1. Introduction: Why Build a Chromosome?
Imagine a cell’s natural chromosome as a vast, intricate instruction manual. This manual, written in the language of DNA, contains all the genetic information an organism needs to develop, function, and reproduce. Every time a cell divides, it must perfectly copy and sort these manuals so that each new cell gets a complete set.
So, what is a chromosome? It is a highly organized package of DNA that ensures this massive amount of genetic information is managed correctly. But why would scientists want to build one from scratch? Creating an artificial chromosome is like writing a new, custom chapter for the instruction manual. It allows researchers to deliver large amounts of genetic information into a cell in a stable, controlled way—a feat that opens up exciting possibilities for treating genetic diseases and understanding the fundamental rules of life.

2. Introduction to Artificial Chromosome Technology
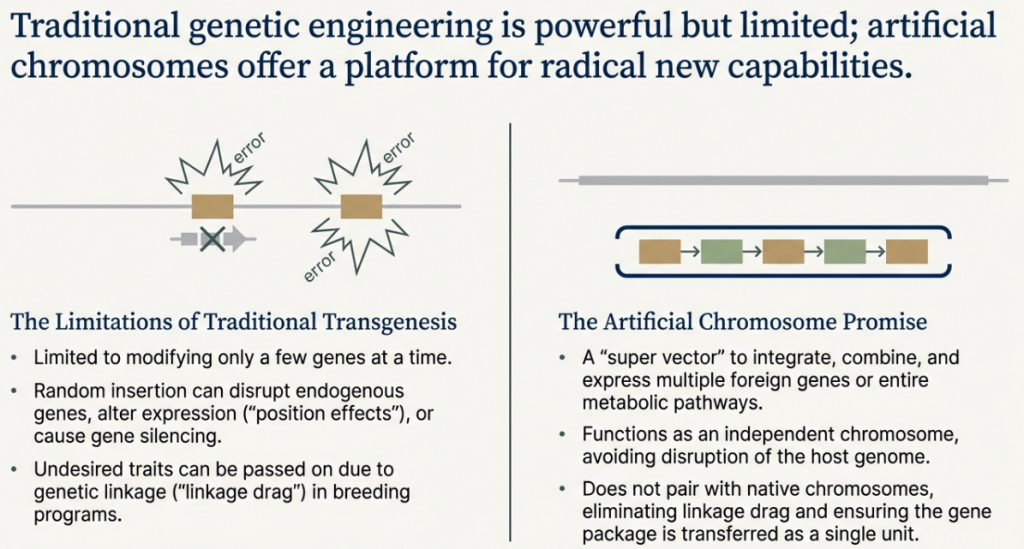
An artificial chromosome is a DNA molecule of predictable structure, assembled in vitro from defined biological components, that is designed to carry large amounts of genetic information and behave with the properties of a natural chromosome.

Its most important feature is that it works as a separate, independent chromosome that doesn’t interfere with the person’s own DNA. Because it is maintained apart from the host’s chromosomes, it acts as a “genomic safe island,” a place where new genes can be added without the risk of disrupting the cell’s existing genetic blueprint.
Compared to other gene therapy methods, like using viruses to deliver genes, artificial chromosomes offer several key advantages:
• Capacity for Large Genes: They can carry enormous genes or even entire sets of genes. This is crucial for diseases caused by faults in very large genes, which are too big for most other gene therapy tools to handle.
• Autonomous Function: They operate independently and are not integrated into the host’s own chromosomes. This avoids a dangerous problem called insertional mutagenesis, where a new gene is accidentally inserted into a healthy gene, potentially causing new health issues like cancer.
• Long-Term Stability: They are built to be stable and are copied along with the cell’s other chromosomes during cell division. This offers the potential for a one-time, long-lasting correction that is passed on to all future daughter cells.
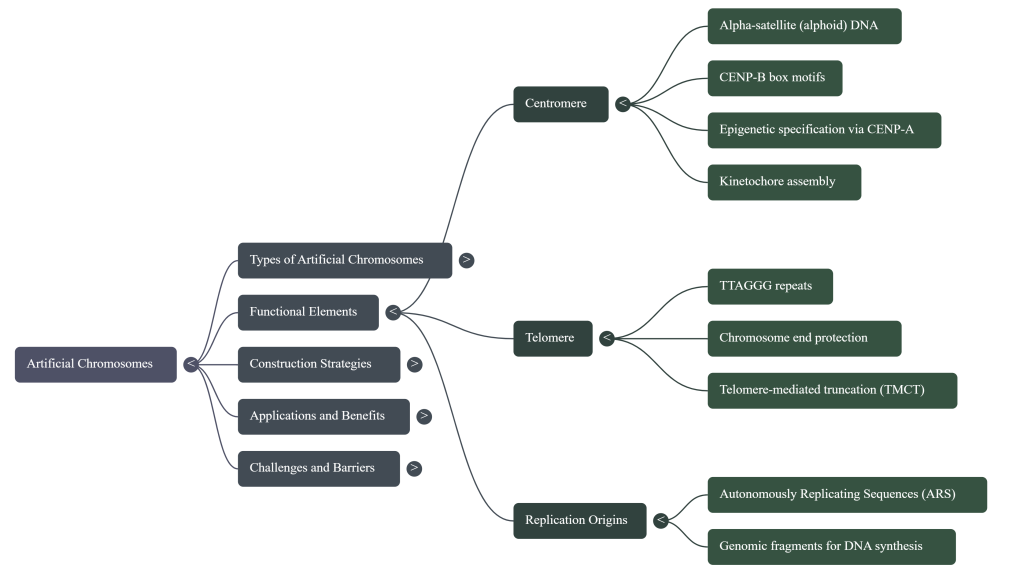
The remarkable stability and autonomy of these chromosomes are made possible by incorporating the three essential DNA components that every natural chromosome relies on to function.
Pioneering research in yeast, later extended to mammals, has established that a functional chromosome requires three essential cis-acting DNA sequences (meaning, sequences that affect only the DNA molecule on which they are located). These components form the fundamental blueprint for constructing any artificial chromosome. Think of these as the non-negotiable hardware for any genetic instruction manual.
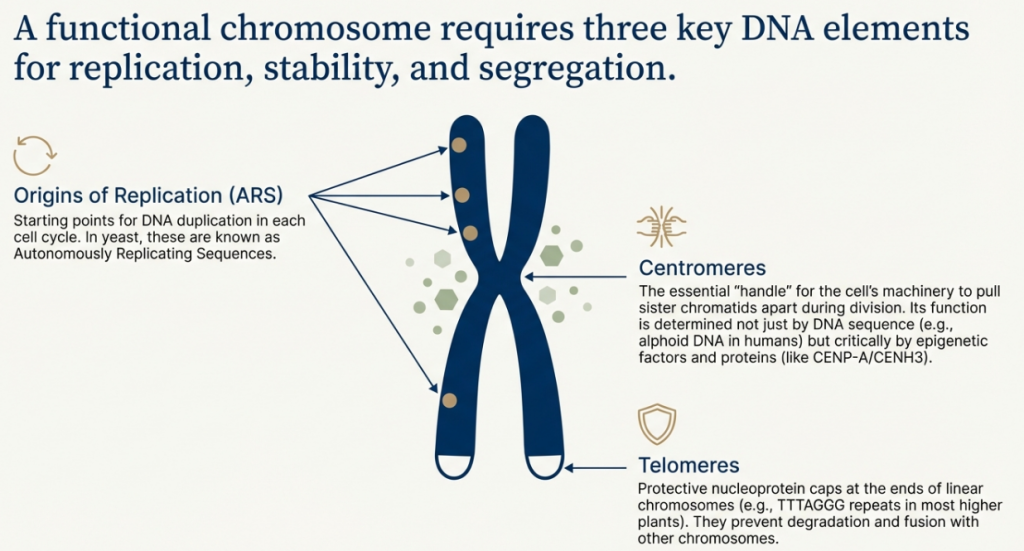
2.1 The Centromere: The Master Coordinator
The centromere is the structural core of a chromosome and its primary role is to serve as the attachment point for the cell’s sorting machinery during division. As the cell prepares to split, a network of protein fibers called spindle microtubules latches onto the centromere.
You can think of the centromere as a “handle” or “gripper” that allows the cell to grab the chromosome and pull it to the correct destination, ensuring each new cell receives exactly one copy. In humans, this region is typically made of highly repetitive DNA sequences known as “alphoid DNA.” However, its function isn’t just about the DNA sequence itself; it’s also determined epigenetically. A special protein called CENP-A marks the location, effectively telling the cell’s machinery, “This is the centromere.” This discovery—that function can be dictated by a protein marker rather than a rigid DNA sequence—is not just a detail; it’s a foundational concept that opens the door to entirely new ways of building chromosomes, as we will see later.
