications.
3. Biological and Technical Hurdles
The process of creating doubled haploids involves complex biological manipulations that present significant barriers:
• Tissue Culture Requirements: The technology often relies on specialized in vitro culturing techniques, such as anther or microspore culture, which require sophisticated equipment and highly specific environmental controls.
• Chromosome Doubling: The conversion of haploids into fertile diploids requires chromosome doubling, which typically necessitates the use of chemical agents like colchicine. These agents can be toxic, and the procedure itself can lead to unintended genetic abnormalities or poor plant survival.
• Meiotic Abnormalities: In certain breeding scenarios, such as interspecific hybridization (crossing different species), abnormal meiosis often leads to sterility, creating a biological bottleneck that makes it difficult to produce viable haploid embryos for doubling.
4. Reduction in Genetic Diversity
Strategic use of DH technology can have long-term biological consequences for a breeding program:
• Unintentional Selection: During the induction process, breeders may unintentionally practice selection for certain loci, which can limit genetic variation in the resulting populations.
• Loss of Recombination: Because DH technology bypasses the multiple generations of selfing used in traditional breeding, it provides fewer opportunities for genetic recombination to break unfavorable linkages.
5. Complexity in Polyploid Species
For polyploid crops like potato, the biological complexity of the genome adds further limits. Haploidization in these species is often used to simplify the genome for research, but high intra-somic competition and the tetraploid nature of the crop can result in a narrow spectrum of useful mutations and lower induction efficiency.
24. Can DH technology be used for polyploid crops like potato?
Yes, doubled haploid (DH) technology can be used for polyploid crops like potato. Reproducible and efficient DH protocols have already been established for potato, along with other major crops such as wheat, rice, maize, rapeseed, and sugar beet.
The application of DH technology in polyploid potato breeding involves the following processes and benefits:
Mechanism in Potato
In potato breeding, the creation of DH lines is a multi-step process designed to achieve complete genetic homozygosity:
• Haploid Induction: Haploids are typically obtained by crossbreeding tetraploid potato varieties (which have four sets of chromosomes, 2n=4x=48) with pollen from specific diploid varieties.
• Chromosome Doubling: These haploids are then converted back into polyploids through chromosome doubling techniques. This often requires the use of chemical agents like colchicine to induce the doubling of the genome.
Benefits for Polyploid Breeding
Polyploid crops like potato have more complex genetics than diploid crops, making traditional breeding more difficult. DH technology addresses these complexities by:
• Expressing Recessive Traits: Homozygous lines created via DH are particularly valuable for disease resistance breeding because they allow for the expression and fixation of recessive alleles that would otherwise be masked in a heterozygous polyploid state.
• Reducing Breeding Cycles: Like in other crops, DH can reduce the time-consuming process of developing stable parental lines from five generations down to just two.
• Simplifying Genetic Research: These lines provide a simplified genetic background for basic research and for identifying specific quantitative trait loci (QTLs).
Challenges and Limitations
Despite its established protocols, the use of DH in potatoes faces specific hurdles:
• Genotype Dependency: The success of haploid induction and plant regeneration is often highly dependent on the specific variety (genotype) being used [277, 138 (ref)].
• Inbreeding Depression: Cultivated tetraploid potatoes are notably heterozygous; forced inbreeding or selfing often results in severely weakened offspring and a lack of flowering, which DH technology aims to bypass by moving immediately to a stable homozygous state.
• Technical Complexity: The process requires specialized tissue culture techniques, such as anther or microspore culture, which can be difficult to optimize across all varieties.
25. How do breeders maintain genetic diversity when using DH?
Maintaining genetic diversity while using doubled haploid (DH) technology is a critical challenge because the process naturally accelerates homozygosity and can unintentionally limit variation. Breeders use several strategic approaches to mitigate this risk and preserve long-term adaptability.
The following strategies are used to maintain diversity in DH programs:
1. Haplotype Embedding
To prevent “selective sweeps”—where a specific genomic region becomes fixed across all lines, rendering a large portion of the surrounding genome unavailable to recombination—breeders deploy high-value genes into multiple different haplotypes. By evaluating breeding germplasm for haplotypic diversity, a breeder can choose several distinct elite recipients to act as parents, ensuring that a new gene is rolled out into the program without depleting the surrounding genetic variation.