2.2 Telomeres: The Protective End Caps
Telomeres are specialized DNA sequences found at the two ends of a linear chromosome. Their function is purely protective. They prevent the chromosome ends from being mistaken for broken DNA, which could lead to them being degraded by cellular enzymes or accidentally fusing with other chromosomes.
The classic analogy for telomeres is the plastic tips on a pair of shoelaces (aglets). Just as aglets prevent the shoelace from fraying and unraveling, telomeres protect the integrity of the genetic information stored on the chromosome.
2.3 Origins of Replication: The Starting Lines for Copying
Before a cell can divide, it must duplicate its entire genome. Origins of replication are specific sites scattered along the length of the chromosome where this copying process begins. Without these sequences, the cell’s DNA-copying machinery wouldn’t know where to start.
These origins function like designated “starting lines” in a race, signaling to the cellular machinery where to begin duplicating the instruction manual to ensure the entire chromosome is copied efficiently and completely.
2.4 Key Components at a Glance
This table provides a quick summary of the three essential parts of any functional chromosome.
| Component | Primary Function | Simple Analogy |
|---|---|---|
| Centromere | Serves as the attachment point for microtubules to sort chromosomes during cell division. | A “handle” for the cell’s machinery to grab. |
| Telomeres | Protect the ends of the chromosome from degradation and fusion. | The plastic tips on a shoelace (aglets). |
| Origins of Replication | Signal the starting points for DNA duplication before cell division. | “Starting lines” for copying the DNA. |
Now that we have identified the essential components in our construction kit, the next step is to understand how scientists assemble them into a working artificial chromosome.
The foundational principles of AC construction were first established in a simple eukaryotic model, yeast, providing a conceptual framework that guided all subsequent efforts in mammals.
3. Foundational Principles from Yeast Artificial Chromosomes (YACs)
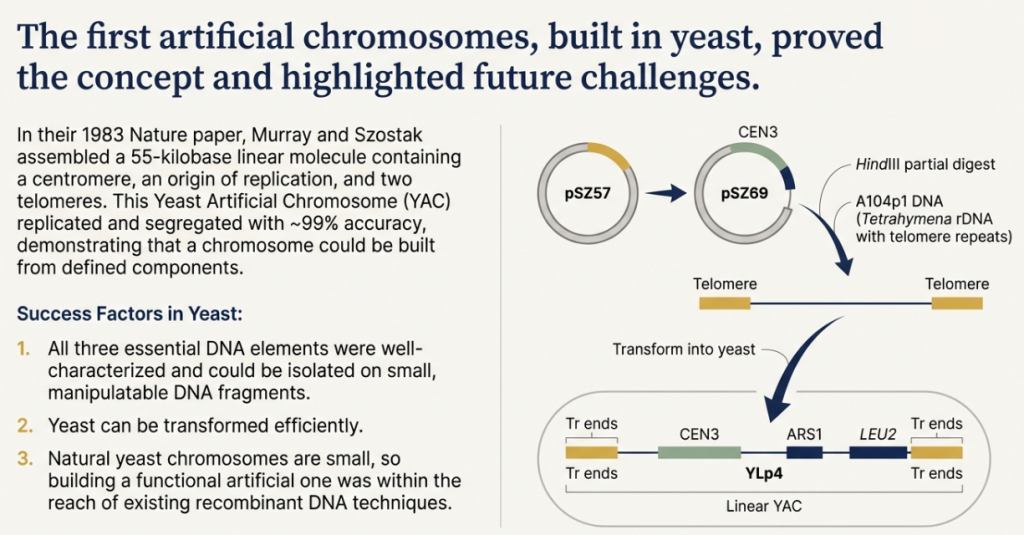
The creation of Yeast Artificial Chromosomes (YACs) in the early 1980s by Murray and Szostak represented the first successful construction of a synthetic chromosome. This achievement was a landmark in molecular biology, demonstrating that a functional chromosome could be assembled from discrete, cloned DNA components. The principles discovered and the methodologies developed in the yeast system provided the fundamental blueprint for all subsequent attempts to build artificial chromosomes in more complex organisms, including mammals. YACs not only proved the concept but also revealed critical design rules governing chromosome behavior.
The methodology for constructing the first YACs was elegant in its approach. The three essential components—cloned telomeres, a centromere sequence (CEN), and an autonomously replicating sequence (ARS)—were first assembled into a circular plasmid vector within E. coli. This plasmid was then transformed into yeast cells. Once inside the yeast, the construct resolves into a linear molecule through the action of the telomere sequences, which form stable ends. This process successfully converted a bacterial plasmid into a linear, self-replicating molecule that behaved like a miniature yeast chromosome.
The YAC system was instrumental in uncovering a fundamental principle of chromosome biology: the direct relationship between physical size and mitotic stability. Early experiments revealed a stark contrast in behavior based on length. Short linear plasmids (<20 kilobases), even when containing a centromere, were highly unstable during mitosis, with segregation frequencies as poor as 0.11 (meaning a loss in 11% of divisions). These constructs were often lost during cell division and, as a compensatory mechanism, were maintained at a high copy number within the cells that retained them. In contrast, when the linear plasmids were elongated to 55 kilobases, their behavior changed dramatically. These larger artificial chromosomes exhibited a high degree of mitotic stability, were maintained at a stable, low copy number (one or two copies per cell), and segregated faithfully, much more closely mimicking natural yeast chromosomes.
The successful engineering of YACs laid the intellectual groundwork for building artificial chromosomes in other species, but the lessons learned also highlighted the immense scale of the challenge ahead, as mammalian chromosomes are two to three orders of magnitude larger and structurally more complex than those of yeast.
The Pioneer: Yeast Artificial Chromosomes (YACs)
The first functional artificial chromosomes were built in yeast cells. This pioneering work was possible for three main reasons:
1. Known Components: The three essential DNA sequences—centromeres, telomeres, and origins of replication—were already well-understood and isolated in yeast.
2. Easy to Manipulate: Yeast cells are simple to grow and can be transformed with new DNA efficiently, making experiments relatively quick and easy to perform.
3. Manageable Size: Natural yeast chromosomes are quite small, so assembling a synthetic molecule large enough to be functional was technically feasible with the tools available at the time.
4. From Yeast to Bacterial Systems
Yeast Artificial Chromosomes (YACs) were the first vectors capable of cloning DNA fragments greater than 500 kb. While revolutionary for genome mapping projects, their utility for therapeutic applications was severely limited by significant technical flaws. YACs were prone to a high degree of chimerism, where a single YAC clone contained fragments from different genomic locations, with rates reported as high as 40%. They were also susceptible to insert rearrangement and general instability within the yeast host, making them unreliable for high-fidelity gene transfer.
The limitations of YACs spurred the development of more stable and manageable systems. Bacterial Artificial Chromosomes (BACs), derived from the single-copy F-factor plasmid of E. coli, and P1-derived Artificial Chromosomes (PACs) quickly became the systems of choice for constructing large-insert genomic libraries. The key differences between these foundational systems are summarized below.
| Feature | YAC | BAC |
|---|---|---|
| Configuration | Linear | Circular |
| Host Organism | Yeast | Bacteria (E. coli) |
| Copy Number | 1 | 1–2 |
| Cloning Capacity | >500 kb, with some constructs reaching up to 2.5 Mb | Up to 350 kb |
| Transformation Efficiency | Low (~10⁷ transformants/μg) | High (~10¹⁰ transformants/μg) |
| Insert Stability | Unstable | Stable |
| Chimerism | High (up to 40%) | None to low |
BACs and PACs offered superior cloning efficiency, greatly improved fidelity with minimal rearrangement, and were significantly easier to purify and handle using standard molecular biology techniques. Their enhanced stability and reliability made them the foundational tools for large-scale genome sequencing and physical mapping. While these systems perfected the cloning of large DNA fragments, they were merely carriers; they lacked the functional elements required for mitotic stability in a mammalian cell, the most challenging of which was the centromere.
5. Methodologies for Mammalian Artificial Chromosome (MAC/HAC) Construction
The construction of stable Mammalian Artificial Chromosomes (MACs) or Human Artificial Chromosomes (HACs) marked a major technological leap beyond the YAC system.
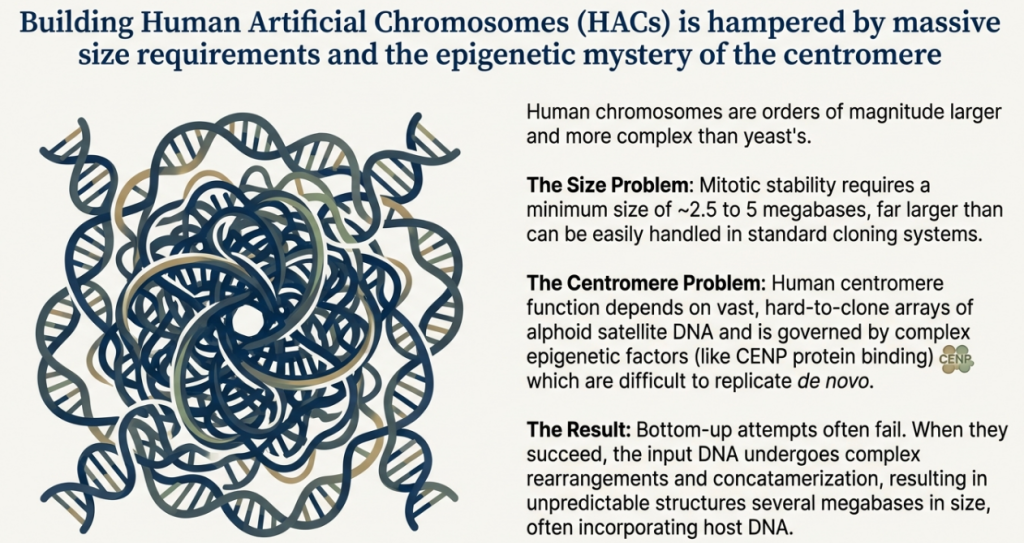

5.1 The Central Challenge: Defining and Engineering the Mammalian Centromere
The centromere stands as the most critical and complex component of a functional artificial chromosome. Its paramount role is to ensure accurate mitotic segregation, and our evolving understanding of how centromere identity is established—from a purely sequence-based (genetic) model to an epigenetically defined one—has directly driven the evolution of HAC design. For decades, the precise nature of the mammalian centromere remained elusive, presenting the single greatest obstacle to building a chromosome from the ground up.
The traditional genetic approach to engineering a centromere focused on alpha-satellite (or alphoid) DNA. This family of highly repetitive DNA, characterized by a 171 base-pair monomer unit, is found in large arrays (from 500 kb to 5 Mb) at the centromeres of all natural human chromosomes. Early HAC construction efforts logically centered on cloning these sequences, assuming they contained the necessary genetic information to specify centromere function. However, this approach was fraught with technical challenges; the long, repetitive nature of alphoid DNA made it extremely difficult to clone and propagate in bacterial systems without incurring deletions and rearrangements. Furthermore, early experiments revealed that the resulting artificial chromosomes were often much larger than the sum of their input DNA components, indicating that significant DNA rearrangement or amplification had occurred. This unpredictability underscored the difficulty of the approach and the need for alternative strategies.
A paradigm shift occurred with the realization that centromere function is primarily defined by epigenetic mechanisms rather than by a specific DNA sequence. It became clear that alphoid DNA is neither strictly necessary nor sufficient for centromere formation. This was powerfully demonstrated by the discovery of “neocentromeres,” functional centromeres that form on non-centromeric, non-repetitive regions of chromosomes. The key epigenetic marker that defines centromere identity and function was identified as CENP-A, a specialized histone H3 variant. The presence of CENP-A-containing nucleosomes is now understood to be the definitive mark that recruits the machinery of the kinetochore, irrespective of the underlying DNA sequence.
Within the context of alphoid DNA, the CENP-B protein plays a significant role. CENP-B is a DNA-binding protein that recognizes a specific 17 bp sequence known as the “CENP-B box,” which is interspersed throughout many alphoid DNA arrays. While not essential for centromere function on its own, CENP-B contributes to the formation and organization of centromeric chromatin on its native alphoid DNA template. This dual understanding of the centromere—as a locus that could be defined genetically by alphoid DNA or epigenetically by CENP-A—gave rise to two distinct philosophical and technical approaches for building a HAC.
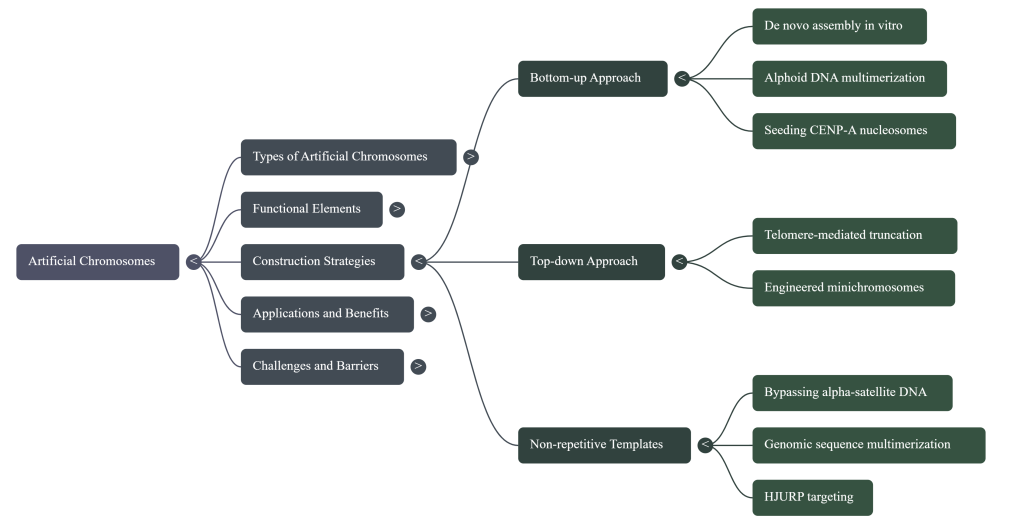
Overcoming the challenges posed by the size and complexity of mammalian chromosomes required the development of two distinct and powerful strategies. The first, a “top-down” approach, leverages existing chromosomes as a scaffold, while the second, a “bottom-up” approach, aims to build a chromosome entirely from defined molecular parts. Each strategy presents a unique set of advantages and technical hurdles.
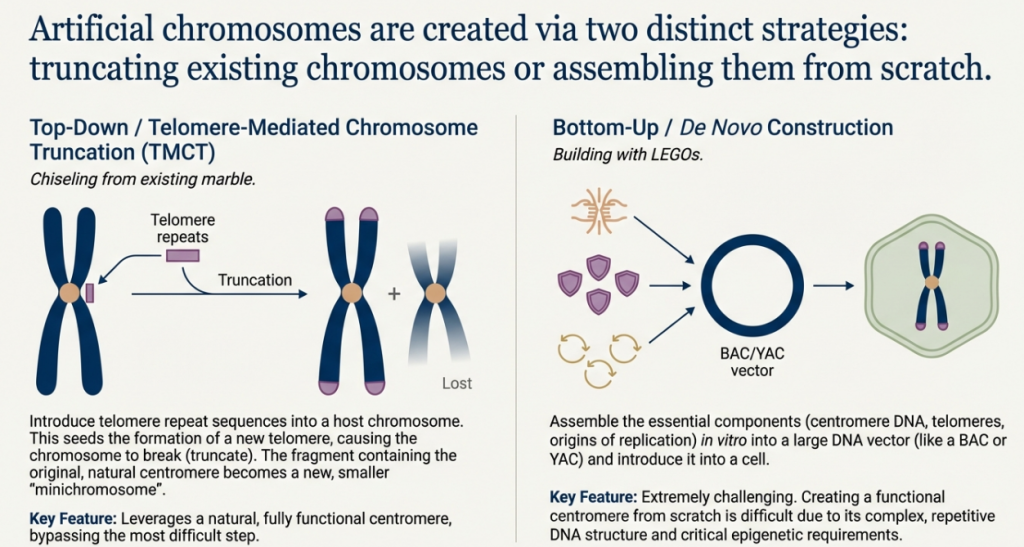
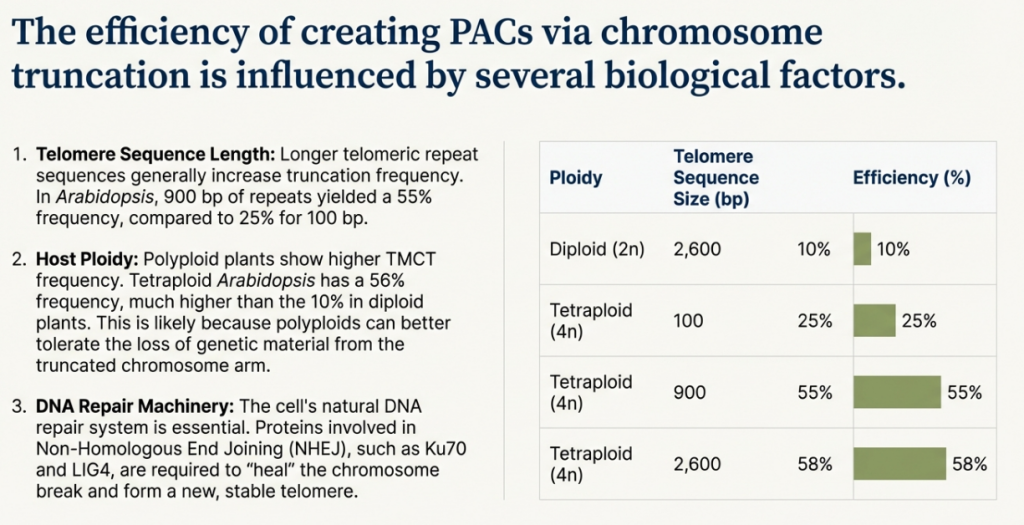
5.2 The “Top-Down” Approach: Telomere-Mediated Chromosome Truncation (TMCT)
Telomere-Mediated Chromosome Truncation (TMCT) is a method for systematically engineering an existing natural chromosome. The process involves introducing cloned telomeric DNA sequences into a specific site on a host chromosome. This targeted insertion provides a substrate for telomerase, leading to the de novo synthesis of a functional telomere at that location, which in turn leads to the truncation of the chromosome arm distal to the integration site. The result is a stable, shortened “mini-chromosome” that retains its original, fully functional native centromere.