2. Utilizing Super-Pangenomes
Breeders leverage super-pangenomes to capture genus-wide genetic diversity, including rare alleles and structural variants that might be overlooked by traditional SNP-based methods or lost during the rapid fixation of DH lines. These comprehensive datasets act as a “surprise package” of hidden diversity, allowing breeders to identify and reintroduce beneficial traits from wild relatives and landraces back into elite cultivars.
3. Integrated Genomic Selection (GS)
While DH rapidly fixes desirable alleles, Genomic Selection is used to manage the overall health of the breeding population.
• Preventing Inbreeding Depression: GS accounts for within-family genetic variance, helping to preserve long-term adaptability while increasing genetic gain.
• Broad Allelic Selection: By using genome-wide markers to predict breeding values, GS models allow breeders to select for a broad base of favorable alleles across the entire genome, rather than focusing solely on a few major loci which could lead to a diversity bottleneck.
4. Careful Selection and Recombination Planning
Because DH production bypasses several generations of traditional selfing, it provides fewer opportunities for genetic recombination to break unfavorable linkages. To counter this:
• Broad Germplasm Foundation: Breeders maintain a diverse germplasm base—including landraces and wild relatives—to ensure wide allelic representation in the initial crosses.
• Parental Profiling: Breeders use QTL profiling to fingerprint potential parents, allowing them to choose parent combinations that are genetically distinct and possess complementary traits, thereby maximizing the diversity of the resulting DH lines.
By integrating DH technology with predictive tools like AI and multi-omics, breeders can rapidly produce homozygous lines while monitoring and maintaining the genetic complexity required for resilient crops.
26. Can CRISPR enhance the development of new CMS parental lines?
CRISPR–Cas9 technology significantly enhances the development of new cytoplasmic male sterility (CMS) parental lines by providing the precision and speed needed to overcome the biological bottlenecks of traditional breeding. While traditional hybrid production for crops like cauliflower and onions relies on identifying and transferring sterile cytoplasm from wild relatives through years of backcrossing, CRISPR–Cas9 allows for more direct genomic interventions.
The technology supports this development through the following mechanisms:
• Rapid Creation of Sterile Lines: CRISPR can be used to knock out specific genes responsible for male fertility directly in elite cultivars. This approach allows researchers to create male-sterile “A-lines” as early as the T0 generation, effectively bypassing the 5 to 8 generations of selfing and crossing required by conventional methods.
• Synergy with Speed Breeding: When CRISPR-edited plants are grown under speed breeding conditions, it is possible to obtain stable, homozygous male-sterile phenotypes in less than a year. This combination addresses the “bottleneck” created by the long reproductive cycles of many crop species.
• Refining CMS Systems: Traditional CMS systems, such as the Ogura cytoplasm in radish and cauliflower, often suffer from physiological defects like chlorosis or temperature sensitivity. Advanced CRISPR tools like base editing and prime editing could potentially be used to precisely modify these mitochondrial or nuclear genomes to eliminate such defects while preserving the sterility trait.
• Improving the “Three-Line” System: Efficient hybrid production requires the coordinated development of A-lines (male sterile), B-lines (maintainers), and R-lines (restorers). CRISPR–Cas9 facilitates the functional characterization of fertility restorer genes, enabling breeders to engineer restorer lines with higher precision and fewer unintended genetic associations, such as “linkage drag”.
• AI-Guided Target Discovery: The integration of artificial intelligence with CRISPR–Cas9 mining of super-pangenomes helps identify novel regulatory motifs and genes linked to male sterility across different species, providing a broader range of targets for engineering next-generation hybrid systems.
By transforming hybrid breeding from an empirical selection process into a programmable design workflow, CRISPR–Cas9 technology enables the faster development of uniform, high-yielding, and resilient F1 hybrid varieties.
27. What are the ethical concerns regarding off-target CRISPR effects?
Ethical concerns regarding off-target CRISPR effects in plants center on the potential for unintended genomic rearrangements and mutations, which could lead to unforeseen ecological consequences once modified plants are released into natural environments. Because these inaccuracies can modify unintended portions of the genome, there are significant debates regarding their long-term environmental impacts, including the risk of cross-breeding with wild relatives or affecting non-target species.