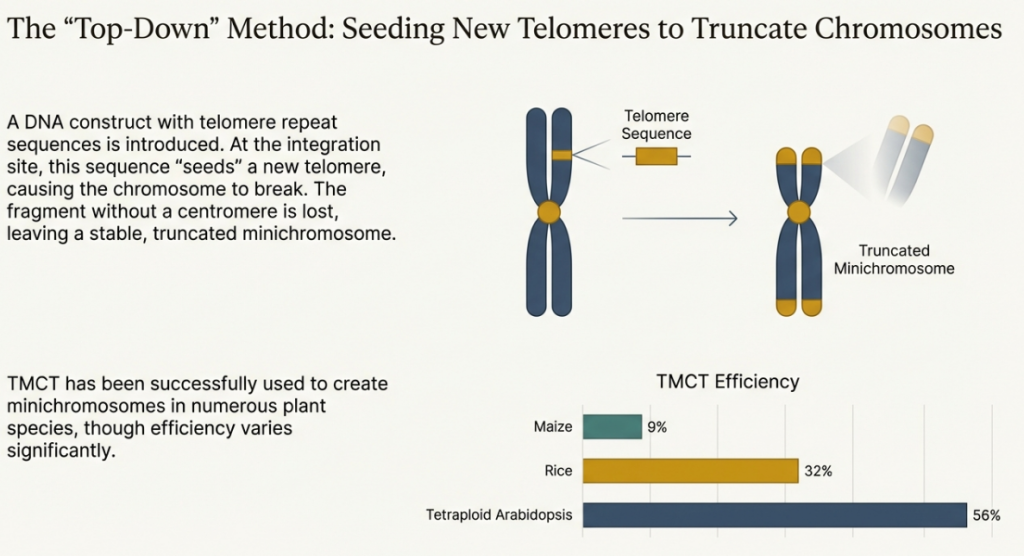

The primary advantage of the TMCT approach is the creation of a vector with a well-defined structure and, most importantly, a proven, natural centromere that guarantees high-fidelity segregation. This bypasses the significant challenge of building a functional centromere from scratch. However, the method is limited by its reliance on an existing chromosome and the fact that truncation inevitably leads to the loss of genetic material, which can be detrimental to the cell if essential genes are located on the discarded fragment.
5.3 The “Bottom-Up” Approach: De Novo Assembly from Cloned DNA
The “bottom-up” strategy represents the ultimate goal of synthetic genomics: the complete assembly of a functional chromosome from discrete, cloned DNA components without using a native chromosome as a template.
5.3.1 Initial Strategies and the Alphoid DNA Challenge
Early attempts at de novo construction involved co-transfecting three key components into human cells: telomeric DNA, a selectable marker gene, and large quantities of human alphoid satellite DNA. Alphoid DNA is the highly repetitive sequence that constitutes the bulk of natural human centromeres. These pioneering experiments successfully generated mini-chromosomes, proving that de novo formation was possible. However, the process was inefficient and difficult to control. The resulting HACs often had complex, unpredictable structures, frequently incorporating host DNA and exhibiting significant rearrangements. The highly repetitive and unstable nature of the input alphoid DNA was identified as a major technical obstacle, making the DNA difficult to clone, handle, and analyze.
5.3.2 YAC-Based Construction with Centromeric DNA
A more controlled and efficient bottom-up method was subsequently developed by Ikeno et al., which used a Yeast Artificial Chromosome as an intermediate vector to carry a defined, 100-kilobase (kb) array of human alphoid DNA. This YAC, containing the centromeric “seed” DNA, was purified and introduced into human cells. This refined approach yielded several critical insights into the requirements for de novo centromere formation and chromosome stability.
| Observation | Implication |
|---|---|
| An α21-I YAC with a regular alphoid repeat structure and CENP-B boxes efficiently formed MACs in a high proportion of cell lines (8 of 24 total), while a more divergent α21-II YAC did not. | The specific sequence composition and higher-order structure of the input centromeric DNA are crucial for initiating de novo centromere formation. This highlights that not all alphoid DNA is equally competent and underscores the importance of cloning strategies (such as using a rad52 yeast host) that can maintain the integrity of long, regular satellite arrays. |
| The resulting MACs were 10- to 50-fold larger (1-5 megabases) than the input 100 kb YAC. Structural analysis via FISH using 24 different human chromosome paint probes showed no acquisition of host chromosome arms. | Simple amplification or multimerization of the input DNA, rather than acquisition of host DNA, is a key cellular mechanism for achieving the minimum physical size necessary for mitotic stability. |
| The formed MACs successfully recruited essential centromere proteins (CENP-B, CENP-C, CENP-E) and segregated with high fidelity (>99% per division). The most stable MACs showed loss rates as low as 0.5% per cell division, approaching the stability of native chromosomes. | This method successfully generates a fully functional centromere and a mitotically stable chromosome de novo from a defined set of molecular components. |
5.4 The Modern Frontier: An Epigenetic Shortcut
A major obstacle in traditional “bottom-up” chromosome construction has always been the centromere. The highly repetitive nature of the alphoid DNA that makes up human centromeres is incredibly difficult to clone, sequence, and handle in the lab. This technical challenge represented a significant bottleneck, preventing the efficient and reliable creation of HACs.
5.4.1 Bypassing the Repetitive DNA Problem
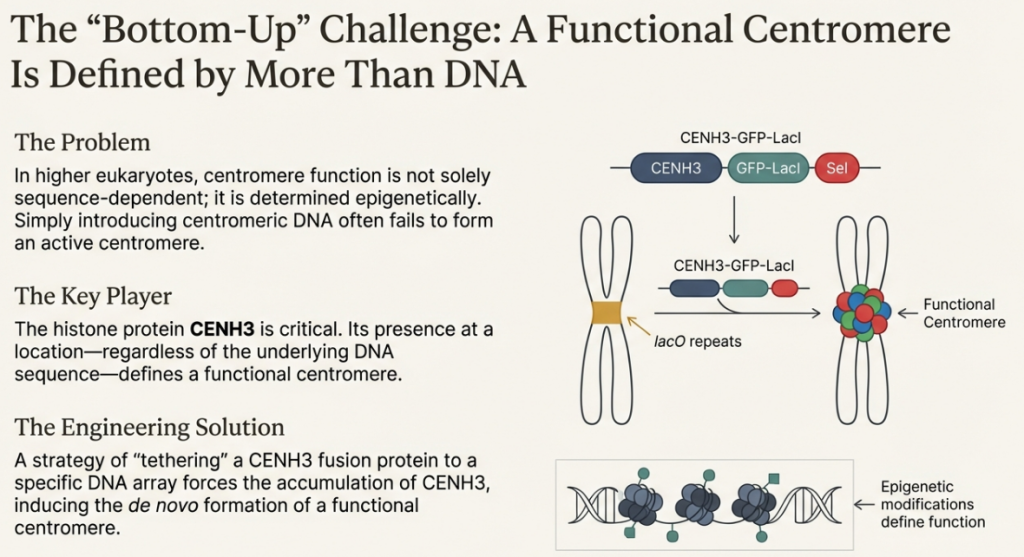
The solution to this problem came from the very epigenetic principle we encountered earlier. The most recent breakthrough in HAC construction is an innovative strategy that bypasses the need for repetitive alphoid DNA altogether. Instead of relying on a specific DNA sequence, scientists can now hijack the cell’s epigenetic machinery.
The key is the centromere protein CENP-A and its dedicated assembly factor, a protein called HJURP. Researchers have engineered systems to artificially target HJURP to a chosen, non-repetitive DNA sequence. This forces the cell’s machinery to deposit CENP-A at that specific spot, effectively “seeding” a new centromere. This protein marker tells the cell, “The centromere is here,” creating a fully functional handle without the need for the underlying repetitive DNA sequence. This innovation frees HAC construction from its biggest technical hurdle and represents a major step forward for synthetic biology.
This approach is transformative because it decouples centromere identity from the constraints of repetitive DNA and enables the construction of truly synthetic, designer chromosomes with fully defined sequences. The development of these diverse construction methodologies has provided powerful tools for creating artificial chromosomes, but their ultimate success depends on a deeper understanding of the specific molecular and structural factors that govern their stability and function.

6. Critical Factors in Artificial Chromosome Stability and Function
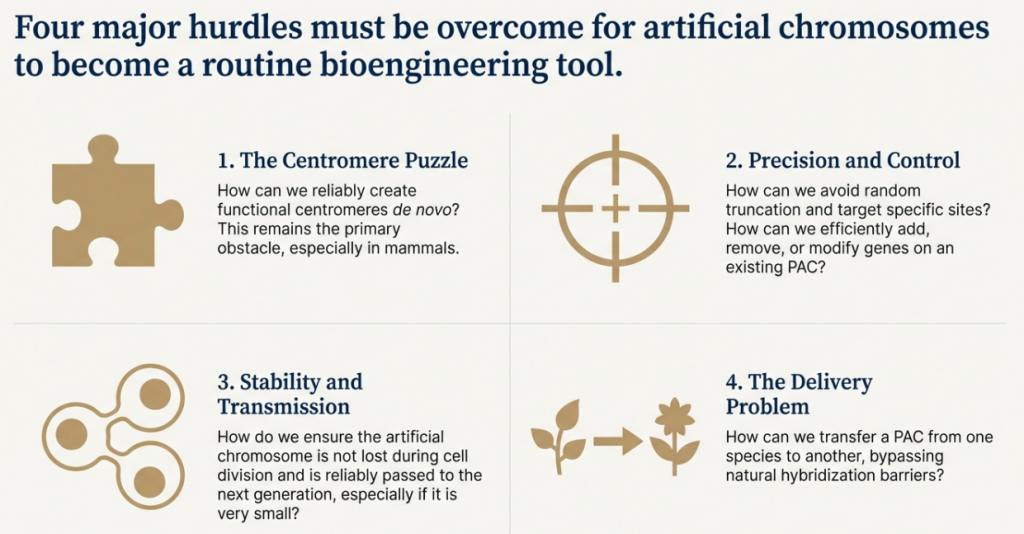
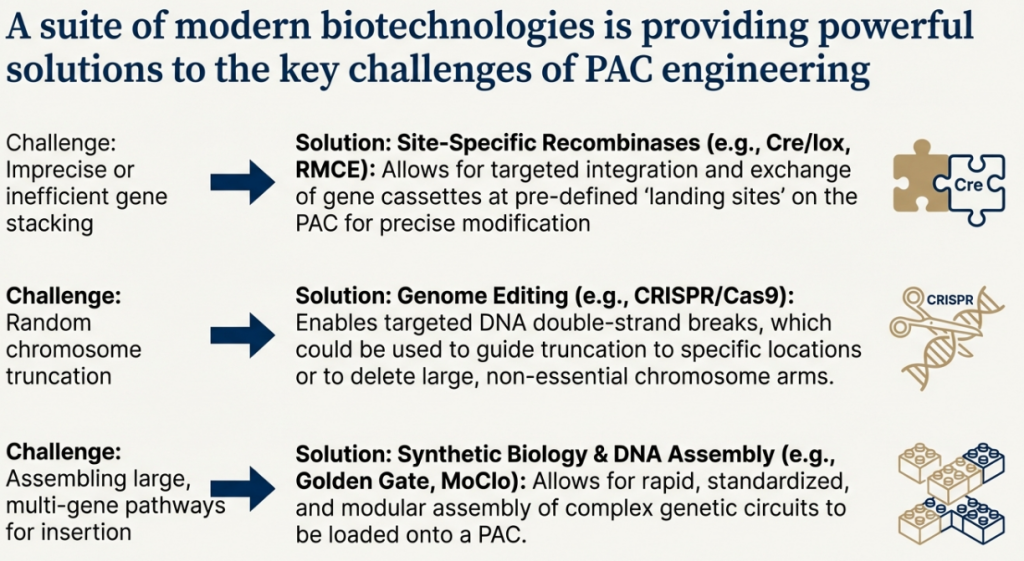
The success of any artificial chromosome platform, whether for research or therapeutic application, is contingent upon mastering the molecular and structural determinants of its stability and function. Building an AC is only the first step; ensuring it behaves predictably as an independent chromosome requires a sophisticated understanding of its core components. This section dissects the central role of the centromere, the critical impact of physical size, and the practical challenges of delivery and functional validation.
6.1 The Centromere: Contrasting Sequence-Dependent and Epigenetic Models
The centromere is the functional heart of the chromosome, and understanding its formation has been central to AC development. Two complementary models explain how a centromere is established.
6.1.1 The Sequence-Dependent Model of Assembly
In traditional HACs built with alphoid DNA, sequence plays a critical initiating role. This model centers on the centromere protein B (CENP-B), which binds specifically to a 17-base-pair motif known as the “CENP-B box” found within alphoid DNA repeats. Evidence strongly indicates that the binding of CENP-B to these boxes is a necessary first step for initiating the de novo assembly of a functional centromere on alphoid DNA. This model is strongly supported by the seminal work of Ikeno et al., where the α21-I YAC, rich in CENP-B boxes, successfully seeded de novo MACs, while the divergent α21-II YAC, which lacked a regular CENP-B box array, failed to do so. While CENP-B is not required for the maintenance of already-established centromeres, its role as a seeding factor is crucial for bottom-up construction using natural centromeric DNA.
6.1.2 The Epigenetic Model of Identity
In contrast, the epigenetic model positions the specialized histone H3 variant, CENP-A, as the master determinant of centromere identity and function. According to this model, the defining feature of a centromere is not a specific DNA sequence, but rather the presence of CENP-A-containing nucleosomes. Once CENP-A is loaded onto a region of chromatin—whether through sequence-dependent mechanisms or artificial tethering—it creates a stable epigenetic mark. This mark is sufficient to recruit the entire downstream cascade of kinetochore proteins required for spindle attachment and proper chromosome segregation, irrespective of the underlying DNA sequence. This understanding has enabled the creation of HACs on completely unique, non-repetitive DNA sequences.
6.2 The Impact of Multimerization and Chromosome Size
A consistent theme, emerging first from the foundational YAC studies and confirmed in mammalian systems, is that a minimum physical size is required for a chromosome to be mitotically stable. This size requirement highlights the dramatic increase in scale from yeast to mammals; whereas YACs achieved stability at 55 kb, stable MACs require a size two orders of magnitude larger, in the multi-megabase range. The estimate for a stable mammalian chromosome is approximately 10 megabases (Mb).
This size requirement helps explain the consistent observation that de novo MACs undergo significant multimerization. When a relatively small DNA construct (e.g., a 100 kb YAC) is introduced into a cell to form a MAC, it undergoes amplification to produce a final chromosome that is many times larger, often 1–5 Mb. While still below the theoretical ~10 Mb optimum for native chromosome stability, this amplification is a necessary cellular mechanism to create a molecule physically large enough to engage the mitotic spindle and segregate with sufficient, albeit imperfect, accuracy.
6.3 Delivery and Validation of Function
A major technical hurdle in AC technology is the physical delivery of such massive DNA molecules into recipient cells. The most established method is Microcell-Mediated Chromosome Transfer (MMCT). This technique allows for the transfer of an individual chromosome from a donor cell to a recipient cell. While highly functional, MMCT is a slow, labor-intensive, and inefficient process, making it a significant bottleneck for high-throughput applications and clinical translation.
This multi-step process involves:
1. Inducing the formation of micronuclei in the donor cell line (which contains the HAC) by disrupting the mitotic spindle with drugs like colcemid. Each micronucleus typically contains one or a few chromosomes.
2. Treating the cells with cytochalasin B to disrupt the actin cytoskeleton, causing the micronuclei to be extruded from the cell, encased in a small amount of cytoplasm and plasma membrane. These are known as microcells.
3. Isolating the microcells via centrifugation and filtration.
4. Fusing the purified microcells with recipient cells, usually with an agent like polyethylene glycol (PEG), to transfer the HAC.
Despite its utility, the low efficiency and laborious nature of MMCT remain the primary technical bottleneck hindering the translation of HAC technology into widespread clinical and industrial applications. This has spurred research into alternative and improved transfer methods, including Micronucleated Whole Cell Fusion (MWCF), Isolated Metaphase Chromosome Transfer (iMCT), and hybrid systems that use viral-like particles to package and deliver HACs more efficiently.