Additional ethical and biosafety dimensions include:
• Framework-Specific Risks: In advanced applications like CRISPR-mediated Horizontal Gene Transfer (CRISPR-HGT), the use of microbial modules introduces risks such as immunogenicity, biological network disruption, or unintended metabolic burdens.
• Stability and Biodiversity: Legitimate concerns remain regarding the stability of complex trait expression across various environments and the potential for unintended gene flow to destabilize native ecosystems. This is particularly complex in perennial forestry species due to their long generation times and deep integration into natural habitats.
• Regulatory and Conceptual Blurring: The precision of CRISPR blurs the line between traditional breeding and genetic engineering, raising questions about whether edited crops should be viewed as “natural” and how they should be regulated globally.
• Governance and Equity: The uncertainty surrounding off-target effects emphasizes the need for inclusive governance frameworks that ensure transparency, public engagement, and equitable access to these technologies. There is also a socioeconomic concern that disparities in managing these technical risks could widen the genomic divide, leaving developing nations with less capacity to monitor or mitigate unintended consequences.
To address these concerns, comprehensive science-based risk assessments and harmonized regulatory policies are considered essential for the responsible and socially acceptable deployment of genome-edited crops.
28. How do researchers identify and minimize unintended CRISPR off-target effects?
Researchers identify and minimize unintended CRISPR off-target effects through a combination of predictive computational modeling, molecular engineering, and refined delivery strategies. While the precision of CRISPR–Cas9 is a significant breakthrough, minimizing unintended cleavage at genomic sites similar to the target sequence remains a critical technical challenge for biosafety and regulatory acceptance.
Identifying Off-Target Effects
To identify potential off-target sites, researchers primarily use the following methods:
• Computational Prediction: Specialized software tools predict off-target potential by analyzing genome-wide sequence similarities and features such as nucleotide composition and GC content.
• AI and Machine Learning: Advanced models, including DeepCRISPR, CRISPRon, and CRISPR-PLANT v2, are used to enhance the accuracy of off-target predictions and improve the design of highly specific single-guide RNAs (sgRNAs).
• Empirical Validation: Computational predictions are often combined with empirical laboratory validation to confirm whether unintended cleavage actually occurs at the predicted sites.
Minimizing Off-Target Effects
Researchers employ several strategies to increase targeting accuracy and reduce genomic instability:
1. sgRNA Optimization The design of the single-guide RNA (sgRNA) is a primary determinant of specificity. Optimization involves selecting sequences that achieve maximum on-target activity while minimizing similarity to other regions of the genome. Factors considered in this process include chromatin accessibility, secondary structure, and the specific genomic context of the target site.
2. Engineered Cas9 Variants To address non-specific DNA interactions, scientists have developed specialized versions of the Cas9 protein:
• High-Fidelity Cas9: These enzymes incorporate amino acid substitutions designed specifically to reduce non-specific binding, thereby improving accuracy.
• Nickase Cas9: These variants generate single-strand breaks (nicks) instead of double-strand breaks (DSBs). Using a “paired nicking” strategy—where two different sgRNAs direct two separate nicks—significantly enhances specificity.
• Stringent PAM Recognition: New variants of other nucleases, such as Cas12a, are being designed for lower off-target effects through more stringent recognition of the protospacer adjacent motif (PAM).
3. Base and Prime Editing Newer iterations of CRISPR technology, such as base editing and prime editing, minimize risks by allowing for precise nucleotide substitutions, insertions, or deletions without inducing double-strand breaks. This approach reduces the likelihood of unintended mutations and genomic rearrangements that can occur during the natural repair of a broken DNA strand.
4. DNA-Free Delivery Strategies Unintended effects are also minimized by controlling how the CRISPR components are delivered to the plant cell:
• Ribonucleoprotein (RNP) Delivery: Instead of delivering DNA that the cell must transcribe and translate, researchers deliver pre-assembled Cas9-sgRNA complexes.
• Transient Expression: RNPs offer a transient delivery platform that reduces the risk of unintended transgene integration into the host genome, which is a major concern for biosafety and regulatory approval.
• Protoplast Transfection: This method allows for DNA-free editing, further decreasing the potential for long-term genomic instability.
29. What are the ecological risks of releasing genome-edited crops into nature?
The release of genome-edited crops into natural environments presents several ecological risks, primarily stemming from unintended genetic changes and the potential for these modifications to spread beyond their intended agricultural setting.
The key ecological risks identified in the sources include:
1. Unintended Genetic Changes and Stability
• Off-Target Mutations: Despite advances in precision, there is a risk of modifying unintended portions of the plant genome. These inaccuracies could lead to unforeseen consequences if edited plants are released into the environment.
• Genomic Rearrangements: Genome editing can cause unintended genomic rearrangements that may impact the plant’s long-term performance or environmental interactions.
• Trait Stability: Legitimate concerns exist regarding the stability of complex trait expression across diverse and unpredictable environmental conditions.
2. Unintended Gene Flow and Biodiversity Impacts
• Cross-Breeding with Wild Relatives: A significant concern is the risk of modified plants cross-breeding with wild relatives, potentially introducing novel traits into natural populations.
• Destabilizing Native Ecosystems: There is a potential for unintended gene flow to destabilize native ecosystems. This is particularly complex in perennial forestry species due to their long generation times, extensive pollen-mediated gene flow, and deep integration into natural habitats.
• Spread Across Landscapes: In forestry, traits may spread across entire landscapes before their full ecological impacts are realized, making containment and post-release monitoring significantly more challenging.
• Herbicide Resistance Spillover: One specific risk of gene flow is the potential for herbicide resistance to spill over into wild plant populations, creating “superweeds”.
3. Impacts on Non-Target Species and Biotic Interactions
• Impact on Non-Target Species: Modified crops may have unintended impacts on non-target species, such as beneficial insects or soil microorganisms.
• Disrupting Symbioses: In advanced frameworks like CRISPR-mediated Horizontal Gene Transfer (CRISPR-HGT), reintroducing microbial genes could inadvertently shift plant resource allocation or disrupt established symbiotic relationships, such as those with arbuscular mycorrhizal fungi.
4. Risks Specific to Advanced Frameworks (CRISPR-HGT)
• Microbial Gene Risks: Reintroducing microbial modules into plants can introduce inherently novel risks, such as immunogenicity, biological network disruption, or unintended metabolic burdens.
• Pleiotropic Effects: Stacking multiple microbial modules could increase pleiotropy (where one gene affects multiple traits) and place a higher metabolic burden on the plant, potentially affecting its overall fitness under benign conditions.
To manage these risks, the sources emphasize the necessity of science-based risk assessments and inclusive governance frameworks that involve early dialogue among scientists, regulators, and the public to ensure responsible deployment.
30. Can AI better predict long-term ecological impacts of off-target effects?
Yes, artificial intelligence (AI) is increasingly being used to predict and mitigate the long-term ecological impacts and off-target effects of genome editing. While traditional methods rely on empirical observation, the integration of AI transforms genome editing into a predictive system capable of modeling complex biological and environmental interactions.
Predictive Modeling of Off-Target Effects
AI and machine learning are primary tools for identifying potential off-target mutations before they occur.
• Precision Tools: Researchers use advanced software such as DeepCRISPR, CRISPRon, and CRISPR-PLANT v2 to enhance guide RNA (sgRNA) design and predict off-target potential with high accuracy.
• Data Integration: These models analyze genome-wide sequence similarities, nucleotide composition, and chromatin accessibility to identify sites where unintended cleavage is most likely to happen.
Anticipating Ecological Trade-offs
In advanced frameworks like CRISPR-mediated Horizontal Gene Transfer (CRISPR-HGT), AI plays a critical role in evaluating how new genetic modules will behave in natural ecosystems.
• Functional Integration: Machine learning models trained on multi-omics and phenotypic datasets can predict the compatibility of introduced genes with existing plant regulatory and metabolic networks.
• Predicting Trade-offs: AI helps researchers anticipate trade-offs among stacked traits, such as identifying if a gene that provides drought tolerance might inadvertently increase the plant’s metabolic burden or disrupt beneficial symbioses under normal conditions.
• Stability of Traits: Predictive modeling is used to ensure the stability of complex trait expression across diverse and unpredictable environmental conditions, which is a major concern for the release of edited crops into nature.
Risk Assessment in Complex Species
For perennial forestry species, where generation times are long and ecological integration is deep, AI-guided prediction is vital.