Once an AC has been constructed and delivered, its function must be rigorously validated. The key criteria for a successful artificial chromosome are:
1. Autonomous Replication and Segregation: The AC must be maintained as a stable, independent genetic entity, separate from the host chromosomes, through multiple cell generations.
2. Kinetochore Assembly: It must demonstrate the formation of a functional centromere by binding key structural proteins, including the epigenetic marker CENP-A, the initial seeding factor CENP-B (in sequence-dependent models), and essential downstream components like CENP-C and CENP-E.
3. Mitotic Stability: It must be inherited reliably by daughter cells. This is quantified by measuring its loss rate over an extended period of cell culture without any selective pressure. A stable AC should exhibit a very low loss rate, typically less than 2% per cell division.
Fluorescence In Situ Hybridization (FISH) is the primary tool for visualizing HACs within a cell. Using fluorescently labeled DNA probes specific to the HAC’s components (e.g., the input BAC/YAC vector or alphoid DNA), researchers can confirm that the HAC is episomal (i.e., not integrated into a host chromosome), estimate its size relative to native chromosomes, and verify its composition.
Mastering these technical factors—from controlled centromere assembly to efficient delivery—is essential for moving artificial chromosome technology from a powerful research tool toward its ultimate goal as a safe and effective platform for human therapeutics.
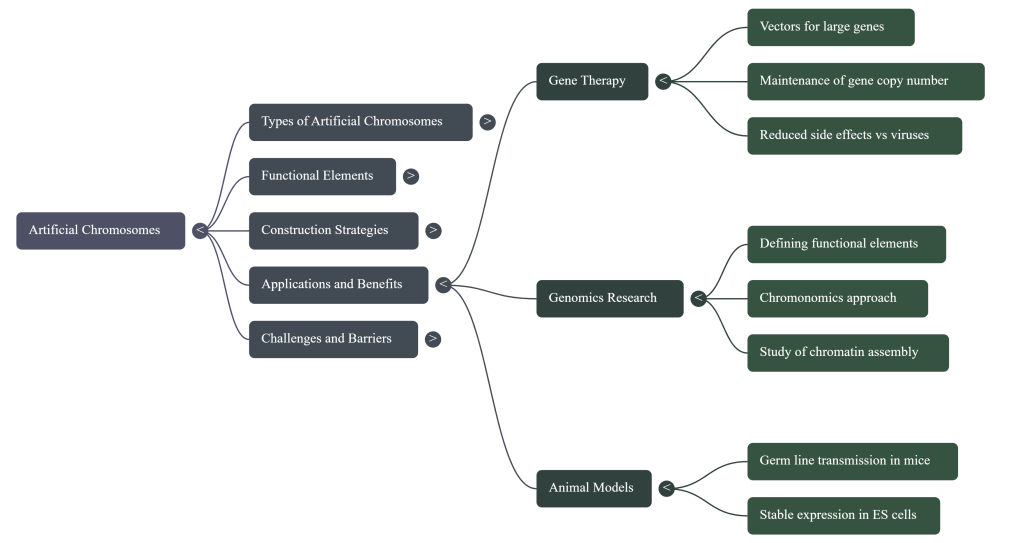
7. Applications and Future Prospects
Beyond its role as an invaluable tool for studying chromosome biology, HAC technology holds transformative potential to revolutionize gene therapy, drug development, and large-scale synthetic biology. As a safe, high-capacity vector, the HAC platform is poised to address challenges that have been intractable with conventional genetic engineering tools.
7.1 Gene and Stem Cell Therapy
HACs represent an ideal vector platform for gene therapy, primarily due to their ability to carry very large genes and their autonomous, episomal maintenance. A prime example is their application for Duchenne Muscular Dystrophy (DMD).
Correcting the Unfixable: The Case of Duchenne Muscular Dystrophy
Duchenne Muscular Dystrophy (DMD) is a devastating genetic disease caused by mutations in the massive dystrophin gene, which is 2.4 megabases (Mb) in size. The sheer size of this gene has made it impossible to correct with traditional gene therapy vectors. In a major breakthrough, scientists have successfully constructed a Human Artificial Chromosome—called a DYS-HAC—that carries the entire, functional dystrophin gene. This DYS-HAC allows for the correct expression of the dystrophin gene, faithfully replicating how the body naturally controls it in different tissues and at different developmental stages. This achievement provides powerful proof-of-principle that HACs can be used to treat complex genetic diseases.
The synergy between HACs and stem cell therapies is another promising frontier. HACs carrying therapeutic genes have been successfully introduced into human mesenchymal stem cells and multipotent germline stem (mGS) cells. This combination allows for the genetic rescue of deficient cells ex vivo, which can then be used for regenerative medicine, providing a powerful platform for treating a wide range of monogenic diseases.
7.2 Transgenic Animal Models
HACs are enabling the creation of “trans-chromosomic” animal models that are “humanized” for specific genetic traits. One of the most successful applications is the development of mice that can produce fully human antibodies. By creating mice with a HAC carrying the human immunoglobulin gene loci and knocking out the corresponding mouse genes, researchers have generated animals that, when challenged with an antigen, produce a diverse repertoire of high-affinity human antibodies. These can be harvested to create monoclonal antibody therapeutics for human use.
Modeling Complex Disease: New Insights into Down Syndrome
Artificial chromosomes are not only for treatment; they are also invaluable research tools. Scientists can use them to study conditions caused by having an extra chromosome, such as Down syndrome, which results from an extra copy of human chromosome 21. Researchers have created “trans-chromosomic” mice that carry an extra, freely-segregating copy of human chromosome 21. These animal models develop symptoms comparable to those seen in humans with Down syndrome. This allows scientists to study the condition in a controlled way and helps them map and identify which specific genes on chromosome 21 contribute to the disorder’s characteristics.
7.3 Synthetic Genomics and Basic Research
HAC technology is a cornerstone of the ambitious Human Genome Project-Write (HGP-Write), which aims to synthesize entire human chromosomes from scratch. HACs provide the foundational platform for designing, building, and testing synthetic human chromosomes, enabling researchers to probe the fundamental principles of genome organization and function.
In cancer research, HACs have been adapted into powerful screening tools. A fluorescently tagged HAC can be used in assays to identify drugs that exploit Chromosome Instability (CIN)—a hallmark of many cancers—by selectively killing cells that exhibit high rates of chromosome loss. These HAC-based systems can also be used with siRNA libraries to uncover novel human CIN genes, providing new targets for cancer therapy.
8.0 Conclusion: The Future of Synthetic Chromosome Engineering
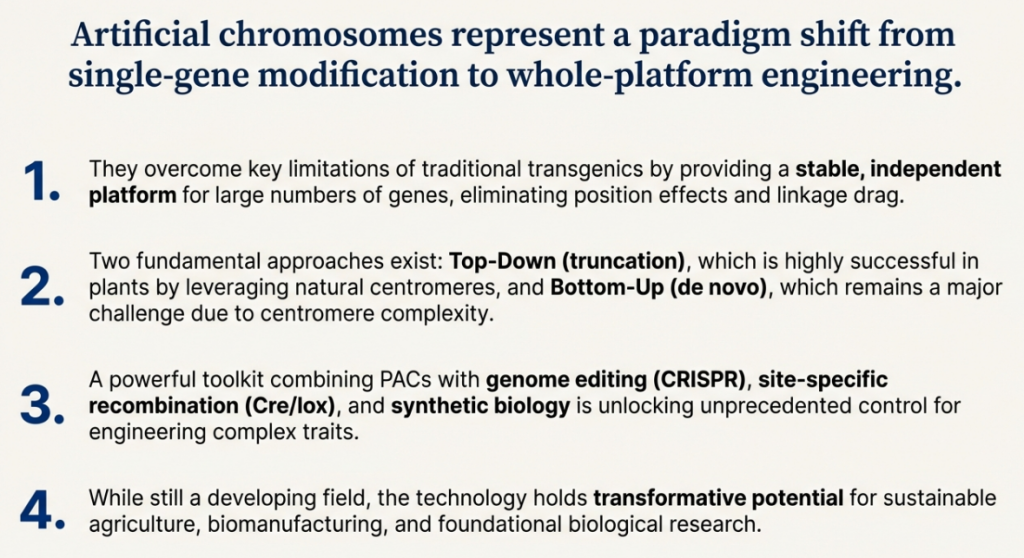
The field of artificial chromosome construction has undergone a remarkable evolution. It has progressed from early, unpredictable methods that relied on the transfection of large, unmanageable stretches of highly repetitive DNA, to modern, highly controlled strategies that leverage a sophisticated, epigenetic understanding of centromere identity. The ability to now build HACs on unique, non-repetitive DNA sequences represents a paradigm shift, opening the door to the creation of truly synthetic, designer chromosomes with fully defined genetic content.