• Long-term Monitoring: AI can help model the spread of edited traits across landscapes over longer ecological timescales than is possible for annual crops.
• Minimizing Ecological Risk: By integrating high-resolution phenotyping with AI, breeders can optimize “trait architecture” to minimize the risk of modified plants cross-breeding with wild relatives or destabilizing native ecosystems.
Limitations and Necessity of Validation
Despite the power of AI, the sources emphasize that experimental confirmation remains essential. While AI can predict potential outcomes and identify the most promising candidates for field deployment, it cannot yet fully replace rigorous, science-based risk assessments and laboratory validation to confirm model-derived insights.
References
Ahmar, S., Gill, R. A., Jung, K. H., Faheem, A., Qasim, M. U., Mubeen, M., & Zhou, W. (2020). Conventional and molecular techniques from simple breeding to speed breeding in crop plants: Recent advances and future outlook. In International Journal of Molecular Sciences (Vol. 21, Number 7). MDPI AG. https://doi.org/10.3390/ijms21072590
Atanassova, A., & Keiper, F. (2018). Plant breeding innovation: A global regulatory perspective. In Cereal Chemistry (Vol. 95, Number 1, pp. 8–16). Wiley-Blackwell. https://doi.org/10.1002/cche.10021
Bradshaw, J. E. (2017). Plant breeding: past, present and future. Euphytica, 213(3). https://doi.org/10.1007/s10681-016-1815-y
Buch, K., Kaushik, A., Mishra, U., Beese, S., Samanta, S., Singh, R., & Mubeen. (2023). Unravelling the Complexity of Plant Breeding through Modern Genetic Techniques and Tools: A review. International Journal of Plant & Soil Science, 35(21), 97–105. https://doi.org/10.9734/ijpss/2023/v35i213950
Cobb, J. N., Biswas, P. S., & Platten, J. D. (2019). Back to the future: revisiting MAS as a tool for modern plant breeding. In Theoretical and Applied Genetics (Vol. 132, Number 3, pp. 647–667). Springer Verlag. https://doi.org/10.1007/s00122-018-3266-4
Edukondalu, B., Aswini, N., Amaresh, Krishnappa, G., Soundharya, B., Nikhitha, G., Pathy, T. L., Krishna, K., Hari, Y., & Vinayaka. (2026). Accelerating genetic gain through integrated genomic selection in crop plants. In Journal of Applied Genetics. Springer Science and Business Media Deutschland GmbH. https://doi.org/10.1007/s13353-025-01034-7
KATSUTA, M., KATSU, K., & SUZUKI, T. (2026). Current Status and Prospects for Perennial Crop Breeding. Japan Agricultural Research Quarterly: JARQ, 24S15. https://doi.org/10.6090/jarq.24S15
Raza, A., Li, Y., Jan, F., Fernandez, C. G. T., Mir, R. R., Hu, Z., & Varshney, R. K. (2026). From the genome to super-pangenome: a new paradigm for accelerated crop improvement. Npj Science of Plants, 2(1), 4. https://doi.org/10.1038/s44383-025-00019-z
Sen, M. K., Roy, A., Varshney, R. K., & Chakraborty, A. (2026). Engineering next-generation crops through CRISPR-mediated horizontal gene transfer. In New Phytologist. John Wiley and Sons Inc. https://doi.org/10.1111/nph.70951
Sidhu, N. S., Prasad, G., Sowmya, M. S., Kavya, M. E., Krishnappa, G., & Pathy, T. L. (2025). Genome-Wide Association Studies for Next-Generation Crop Breeding in the Postgenomic Era. In Plant Breeding. John Wiley and Sons Inc. https://doi.org/10.1111/pbr.70017
Sokra, I. (2026). Attribution-Share Alike 4.0 International CC BY-SA 4.0 CRISPR-Cas9 Applications in Plant Science: Advances, Challenges, and Future Perspectives. Journal of Agriculture and Technology, 2(1), 129–139. https://doi.org/10.5281/zenodo.17983250
Yoosefzadeh-Najafabadi, M. (2025). Merging traditional practices and modern technology through computational plant breeding. In Plant Physiology (Vol. 199, Number 1). American Society of Plant Biologists. https://doi.org/10.1093/plphys/kiaf355