The potential of ACs as next-generation vectors for gene therapy and biomedical research is immense. Their key advantage is a carrying capacity that is orders of magnitude larger than any other vector system. This enables them to carry not just single genes, but entire genomic loci with all their native regulatory elements. This has already been demonstrated with the successful delivery of the massive 2.4 megabase DYSTROPHIN gene, a feat impossible with conventional vectors, offering new hope for treating diseases like Duchenne muscular dystrophy. Furthermore, ACs are invaluable tools for creating “trans-chromosomic” or humanized animal models, which carry human genes or entire chromosomes, providing unparalleled systems for studying human disease, drug metabolism, and toxicology.
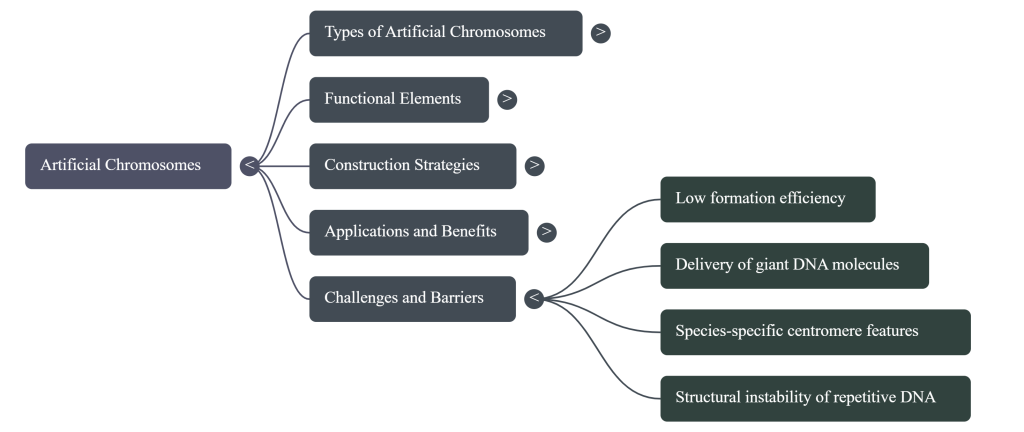
Despite this progress, significant challenges must be addressed before artificial chromosomes can be widely implemented in a clinical setting. The most critical hurdles that remain are the development of robust, efficient, and scalable delivery systems to replace methods like MMCT, and ensuring that therapeutic genes loaded onto these vectors are expressed at the correct levels, in the correct tissues, and for the life of the patient. The single most significant challenge is a practical one: delivery. Scientists must solve the problem of how to get these massive, custom-built DNA molecules into a patient’s cells safely and efficiently. As one leading researcher noted, a number of barriers block their introduction as gene therapy vectors, not the least of which is the necessity for new delivery systems capable of introducing these giant DNA molecules intact into recipient cells.

Despite these challenges, human artificial chromosomes represent a powerful and exciting frontier in medicine. Overcoming these hurdles is therefore the central mission of the field. The development of elegant, efficient delivery platforms and the mastery of long-term epigenetic regulation of gene expression will be the twin pillars upon which the clinical translation of this transformative technology is built.

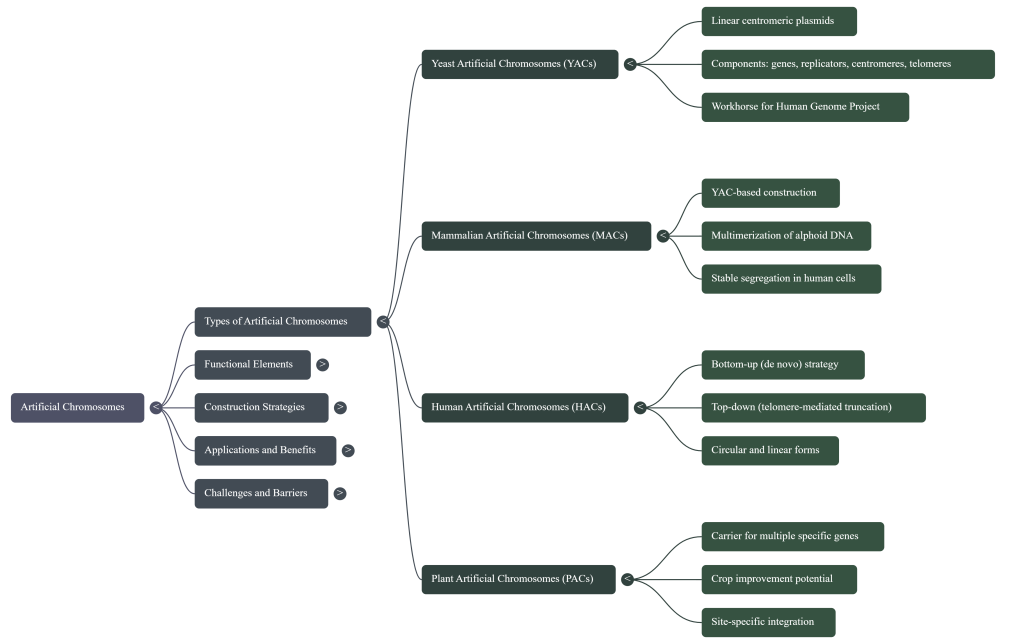
Summary: Types
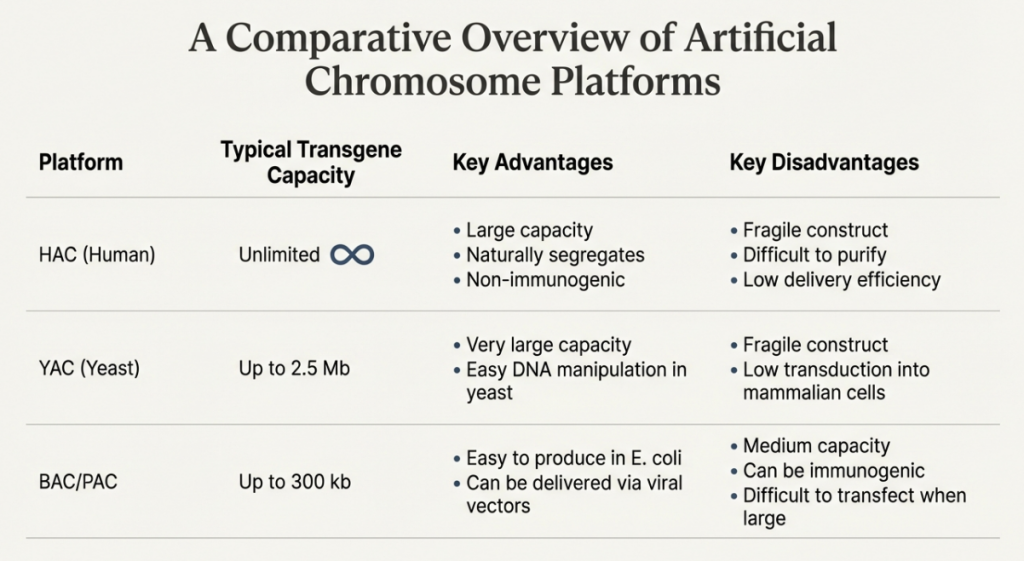
Artificial chromosomes are laboratory-constructed vectors designed to perform the vital functions of natural chromosomes, such as autonomous replication and stable inheritance, while carrying large segments of engineered DNA. They are used as alternatives to raw chromosomes in genetic engineering for cloning, mapping genes, and studying chromosomal functions.

Bacterial Artificial Chromosomes (BACs)
BACs are DNA constructs based on the F-plasmid (fertility plasmid) of Escherichia coli.
• Host and Shape: They are maintained in E. coli as large, circular, single-copy plasmids.
• Capacity: They typically carry inserts between 100 and 350 kb.
• Key Components: Common components include oriS and repE for replication control, parA and parB for stable partitioning during cell division, and selectable markers like chloramphenicol resistance.
• Applications: BACs were essential for the Human Genome Project, used to construct large-insert genomic libraries and map genes onto chromosomes via BAC arrays. They are more stable than other vectors and carry a lower risk of DNA rearrangement.
P1-Derived Artificial Chromosomes (PACs)
PACs were developed in the 1990s as a cloning system derived from the P1 bacteriophage.
• Capacity: Like BACs, they can carry inserts ranging from 100 to 300 kb.
• Structure: They incorporate features of both P1 and F-factor systems and use a packaging site (pac) for in vitro packaging into phage particles.
• Applications: They are frequently used for the genome analysis of complex plants and animals and for studying “phage therapy” and antibiotic mechanisms.
Yeast Artificial Chromosomes (YACs)
YACs were the first artificial chromosomes to be constructed, utilizing the machinery of the budding yeast Saccharomyces cerevisiae.
• Structure and Capacity: They are linear molecules that can clone extremely large DNA fragments, typically from 100 kb up to 3,000 kb (3 Mb).
• Key Components: They must contain a centromere (CEN) for segregation, telomeres (TEL) for protection, and an origin of replication (ORI/ARS).
• Challenges: While they handle the largest inserts, they are often unstable and prone to chimerism (losing parts of the DNA or fusing unrelated sequences) during propagation.
Mammalian (MACs) and Human Artificial Chromosomes (HACs)
MACs and HACs are designed to function as independent genetic elements within mammalian and human cells, respectively.
• Construction Approaches:
◦ Top-down: Created by the truncation of native chromosomes (like human chromosome 21) using telomere-mediated fragmentation.
◦ Bottom-up: Assembled de novo from cloned constituents, primarily centromeric alpha-satellite (alphoid) DNA, telomeres, and replication origins.
• Features: They range in size from 0.5 to 10 Mb. A cell containing an HAC effectively has a 47th chromosome that replicates once per cell cycle without integrating into the host genome, thereby avoiding insertional mutagenesis.
• Applications: They are highly promising for gene and cell therapy (e.g., treating hemophilia or Duchenne muscular dystrophy), studying chromosome instability in cancer, and producing humanized animal models.
Plant Artificial Chromosomes (PACs)
In plants, these are often referred to as minichromosomes.
• Function: They act as “super vectors” that can stack multiple foreign genes to improve crop traits without interfering with native chromosomes or causing “linkage drag”.
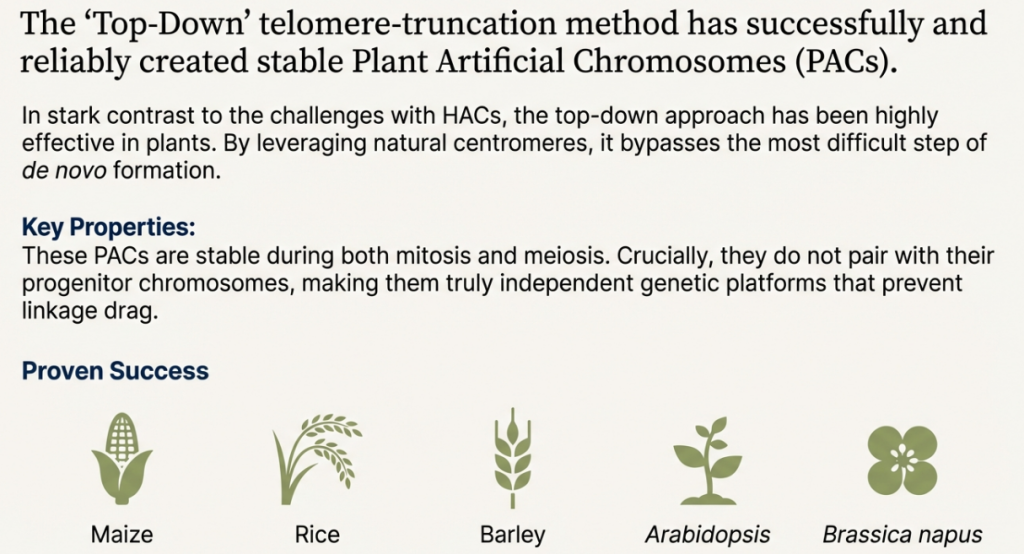
• Construction: Most successful plant minichromosomes have been produced via top-down telomere-mediated chromosomal truncation (TMCT) in species like maize, rice, and Arabidopsis.
• Benefits: They allow for the introduction of complex metabolic pathways for biofuels or medicines and can be transferred between cultivars via sexual hybridization or microprotoplast fusion.
• Challenges: Current plant artificial chromosomes often face issues with transmission efficiency during meiosis, which can lead to the loss of the vector in subsequent generations.

Analogy: If a standard plasmid is like a single sticky note that can be easily lost or stuck anywhere on a page, and a BAC is a single heavy book that requires a specific shelf (E. coli), then an HAC or Plant AC is like an entirely new movable bookcase. It allows you to add a whole new library of information to a room without having to tear down the original walls or reorganize the existing shelves.
Image Summary

References
Asami J, Inoue YU, Terakawa YW, Egusa SF, Inoue T. Bacterial artificial chromosomes as analytical basis for gene transcriptional machineries. Transgenic Res. 2011 Aug;20(4):913-24. doi: 10.1007/s11248-010-9469-3. Epub 2010 Dec 4. PMID: 21132362; PMCID: PMC3139094.
Basu, J., & Willard, H. F. (2006). Human artificial chromosomes: potential applications and clinical considerations. Pediatric Clinics, 53(5), 843-853.
Brown, William RA, P. Joe Mee, and Ming Hong Shen. “Artificial chromosomes: ideal vectors?.” Trends in biotechnology 18.5 (2000): 218-223. DOI: 10.1016/S0167-7799(00)01438-4
Choi, S., Wing, R.A. (2000). The Construction of Bacterial Artificial Chromosome (BAC) Libraries. In: Gelvin, S.B., Schilperoort, R.A. (eds) Plant Molecular Biology Manual. Springer, Dordrecht. https://doi.org/10.1007/978-94-011-4217-5_1
Coulson, A., Waterston, R., Kiff, J. et al. Genome linking with yeast artificial chromosomes. Nature 335, 184–186 (1988). https://doi.org/10.1038/335184a0
Fachinetti, D., Masumoto, H., & Kouprina, N. (2020). Artificial chromosomes. Experimental Cell Research, 396(1), 112302. https://doi.org/10.1016/j.yexcr.2020.11230
Ikeno, M., Grimes, B., Okazaki, T. et al. Construction of YAC–based mammalian artificial chromosomes. Nat Biotechnol 16, 431–439 (1998). https://doi.org/10.1038/nbt0598-431
Kan, M., Huang, T., & Zhao, P. (2022). Artificial chromosome technology and its potential application in plants. Frontiers in Plant Science, 13, 970943. https://doi.org/10.3389/fpls.2022.970943
Kazuki Y, Oshimura M. Human artificial chromosomes for gene delivery and the development of animal models. Mol Ther. 2011 Sep;19(9):1591-601. doi: 10.1038/mt.2011.136. Epub 2011 Jul 12. PMID: 21750534; PMCID: PMC3182354.
Logsdon, G. A., Gambogi, C. W., Liskovykh, M. A., Barrey, E. J., Larionov, V., Miga, K. H., … & Black, B. E. (2019). Human artificial chromosomes that bypass centromeric DNA. Cell, 178(3), 624-639. DOI: 10.1016/j.cell.2019.06.006
Macnab, S., Whitehouse, A. Progress and prospects: human artificial chromosomes. Gene Ther 16, 1180–1188 (2009). https://doi.org/10.1038/gt.2009.102
Ponomartsev SV, Sinenko SA, Tomilin AN. Human Artificial Chromosomes and Their Transfer to Target Cells. Acta Naturae. 2022 Jul-Sep;14(3):35-45. doi: 10.32607/actanaturae.11670. PMID: 36348716; PMCID: PMC9611860.
Sgaramella, V., Eridani, S. Mammalian artificial chromosomes: A review. Cytotechnology 21, 253–261 (1996). https://doi.org/10.1007/BF00365348
Suzuki, N., Nishii, K., Okazaki, T., & Ikeno, M. (2006). Human artificial chromosomes constructed using the bottom-up strategy are stably maintained in mitosis and efficiently transmissible to progeny mice. Journal of Biological Chemistry, 281(36), 26615-26623. DOI: 10.1074/jbc.M603053200
Tao, Q., Zhao, H., Qiu, L. et al. Construction of a full bacterial artificial chromosome (BAC) library of Oryza sativa genome. Cell Res 4, 127–133 (1994). https://doi.org/10.1038/cr.1994.13
Willard, H. F. (2000). Artificial chromosomes coming to life. Science, 290(5495), 1308-1309. DOI: 10.1126/science.290.5495.1308
Yu W, Yau YY, Birchler JA. Plant artificial chromosome technology and its potential application in genetic engineering. Plant Biotechnol J. 2016 May;14(5):1175-82. doi: 10.1111/pbi.12466. Epub 2015 Sep 15. PMID: 26369910; PMCID: PMC11389009.






Pingback: The Genetic Engineer’s Toolkit: A Beginner’s Guide to Cloning Vectors - Aneknowledge.com
Pingback: Comprehensive Guide to DNA Library Types and High-Throughput Cloning Methodologies - Aneknowledge.com